
Acta Horticulturae Sinica ›› 2021, Vol. 48 ›› Issue (6): 1183-1196.doi: 10.16420/j.issn.0513-353x.2020-0510
• Research Notes • Previous Articles Next Articles
Received:2020-12-28
Revised:2021-04-06
Online:2021-06-25
Published:2021-07-08
Contact:
LI Guisheng
E-mail:gui-sheng@jsu.edu.cn
CLC Number:
LI Guisheng. Comparative Analysis of‘Jinyan'and‘Hongyang'Kiwifruit Transcriptomes[J]. Acta Horticulturae Sinica, 2021, 48(6): 1183-1196.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2020-0510
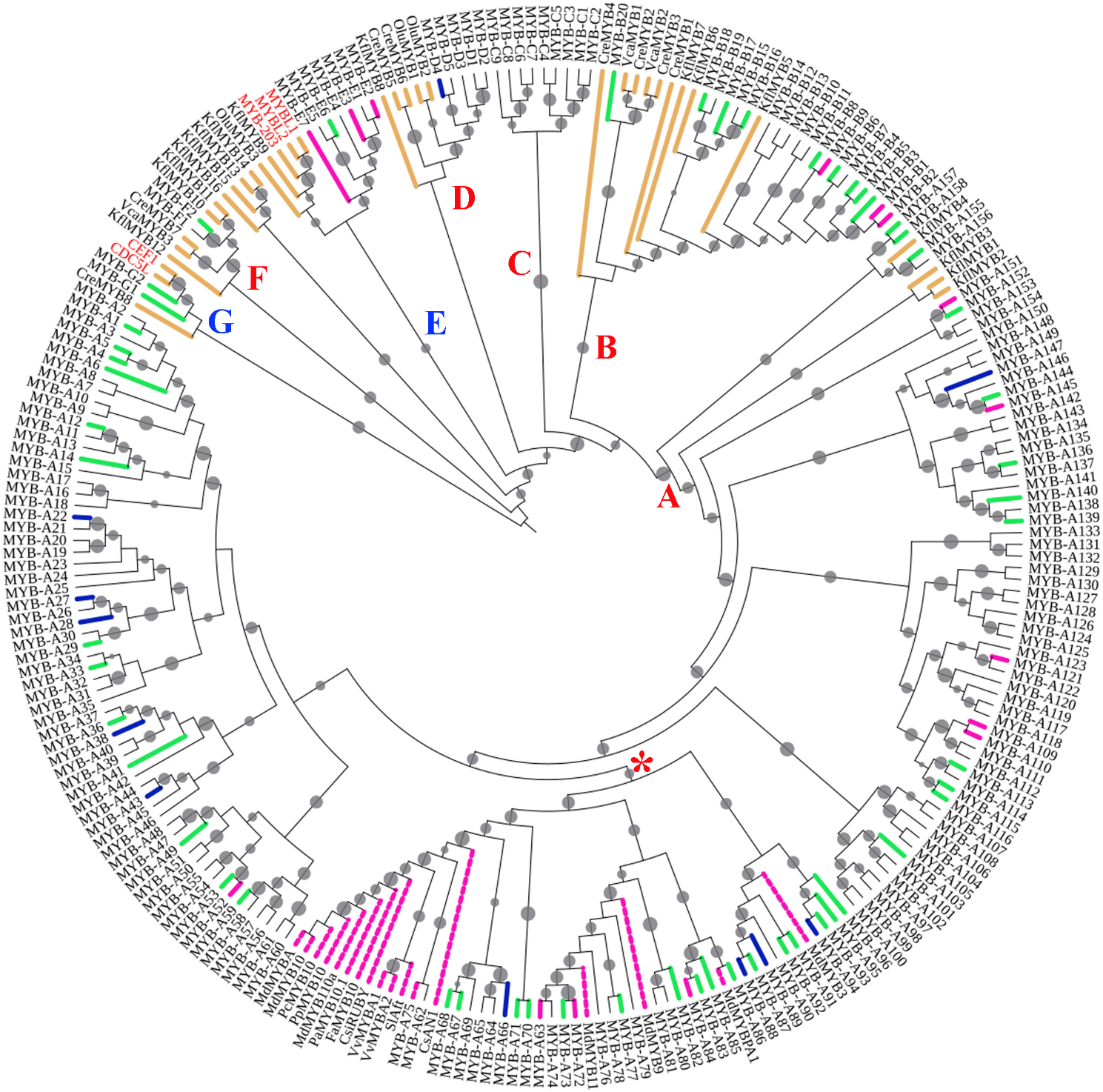
Fig. 1 The phylogenetic analysis of kiwifruit MYB genes Algae genes were indicated by yellow,MYB genes studied and related to anthocyanin synthesis were indicated by dashed pink,genes strongly expressed in‘Hongyang'fruit were indicated by pink,and genes with low expression were indicated by blue solid lines,while genes lacking significant expression difference were indicated by green lines. Fungus and animal gene names were in red. Red letters such as A,B,C,D,and F along branches indicated individual clades of the R2R3 type,while blue letters such as E and G indicated clades of other types. Red asterisks indicated the cluster involved in anthocyanin synthesis. Filled circles on branches indicated bootstrap values above 50%,with a larger circle indicating a higher value.

Fig. 2 Multi-alignments of MYB genes involved in anthocyanin synthesis Two MYB domains were indicated by the grey and the black long bar below the alignment,while the first box indicated the bHLH binding region,and the second box the characteristic motif of MYB proteins related to anthocyanin synthesis.

Fig. 3 The clustering heatmap of genes related to anthocyanin synthesis “*”indicated genes significantly differently expressed between‘Hongyang'and‘Jinyan',with it highlighted in red for MYB genes,and in green for other genes. C1-C4 indicated four major clusters.
| [1] |
Ampomah-Dwamena C, Thrimawithana A H, Dejnoprat S, Lewis D, Espley R V, Allan A C. 2019. A kiwifruit(Actinidia deliciosa)R2R3-MYB transcription factor modulates chlorophyll and carotenoid accumulation. New Phytol, 221 (1):309-325.
doi: 10.1111/nph.15362 |
| [2] | Bi Siqi, An Jianping, Wang Xiaofei, Hao Yujin, Rui Lin, Li Tong, Han Yuepeng, You Chunxiang. 2019. Ethylene response factor MdERF 3 promotes anthocyanin and proanthocyanidin accumulation in apple. Acta Horticulturae Sinica, 46 (12):2277-2285. (in Chinese) |
| 毕思琦, 安建平, 王小非, 郝玉金, 芮麟, 李彤, 韩月彭, 由春香. 2019. 苹果乙烯响应因子MdERF3促进花青苷和原花青苷积累. 园艺学报, 46 (12):2277-2285. | |
| [3] |
Bovy A,de Vos R, Kemper M, Schijlen E, Almenar Pertejo M, Muir S, Collins G, Robinson S, Verhoeyen M, Hughes S, Santos-Buelga C, van Tunen A. 2002. High-flavonol tomatoes resulting from the heterologous expression of the maize transcription factor genes LC and C1. Plant Cell, 14 (10):2509-2526.
doi: 10.1105/tpc.004218 URL |
| [4] |
Butelli E, Licciardello C, Zhang Y, Liu J, Mackay S, Bailey P, Reforgiato-Recupero G, Martin C. 2012. Retrotransposons control fruit-specific,cold-dependent accumulation of anthocyanins in blood oranges. Plant Cell, 24 (3):1242-1255.
doi: 10.1105/tpc.111.095232 URL |
| [5] |
Butelli E, Titta L, Giorgio M, Mock H P, Matros A, Peterek S, Schijlen E G, Hall R D, Bovy A G, Luo J, Martin C. 2008. Enrichment of tomato fruit with health-promoting anthocyanins by expression of select transcription factors. Nat Biotechnol, 26 (11):1301-1308.
doi: 10.1038/nbt.1506 URL |
| [6] | Cao Yuwei, Xu Leifeng, Yang Panpan, Xu Hua, He Guoren, Tang Yuchao, Ren Junfang, Ming Jun. 2019. Differential expression of three R2R3-MYBs genes regulating anthocyanin pigmentation patterns in Lilium spp. Acta Horticulturae Sinica, 46 (5):955-963. (in Chinese) |
| 曹雨薇, 徐雷锋, 杨盼盼, 徐华, 何国仁, 唐玉超, 任君芳, 明军. 2019. 百合花青素苷呈色类型中3种R2R3-MYBs基因的差异表达. 园艺学报, 46 (5):955-963. | |
| [7] |
Chiu L W, Zhou X, Burke S, Wu X, Prior R L, Li L. 2010. The purple cauliflower arises from activation of a MYB transcription factor. Plant Physiol, 154 (3):1470-1480.
doi: 10.1104/pp.110.164160 pmid: 20855520 |
| [8] |
Colanero S, Perata P, Gonzali S. 2020. What's behind purple tomatoes? insight into the mechanisms of anthocyanin synthesis in tomato fruits. Plant Physiol, 182 (4):1841-1853.
doi: 10.1104/pp.19.01530 pmid: 31980573 |
| [9] |
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L. 2010. MYB transcription factors in Arabidopsis. Trends Plant Sci, 15 (10):573-581.
doi: 10.1016/j.tplants.2010.06.005 URL |
| [10] |
Espley R V, Brendolise C, Chagne D, Kutty-Amma S, Green S, Volz R, Putterill J, Schouten H J, Gardiner S E, Hellens R P, Allan A C. 2009. Multiple repeats of a promoter segment causes transcription factor autoregulation in red apples. Plant Cell, 21 (1):168-183.
doi: 10.1105/tpc.108.059329 URL |
| [11] |
Fu Z Z, Shang H Q, Jiang H, Gao J, Dong X Y, Wang H J, Li Y M, Wang L M, Zhang J, Shu Q Y, Chao Y C, Xu M L, Wang R, Wang L S, Zhang H C. 2020. Systematic identification of the light-quality responding anthocyanin synthesis-related transcripts in petunia petals. Horticultural Plant Journal, 6 (6):428-438.
doi: 10.1016/j.hpj.2020.11.006 URL |
| [12] | Gambi F, Pilkington S M, McAtee P A, Donati I, Schaffer R J, Montefiori M, Spinelli F, Burdon J. 2018. Fruit of three kiwifruit(Actinidia chinensis)cultivars differ in their degreening response to temperature after harvest. Postharvest Biology & Technology, 141:16-23. |
| [13] |
Gates D J, Strickler S R, Mueller L A, Olson B J, Smith S D. 2016. Diversification of R2R3-MYB transcription factors in the tomato family Solanaceae. Journal Mol Evol, 83 (1-2):26-37.
doi: 10.1007/s00239-016-9756-6 URL |
| [14] |
Gonzalez A, Zhao M, Leavitt J M, Lloyd A M. 2008. Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex inArabidopsis seedlings. Plant Journal, 53 (5):814-827.
doi: 10.1111/tpj.2008.53.issue-5 URL |
| [15] |
Ho L C. 1996. The mechanism of assimilate partitioning and carbohydrate compartmentation in fruit in relation to the quality and yield of tomato. Journal Exp Bot, 47:1239-1243.
doi: 10.1093/jxb/47.Special_Issue.1239 URL |
| [16] |
Holton T A, Cornish E C. 1995. Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell, 7 (7):1071-1083.
doi: 10.2307/3870058 URL |
| [17] |
Hsu C C, Chen Y Y, Tsai W C, Chen W H, Chen H H. 2015. Three R2R3-MYB transcription factors regulate distinct floral pigmentation patterning in Phalaenopsis spp. Plant Physiol, 168 (1):175-191.
doi: 10.1104/pp.114.254599 URL |
| [18] |
Hsu C C, Su C J, Jeng M F, Chen W H, Chen H H. 2019. A HORT1 retrotransposon insertion in the PeMYB11 promoter causes Harlequin/Black flowers in Phalaenopsis orchids. Plant Physiol, 180 (3):1535-1548.
doi: 10.1104/pp.19.00205 URL |
| [19] | Huang S, Ding J, Deng D, Tang W, Sun H, Liu D, Zhang L, Niu X, Zhang X, Meng M, Yu J, Liu J, Han Y, Shi W, Zhang D, Cao S, Wei Z, Cui Y, Xia Y, Zeng H, Bao K, Lin L, Min Y, Zhang H, Miao M, Tang X, Zhu Y, Sui Y, Li G, Yue J, Sun J, Liu F, Zhou L, Lei L, Zheng X, Liu M, Huang L, Song J, Xu C, Li J, Ye K, Zhong S, Lu B R, He G, Xiao F, Wang H L, Zheng H, Fei Z, Liu Y. 2013. Draft genome of the kiwifruit Actinidia chinensis. Nat Commun, 4:10.1038/ncomms3640. |
| [20] |
Inagaki Y, Hisatomi Y, Suzuki T, Kasahara K, Iida S. 1994. Isolation of a Suppressor-mutator/Enhancer-like transposable element, Tpn1,from Japanese morning glory bearing variegated flowers. Plant Cell, 6 (3):375-383.
pmid: 8180498 |
| [21] |
Itoh Y, Higeta D, Suzuki A, Yoshida H, Ozeki Y. 2002. Excision of transposable elements from the chalcone isomerase and dihydroflavonol 4-reductase genes may contribute to the variegation of the yellow-flowered carnation(Dianthus caryophyllus). Plant Cell Physiol, 43 (5):578-585.
doi: 10.1093/pcp/pcf065 URL |
| [22] |
Johnson E T, Ryu S, Yi H, Shin B, Cheong H, Choi G. 2001. Alteration of a single amino acid changes the substrate specificity of dihydroflavonol 4-reductase. Plant Journal, 25 (3):325-333.
pmid: 11208024 |
| [23] |
Krga I, Milenkovic D. 2019. Anthocyanins:from sources and bioavailability to cardiovascular-health benefits and molecular mechanisms of action. Journal Agric Food Chem, 67 (7):1771-1783.
doi: 10.1021/acs.jafc.8b06737 URL |
| [24] | Li B, Xia Y, Wang Y, Qin G, Tian S. 2017a. Characterization of genes encoding key enzymes involved in anthocyanin metabolism of kiwifruit during storage period. Front Plant Sci, 8:10.3389/fpls.2017.00341. |
| [25] | Li M, Verboven P, Buchsbaum A, Cantre D, Nicola B, Heyes J, Mowat A, East A. 2015. Characterizing kiwifruit(Actinidia sp.)near skin cellular structures using optical coherence tomography. Postharvest Biology & Technology, 110:247-256. |
| [26] |
Li W, Ding Z, Ruan M, Yu X, Peng M, Liu Y. 2017b. Kiwifruit R2R3-MYB transcription factors and contribution of the novel AcMYB75 to red kiwifruit anthocyanin biosynthesis. Sci Rep, 7 (1):10.1038/s41598-017-16905-1.
doi: 10.1038/s41598-017-00036-8 URL |
| [27] |
Li W, Liu Y, Zeng S, Xiao G, Wang G, Wang Y, Peng M, Huang H. 2015. Gene expression profiling of development and anthocyanin accumulation in kiwifruit(Actinidia chinensis)based on transcriptome sequencing. PLoS ONE, 10 (8):e0136439.
doi: 10.1371/journal.pone.0136439 URL |
| [28] |
Li Y, Cui W, Wang R, Lin M, Zhong Y, Sun L, Qi X, Fang J. 2019. MicroRNA858-mediated regulation of anthocyanin biosynthesis in kiwifruit (Actinidia arguta)based on small RNA sequencing. PLoS ONE, 14 (5):e0217480.
doi: 10.1371/journal.pone.0217480 URL |
| [29] |
Li Y, Fang J, Qi X, Lin M, Zhong Y, Sun L. 2018. A key structural gene, AaLDOX,is involved in anthocyanin biosynthesis in all red-fleshed kiwifruit(Actinidia arguta)based on transcriptome analysis. Gene, 648:31-41.
doi: 10.1016/j.gene.2018.01.022 URL |
| [30] | Liu Y, Qi Y, Zhang A, Wu H, Liu Z, Ren X. 2019. Molecular cloning and functional characterization of AcGST1,an anthocyanin-related glutathione S-transferase gene in kiwifruit (Actinidia chinensis). Plant Mol Biol, 100 (4-5):451-465. |
| [31] | Liu Y, Zhou B, Qi Y, Chen X, Liu C, Liu Z, Ren X. 2017. Expression differences of pigment structural genes and transcription factors explain flesh coloration in three contrasting kiwifruit cultivars. Front Plant Sci, 8:10.3389/fpls.2017.01507. |
| [32] | Lu Jing, Ma Qijun, Liu Xiao, Kang Hui, Liu Yajing, Hao Yujin, You Chunxiang. 2019. Studies on the regulation of anthocyanin accumulation by apple sucrose transporter MdSUT2. Acta Horticulturae Sinica, 46 (1):1-10. (in Chinese) |
| 路静, 马齐军, 刘晓, 康慧, 刘亚静, 郝玉金, 由春香. 2019. 苹果蔗糖转运蛋白MdSUT2调控花青苷积累的研究. 园艺学报, 46 (1):1-10. | |
| [33] |
Man Y P, Wang Y C, Li Z Z, Jiang Z W, Yang H L, Gong J J, He S S, Wu S Q, Yang Z Q, Zheng J, Wang Z Y. 2015. High-temperature inhibition of biosynthesis and transportation of anthocyanins results in the poor red coloration in red-fleshed Actinidia chinensis. Physiol Plant, 153 (4):565-583.
doi: 10.1111/ppl.2015.153.issue-4 URL |
| [34] |
McClure M A, Smith C, Elton P. 1996. Parameterization studies for the SAM and HMMER methods of hidden Markov model generation. Proc Int Conf Intell Syst Mol Biol, 4:155-164.
pmid: 8877515 |
| [35] |
Mehrtens F, Kranz H, Bednarek P, Weisshaar B. 2005. The Arabidopsis transcription factor MYB12 is a flavonol-specific regulator of phenylpropanoid biosynthesis. Plant Physiol, 138 (2):1083-1096.
pmid: 15923334 |
| [36] |
Meng J X, Yan Gao Y, Han M L, Liu P Y, Yang C, Shen T, Li H H. 2020. In vitro anthocyanin induction and metabolite analysis in Malus spectabilis leaves under low nitrogen conditions. Horticultural Plant Journal, 6 (5):284-292.
doi: 10.1016/j.hpj.2020.06.004 URL |
| [37] |
Montefiori M, Espley R V, Stevenson D, Cooney J, Datson P M, Saiz A, Atkinson R G, Hellens R P, Allan A C. 2010. Identification and characterisation of F3GT1 and F3GGT1,two glycosyltransferases responsible for anthocyanin biosynthesis in red-fleshed kiwifruit(Actinidia chinensis). Plant Journal, 65 (1):106-118.
doi: 10.1111/j.1365-313X.2010.04409.x URL |
| [38] |
Povero G, Gonzali S, Bassolino L, Mazzucato A, Perata P. 2011. Transcriptional analysis in high-anthocyanin tomatoes reveals synergistic effect of Aft and atv genes. Journal Plant Physiol, 168 (3):270-279.
doi: 10.1016/j.jplph.2010.07.022 URL |
| [39] |
Price M N, Dehal P S, Arkin A P. 2009. FastTree:computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol, 26 (7):1641-1650.
doi: 10.1093/molbev/msp077 URL |
| [40] |
Ruiz-Sanchez E, Sosa V. 2015. Origin and evolution of fleshy fruit in woody bamboos. Mol Phylogenet Evol, 91:123-134.
doi: 10.1016/j.ympev.2015.05.020 pmid: 26048705 |
| [41] | Shinozaki Y, Nicolas P, Fernandez-Pozo N, Ma Q, Evanich D J, Shi Y, Xu Y, Zheng Y, Snyder S I, Martin L B B, Ruiz-May E, Thannhauser T W, Chen K, Domozych D S, Catala C, Fei Z, Mueller L A, Giovannoni J J, Rose J K C. 2018. High-resolution spatiotemporal transcriptome mapping of tomato fruit development and ripening. Nat Commun, 9 (1):10.1038/s41467-017-02782-9. |
| [42] | Song Yang, Liu Hongdi, Wang Haibo, Zhang Hongjun, Liu Fengzhi. 2019. Molecular cloning and functional characterization of anthocyanin synthesis related genes VcTTG1of blueberry. Acta Horticulturae Sinica, 46 (7):1270-1278. (in Chinese) |
| 宋杨, 刘红弟, 王海波, 张红军, 刘凤之. 2019. 越橘花青苷合成相关基因 VcTTG1的克隆与功能鉴定. 园艺学报, 46 (7):1270-1278. | |
| [43] |
Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B. 2007. Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant Journal, 50 (4):660-677.
pmid: 17419845 |
| [44] |
Sun C, Deng L, Du M, Zhao J, Chen Q, Huang T, Jiang H, Li C, Li C. 2020. A transcriptional network promotes anthocyanin biosynthesis in tomato flesh. Mol Plant, 13 (1):42-58.
doi: 10.1016/j.molp.2019.10.010 URL |
| [45] |
Thompson J D, Higgins D G, Gibson T J. 1994. CLUSTAL W:improving the sensitivity of progressive multiple sequence alignment through sequence weighting,position-specific gap penalties and weight matrix choice. Nucleic Acids Res, 22 (22):4673-4680.
pmid: 7984417 |
| [46] |
Wang L, Tang W, Hu Y, Zhang Y, Sun J, Guo X, Lu H, Yang Y, Fang C, Niu X, Yue J, Fei Z, Liu Y. 2019. A MYB/bHLH complex regulates tissue-specific anthocyanin biosynthesis in the inner pericarp of red-centered kiwifruit Actinidia chinensis cv. Hongyang. Plant Journal, 99 (2):359-378.
doi: 10.1111/tpj.2019.99.issue-2 URL |
| [47] |
Xu Z S, Yang Q Q, Feng K, Xiong A S. 2019. Changing carrot color:insertions in DcMYB7 alter the regulation of anthocyanin biosynthesis and modification. Plant Physiol, 181 (1):195-207.
doi: 10.1104/pp.19.00523 URL |
| [48] | Yin Cui-bo. 2008. Preliminary studies on biological characters and the rule of fruit growth and development of Hongyang kiwifruit[M. D. Dissertation]. Chengdu: Sichuan Agricultural University. (in Chinese) |
| 尹翠波. 2008. 红阳猕猴桃生物学特性及果实生长发育规律的初步研究[硕士论文]. 成都: 四川农业大学. | |
| [49] |
Yu M, Man Y, Wang Y. 2019. Light- and temperature-induced expression of an R2R3-MYB gene regulates anthocyanin biosynthesis in red-fleshed kiwifruit. Int Journal Mol Sci, 20 (20):10.3390/ijms20205228.
doi: 10.3390/ijms20010010 URL |
| [50] | Zhong Caihong, Zhang Peng, Han Fei, Li Dawei. 2015. Studies on characterization of fruit development of interspecific hybrid cultivar‘Jinyan'. Journal of Fruit Science, 32 (6):1152-1160. (in Chinese) |
| 钟彩虹, 张鹏, 韩飞, 李大卫. 2015. 猕猴桃种间杂交新品种‘金艳'的果实发育特征. 果树学报, 32 (6):1152-1160. |
| [1] | YE Zimao, SHEN Wanxia, LIU Mengyu, WANG Tong, ZHANG Xiaonan, YU Xin, LIU Xiaofeng, and ZHAO Xiaochun, . Effect of R2R3-MYB Transcription Factor CitMYB21 on Flavonoids Biosynthesis in Citrus [J]. Acta Horticulturae Sinica, 2023, 50(2): 250-264. |
| [2] | YU Tingting, LI Huan, NING Yuansheng, SONG Jianfei, PENG Lulin, JIA Junqi, ZHANG Weiwei, and YANG Hongqiang. Genome-wide Identification of GRAS Gene Family in Apple and Expression Analysis of Its Response to Auxin [J]. Acta Horticulturae Sinica, 2023, 50(2): 397-409. |
| [3] | ZHAI Hanhan, ZHAI Yujie, TIAN Yi, ZHANG Ye, YANG Li, WEN Zhiliang, CHEN Haijiang. Genome-wide Identification of Peach SAUR Gene Family and Characterization of PpSAUR5 Gene [J]. Acta Horticulturae Sinica, 2023, 50(1): 1-14. |
| [4] | YUAN Xin, XU Yunhe, ZHANG Yupei, SHAN Nan, CHEN Chuying, WAN Chunpeng, KAI Wenbin, ZHAI Xiawan, CHEN Jinyin, GAN Zengyu. Studies on AcAREB1 Regulating the Expression of AcGH3.1 During Postharvest Ripening of Kiwifruit [J]. Acta Horticulturae Sinica, 2023, 50(1): 53-64. |
| [5] | SONG Fang, CHEN Qi, YUAN Yanliang, CHEN Sha, YIN Haijun, and JIANG Yingchun, . A New Yellow-fleshed Kiwifruit Cultivar‘Xianwo 1’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 47-48. |
| [6] | QI Yongjie, GAO Zhenghui, MA Na, WANG Qingming, KE Fanjun, CHEN Qian, and XU Yiliu, . A New Yellow Flesh and Resistant to Canker Disease Kiwifruit Cultivar ‘Wannong Jinguo’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 49-50. |
| [7] | ZHANG Huiqin, LOU Guorong, LU Linghong, GU Xianbin, SONG Genhua, and XIE Ming. A New Yellow-fleshed Kiwifruit Cultivar‘Jinyi’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 51-52. |
| [8] | LI Maofu, YANG Yuan, WANG Hua, FAN Youwei, SUN Pei, JIN Wanmei. Analysis the Function of R2R3 MYB Transcription Factor RhMYB113c on Regulating Anthocyanin Synthesis in Rosa hybrida [J]. Acta Horticulturae Sinica, 2022, 49(9): 1957-1966. |
| [9] | GAO Yanlong, WU Yuxia, ZHANG Zhongxing, WANG Shuangcheng, ZHANG Rui, ZHANG De, WANG Yanxiu. Bioinformatics Analysis of Apple ELO Gene Family and Its Expression Analysis Under Low Temperature Stress [J]. Acta Horticulturae Sinica, 2022, 49(8): 1621-1636. |
| [10] | LIU Jinming, GUO Caihua, YUAN Xing, KANG Chao, QUAN Shaowen, NIU Jianxin. Genome-wide Identification of Dof Family Genes and Expression Analysis Sepal Persistent and Abscission in Pear [J]. Acta Horticulturae Sinica, 2022, 49(8): 1637-1649. |
| [11] | XU Haifeng, WANG Zhongtang, CHEN Xin, LIU Zhiguo, WANG Lihu, LIU Ping, LIU Mengjun, ZHANG Qiong. The Analyses of Target Metabolomics in Flavonoid and Its Potential MYB Regulation Factors During Coloring Period of Winter Jujube [J]. Acta Horticulturae Sinica, 2022, 49(8): 1761-1771. |
| [12] | QIAN Jieyu, JIANG Lingli, ZHENG Gang, CHEN Jiahong, LAI Wuhao, XU Menghan, FU Jianxin, ZHANG Chao. Identification and Expression Analysis of MYB Transcription Factors Regulating the Anthocyanin Biosynthesis in Zinnia elegans and Function Research of ZeMYB9 [J]. Acta Horticulturae Sinica, 2022, 49(7): 1505-1518. |
| [13] | LÜ Zhengxin, HE Yanqun, JIA Dongfeng, HUANG Chunhui, ZHONG Min, LIAO Guanglian, ZHU Yi, YUAN Kaichang, LIU Chuanhao, XU Xiaobiao. Genetic Diversity Analysis of Phenotypic Traits for Kiwifruit Germplasm Resources [J]. Acta Horticulturae Sinica, 2022, 49(7): 1571-1581. |
| [14] | ZHANG Lugang, LU Qianqian, HE Qiong, XUE Yihua, MA Xiaomin, MA Shuai, NIE Shanshan, YANG Wenjing. Creation of Novel Germplasm of Purple-orange Heading Chinese Cabbage [J]. Acta Horticulturae Sinica, 2022, 49(7): 1582-1588. |
| [15] | MA Weifeng, LI Yanmei, MA Zonghuan, CHEN Baihong, MAO Juan. Identification of Apple POD Gene Family and Functional Analysis of MdPOD15 Gene [J]. Acta Horticulturae Sinica, 2022, 49(6): 1181-1199. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd
