
Acta Horticulturae Sinica ›› 2022, Vol. 49 ›› Issue (8): 1621-1636.doi: 10.16420/j.issn.0513-353x.2021-0545
• Research Papers • Next Articles
GAO Yanlong, WU Yuxia, ZHANG Zhongxing, WANG Shuangcheng, ZHANG Rui, ZHANG De, WANG Yanxiu*( )
)
Received:2022-04-27
Revised:2022-06-06
Online:2022-08-25
Published:2022-09-05
Contact:
WANG Yanxiu
E-mail:wangxy@gsau.edu.cn
CLC Number:
GAO Yanlong, WU Yuxia, ZHANG Zhongxing, WANG Shuangcheng, ZHANG Rui, ZHANG De, WANG Yanxiu. Bioinformatics Analysis of Apple ELO Gene Family and Its Expression Analysis Under Low Temperature Stress[J]. Acta Horticulturae Sinica, 2022, 49(8): 1621-1636.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2021-0545
| 基因登录号 Accession No. | 基因名称 Gene name | 序列(5′-3′) Sequence | |
|---|---|---|---|
| MD02G1155100 | MdELO1 | F:TCCTACGTGTACTACCTGTCCAGATTC | R:AGAGGAACGAGATGACGGTGAGG |
| MD08G1036600 | MdELO2 | F:CAGTCTACCTCATCTTCATGTCGTTCC | R:AGAGCAGAGTGGTGGAGAGTATGG |
| MD08G1151400 | MdELO3 | F:CCGCCATCGCCGTCTACATAAC | R:GGCTGGAATAGGTCCGAGAGGTAC |
| MD08G1227900 | MdELO4 | F:ATTTGGCTCCACACATCGCAGTC | R:CGGTCACCCTTCTCTTCCACTTTG |
| MD15G1269600 | MdELO5 | F:CCGCCATCGCCGTCTACATAAC | R:GGCTGGAATAGGTCCGAGAGGTAC |
| MD15G1421700 | MdELO6 | F:CAACCTCAACCTCCTCCTCCTCTC | R:GCAAACAGATAATCCAGAAGGGGTAGG |
| MD15G1421900 | MdELO7 | F:CACAGCCGTCCACAACCTCATC | R:AGGTGTTGGGTGGGAAGCAGAG |
Table 1 qRT-PCR primer sequence
| 基因登录号 Accession No. | 基因名称 Gene name | 序列(5′-3′) Sequence | |
|---|---|---|---|
| MD02G1155100 | MdELO1 | F:TCCTACGTGTACTACCTGTCCAGATTC | R:AGAGGAACGAGATGACGGTGAGG |
| MD08G1036600 | MdELO2 | F:CAGTCTACCTCATCTTCATGTCGTTCC | R:AGAGCAGAGTGGTGGAGAGTATGG |
| MD08G1151400 | MdELO3 | F:CCGCCATCGCCGTCTACATAAC | R:GGCTGGAATAGGTCCGAGAGGTAC |
| MD08G1227900 | MdELO4 | F:ATTTGGCTCCACACATCGCAGTC | R:CGGTCACCCTTCTCTTCCACTTTG |
| MD15G1269600 | MdELO5 | F:CCGCCATCGCCGTCTACATAAC | R:GGCTGGAATAGGTCCGAGAGGTAC |
| MD15G1421700 | MdELO6 | F:CAACCTCAACCTCCTCCTCCTCTC | R:GCAAACAGATAATCCAGAAGGGGTAGG |
| MD15G1421900 | MdELO7 | F:CACAGCCGTCCACAACCTCATC | R:AGGTGTTGGGTGGGAAGCAGAG |
| 基因登录号 Accession No. | 基因名称 Gene name | 大小/aa Size | 分子量 Molecular weight | 等电点 Isoelectric point | 亚细胞定位 Subcellular localization | 不稳定系数 Instability index |
|---|---|---|---|---|---|---|
| MD02G1155100 | MdELO1 | 305 | 34 129.36 | 10.18 | 质膜Plasma membrane | 43.13 |
| MD08G1036600 | MdELO2 | 211 | 24 143.86 | 11.62 | 质膜/细胞核Plasma membrane/nucleus | 75.39 |
| MD08G1151400 | MdELO3 | 302 | 33 869.14 | 10.23 | 质膜Plasma membrane | 41.39 |
| MD08G1227900 | MdELO4 | 265 | 30 759.69 | 9.43 | 叶绿体Chloroplast | 40.60 |
| MD15G1269600 | MdELO5 | 302 | 33 869.14 | 10.23 | 质膜Plasma membrane | 41.39 |
| MD15G1421700 | MdELO6 | 270 | 31 346.38 | 9.50 | 叶绿体Chloroplast | 38.86 |
| MD15G1421900 | MdELO7 | 273 | 31 159.93 | 9.53 | 叶绿体Chloroplast | 38.90 |
Table 2 Analysis of physicochemical properties of apple MdELO gene family
| 基因登录号 Accession No. | 基因名称 Gene name | 大小/aa Size | 分子量 Molecular weight | 等电点 Isoelectric point | 亚细胞定位 Subcellular localization | 不稳定系数 Instability index |
|---|---|---|---|---|---|---|
| MD02G1155100 | MdELO1 | 305 | 34 129.36 | 10.18 | 质膜Plasma membrane | 43.13 |
| MD08G1036600 | MdELO2 | 211 | 24 143.86 | 11.62 | 质膜/细胞核Plasma membrane/nucleus | 75.39 |
| MD08G1151400 | MdELO3 | 302 | 33 869.14 | 10.23 | 质膜Plasma membrane | 41.39 |
| MD08G1227900 | MdELO4 | 265 | 30 759.69 | 9.43 | 叶绿体Chloroplast | 40.60 |
| MD15G1269600 | MdELO5 | 302 | 33 869.14 | 10.23 | 质膜Plasma membrane | 41.39 |
| MD15G1421700 | MdELO6 | 270 | 31 346.38 | 9.50 | 叶绿体Chloroplast | 38.86 |
| MD15G1421900 | MdELO7 | 273 | 31 159.93 | 9.53 | 叶绿体Chloroplast | 38.90 |

Fig. 2 Multiple sequence alignment of apple MdELO gene family members Sequence identity of 100% is marked in dark blue,> 75% in pink,and > 50% in light blue. The red underline represents the ELO domain.
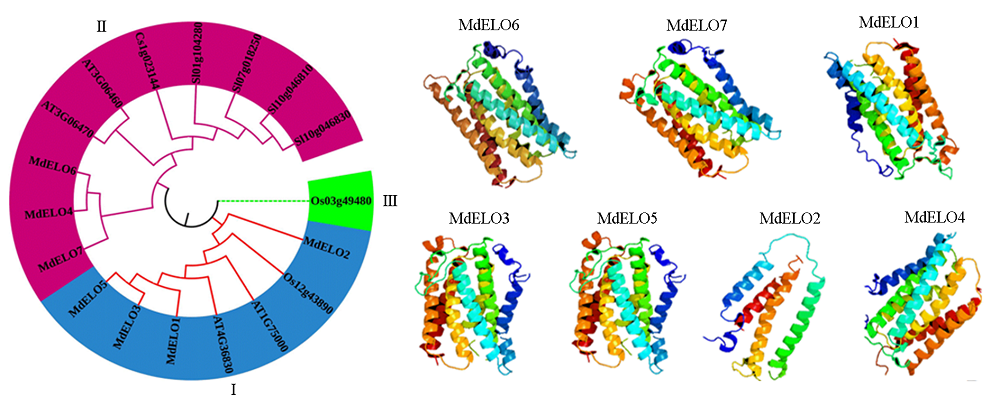
Fig. 3 Phylogenetic tree and tertiary structure of apple MdELO gene family At:Arabidopsis thaliana;Sl:Solanum lycopersicum;Cs:Citrus reticulata Blanco;Os:Oryza sativa.
| 旁系同源基因对位点 Paralogous gene pair sites | Ka | Ks | Ka/Ks |
|---|---|---|---|
| MdELO1/MdELO2 | 0.427957 | 0.867576 | 0.493279 |
| MdELO1/MdELO3 | 0.026568 | 0.236089 | 0.112535 |
| MdELO1/MdELO4 | 0.755620 | 1.160104 | 0.651338 |
| MdELO1/MdELO5 | 0.026568 | 0.236089 | 0.112535 |
| MdELO1/MdELO6 | 0.745098 | 1.256016 | 0.593223 |
| MdELO1/MdELO7 | 0.671194 | 1.008214 | 0.665726 |
| MdELO2/MdELO3 | 0.368452 | 0.856895 | 0.429984 |
| MdELO2/MdELO4 | 0.701287 | 1.012445 | 0.692667 |
| MdELO2/MdELO5 | 0.368452 | 0.856895 | 0.429984 |
| MdELO2/MdELO6 | 0.700478 | 1.012911 | 0.691550 |
| MdELO2/MdELO7 | 0.743613 | 0.838943 | 0.886369 |
| MdELO3/MdELO4 | 0.718204 | 1.073463 | 0.669053 |
| MdELO3/MdELO5 | 0.701591 | 1.069425 | 0.656045 |
| MdELO3/MdELO6 | 0.706619 | 1.203640 | 0.587069 |
| MdELO3/MdELO7 | 0.636910 | 0.936570 | 0.680045 |
| MdELO4/MdELO5 | 0.718204 | 1.073463 | 0.669053 |
| MdELO4/MdELO6 | 0.018535 | 0.220872 | 0.083918 |
| MdELO4/MdELO7 | 0.244042 | 0.942442 | 0.258947 |
| MdELO5/MdELO6 | 0.706619 | 1.203640 | 0.587069 |
| MdELO5/MdELO7 | 0.636910 | 0.936570 | 0.680045 |
| MdELO6/MdELO7 | 0.246400 | 0.868520 | 0.283700 |
Table 3 Ka/Ks of MdELO gene
| 旁系同源基因对位点 Paralogous gene pair sites | Ka | Ks | Ka/Ks |
|---|---|---|---|
| MdELO1/MdELO2 | 0.427957 | 0.867576 | 0.493279 |
| MdELO1/MdELO3 | 0.026568 | 0.236089 | 0.112535 |
| MdELO1/MdELO4 | 0.755620 | 1.160104 | 0.651338 |
| MdELO1/MdELO5 | 0.026568 | 0.236089 | 0.112535 |
| MdELO1/MdELO6 | 0.745098 | 1.256016 | 0.593223 |
| MdELO1/MdELO7 | 0.671194 | 1.008214 | 0.665726 |
| MdELO2/MdELO3 | 0.368452 | 0.856895 | 0.429984 |
| MdELO2/MdELO4 | 0.701287 | 1.012445 | 0.692667 |
| MdELO2/MdELO5 | 0.368452 | 0.856895 | 0.429984 |
| MdELO2/MdELO6 | 0.700478 | 1.012911 | 0.691550 |
| MdELO2/MdELO7 | 0.743613 | 0.838943 | 0.886369 |
| MdELO3/MdELO4 | 0.718204 | 1.073463 | 0.669053 |
| MdELO3/MdELO5 | 0.701591 | 1.069425 | 0.656045 |
| MdELO3/MdELO6 | 0.706619 | 1.203640 | 0.587069 |
| MdELO3/MdELO7 | 0.636910 | 0.936570 | 0.680045 |
| MdELO4/MdELO5 | 0.718204 | 1.073463 | 0.669053 |
| MdELO4/MdELO6 | 0.018535 | 0.220872 | 0.083918 |
| MdELO4/MdELO7 | 0.244042 | 0.942442 | 0.258947 |
| MdELO5/MdELO6 | 0.706619 | 1.203640 | 0.587069 |
| MdELO5/MdELO7 | 0.636910 | 0.936570 | 0.680045 |
| MdELO6/MdELO7 | 0.246400 | 0.868520 | 0.283700 |
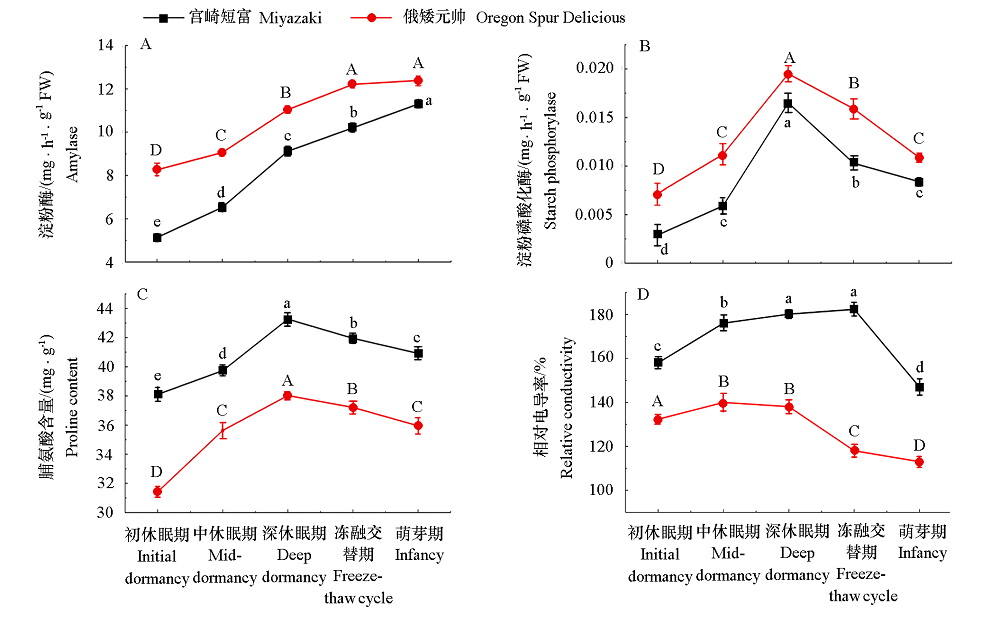
Fig. 8 Changes of amylase,starch phosphorylase,Proline and relative conductivity of‘Miyazaki’and ‘Oregon Spur Delicious’branches in different overwintering periods Different lowercase letters represent the significant differences of‘Miyazaki’at different times;Different capital letters represent the significant differences of‘Oregon Spur Delicious’at different times(P < 0.05).
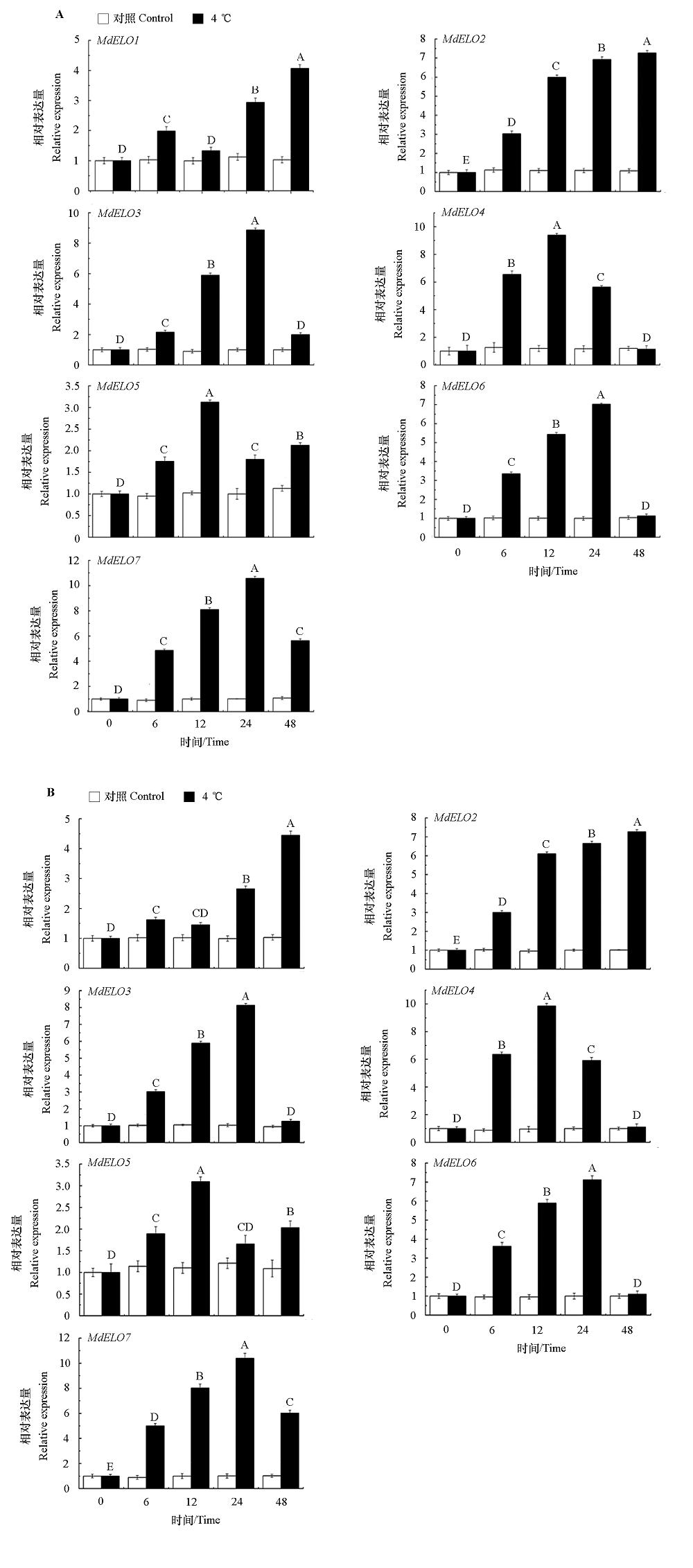
Fig. 9 Expression analysis of apple MhELO gene family in roots(A)and leaves(B)under low temperature stress Different uppercase letters represent significant differences between treatment groups under different root treatment times(P < 0.01).
| [1] |
Achuba F I, Okoh P N. 2015. Effects of petroleum products in soil on α-amylase,starch phosphorylase and peroxidase activities in cowpea and maize seedlings. American Journal of Experimental Agriculture, 6 (6):112-120.
doi: 10.9734/AJEA/2015/9750 URL |
| [2] |
Adamski N M, Bush M S, Simmonds J, Turner A S, Mugford S G, Jones A, Findlay K, Pedentchouk N, Uauy C. 2013. The inhibitor of wax 1 locus(Iw1)prevents formation of β-and OH-β-diketones in wheat cuticular waxes and maps to a sub-cM interval on chromosome arm 2BS. The Plant Journal, 74 (6):989-1002.
doi: 10.1111/tpj.12185 URL |
| [3] |
Belding R D, Blankenship S M, Young E, Leidy R B. 1998. Composition and variability of epicuticular waxes in apple cultivars. Journal of the American Society for Horticultural Science, 123 (3):348-356.
doi: 10.21273/JASHS.123.3.348 URL |
| [4] |
Cahoon E B, Shockey J M, Dietrich C R, Gidda S K, Mullen R T, Dyer J M. 2007. Engineering oilseeds for sustainable production of industrial and nutritional feedstocks:solving bottlenecks in fatty acid flux. Current Opinion in Plant Biology, 10 (3):236-244.
pmid: 17434788 |
| [5] |
Cai Xiao-feng, Zhang Yu-yang. 2013. Genome-wide analysis of plant-specific dof transcription factor family in tomato. Journal of Integrative Plant Biology, 55 (6):552-566. (in Chinese)
doi: 10.1111/jipb.12043 URL |
| 蔡晓峰, 张玉阳. 2013. 番茄植物特异性多夫转录因子家族的全基因组分析. 植物生物学综合杂志, 55 (6):552-566. | |
| [6] |
Cinti D L, Cook L, Nagi M N, Suneja S K. 1992. The fatty acid chain elongation system of mammalian endoplasmic reticulum. Progress in Lipid Research, 31 (1):1-51.
pmid: 1641395 |
| [7] |
Dietbert E, Jorg J, Sauter. 2000. Seasonal changes of activity of a starch granule bound endoamylase and of a starch phosphorylase in poplar wood (Populus × canadensis Moench‘robusta’)and their possible regulation by temperature and phytohormones. Journal of Plant Physiology, 156 (5):731-740.
doi: 10.1016/S0176-1617(00)80239-4 URL |
| [8] |
Eigenbrode S D, Espelie K E. 1995. Effects of plant epicuticular lipids on insect herbivores. Annual Review of Entomology, 40 (1):171-194.
doi: 10.1146/annurev.en.40.010195.001131 URL |
| [9] | Fehling E, Mukherjee K D. 1991. Acyl-CoA elongase from a higher plant(Lunaria annua):metabolic intermediates of very-long chain acyl-CoA products and substrate specificity. Biochimica Biophysica Acta(BBA)-Lipids and Lipid Metabolism, 1082 (3):239-246. |
| [10] |
Jaganathan G K. 2017. Physiological mechanisms only tell half story:multiple biological processes are involved in regulating freezing tolerance of imbibed Lactuca sativa seeds. Scientific Reports, 7 (1):44166.
doi: 10.1038/srep44166 URL |
| [11] |
Jee E R, Peter M P, William W, Van O. 2007. Using the method of homogenization to calculate the effective diffusivity of the stratum corneum. Journal of Membrane Science, 293 (1):174-182.
doi: 10.1016/j.memsci.2007.02.018 URL |
| [12] |
Jin Z, Chandrasekaran U, Liu A. 2014. Genome-wide analysis of the dof transcription factors in castor bean(Ricinus communis L.). Genes and Genomics, 36 (4):527-537.
doi: 10.1007/s13258-014-0189-6 URL |
| [13] | Kazuyoshi T, Kiyohiro N, Ken S. 1978. Difference between the size of waxy and non-waxy kernel in the F1 rice plant.:unbalanced growth in floral glumes and caryopsis in rice. Japanese Journal of Breeding, 28 (3):225-233. |
| [14] |
Kohlwein S D, Eder S, Oh C S, Martin C E, Gable K, Bacikova D, Dunn T. 2001. Tsc13p is required for fatty acid elongation and localizes to a novel structure at the nuclear-vacuolar interface in SacChromyces cerevisiae. Molecular and Cellular Biology, 21 (1):109-125.
pmid: 11113186 |
| [15] | Krah C Y, Kumah P, Mardjan S, Njume A C. 2020. Effect of different threshing methods on the physical chracteristics of two varieties of paddy rice. IOP Conference Series: Earth and Environmental Science, 542 (1): 012022 (9pp). |
| [16] | Lassner M W, Lardizabal K, Metz J G. 1996. A jojoba beta-Ketoacyl-CoA synthase cDNA complements the canola fatty acid elongation mutation in transgenic plants. Th e Plant Cell, 8 (2):281-292. |
| [17] | Lescot M. 2002. Plant CARE,a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Research, 30 (1):325-327. |
| [18] | Makarenko S P, Konenkina T A, Salyaev R K. 2000. Chracteristics of fatty acid composition of lipids in higher plant vacuolar membranes. Membrane & Cell Biology, 13 (5):687-695. |
| [19] |
Millar A A, Clemens S, Zachgo S, Giblin E M, Taylor D C, Kunst L. 1999. CUT1,an arabidopsis gene required for cuticular wax biosynthesis and pollen fertility,encodes a very-long-chain fatty acid condensing enzyme. The Plant Cell, 11 (5):825-838.
doi: 10.1105/tpc.11.5.825 URL |
| [20] |
Millar A A, Kunst L. 1997. Very-long-chain fatty acid biosynthesis is controlled through the expression and specificity of the condensing enzyme. The Plant Journal:for Cell and Molecular Biology, 12 (1):121-31.
doi: 10.1046/j.1365-313X.1997.12010121.x URL |
| [21] |
Nijhawan A, Jain M, Tyagi A K, Khurana J P. 2008. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiology, 146 (2):333-350.
doi: 10.1104/pp.107.112821 pmid: 18065552 |
| [22] | Riederer M, Schneider G. 1990. The effect of the environment on the permeability and composition of citrus leaf cuticles:II. Composition of soluble cuticular lipids and correlation with transport properties. Planta, 180 (2):147-153. |
| [23] |
Shepherd T, Griffiths D W. 2006. The effects of stresson plant cuticular waxes. New Phytol, 171 (3):469-499.
pmid: 16866954 |
| [24] | Sun Bai-lin, Ma Li, Zeng Xiu-cun, Niu Zao-xia, Wang Wan-peng, Hu Fang-di, Lu Xiao-ming, Qi Wei-liang, Wu Jun-yan, Liu Li-jun, Li Xue-cai, Sun Wan-cang, Liao Chong-qing, Wang Xue-fang, Li Xiao-ze, Fang Yan. 2021. Differential expression analysis of fatty acid components and FAD 3 in winter rapeseed under low temperature stress. Agricultural Research in the Arid Areas, 39 (1):65-74. (in Chinese) |
| 孙柏林, 马骊, 曾秀存, 牛早霞, 王万鹏, 呼芳娣, 路晓明, 祁伟亮, 武军艳, 刘丽君, 李学才, 孙万仓, 缪纯庆, 王学芳, 李孝泽, 方彦. 2021. 低温胁迫下白菜型冬油菜脂肪酸组分及FAD3的差异表达分析. 干旱地区农业研究, 39 (1):65-74. | |
| [25] |
Tieman D M, Klee H J. 1999. Differential expression of two novel members of the tomato ethylene-receptor family. Plant Physiology, 120 (1):165-172.
pmid: 10318694 |
| [26] |
Tory A, Hendry, Joseph W, Brown, Geordan T. 2010. Angiosperm phylogeny inferred from sequences of four mitochondrial genes. Journal of Systematics and Evolution, 48 (6):391-425.
doi: 10.1111/j.1759-6831.2010.00097.x URL |
| [27] |
Velasco R, Zharkikh A, Affourtit J, Dhingra A, Cestaro A, Kalyanaraman A, Fontana P, Bhatnagar S K, Troggio M, Pruss D. 2010. The genome of the domesticated apple(Malus domestica Borkh.). Nature Genetics, 42:833-839.
doi: 10.1038/ng.654 URL |
| [28] |
Wang Q, Cheng T R, Yu X N, Jaime A, Teixeira D S, Byrne D H. 2014. Physiological and biochemical responses of six herbaceous peony cultivars to cold stress. South African Journal of Botany, 94:140-148.
doi: 10.1016/j.sajb.2014.05.012 URL |
| [29] | Wilhelm B, Christoph N, David C. 2008. Classification and terminology of plant epicuticular waxes. Botanical Journal of the Linnean Society,(3):237-260. |
| [30] | Xie Feng-lian. 2019. Comparison of expression differences of related genes in brown adipose tissue of yak in different seasons and comparison of fatty acid composition[M. D. Dissertation]. Xining: Qinghai University. (in Chinese) |
| 谢凤莲. 2019. 不同季节牦牛机体棕色脂肪组织相关基因表达差异及其脂肪酸组成成分的比较[硕士论文]. 西宁: 青海大学. | |
| [31] | Xu Guang-ping, Zhang Zhi-yong, Ren Zhong-hong, Zhang Zhi-wei. 2021. Bioinformatics analysis of the structure and function of black sea bream ELO gene-encoded protein. Advances in Biotechnology, 11 (3):344-352. (in Chinese) |
| 许广平, 张志勇, 任忠宏, 张志伟. 2021. 黑鲷ELO基因编码蛋白结构与功能的生物信息学分析. 生物技术进展, 11 (3):344-352. | |
| [32] | Xu J Y, Xue C C, Xue D, Zhao J M, Gai J Y, Guo N, Xing H, Swati P D. 2013a. Overexpression of GmHsp90s,a heat shock protein 90(HSP90)gene family cloning fromsoybean,decrease damage of abiotic stresses in Arabidopsis thaliana. PLoS ONE, 8 (7):e69810. |
| [33] |
Xu Q, Chen L L, Ruan X A, Chen D J, Zhu A D, Chen C L, Denis B, Jiao W B, Hao B H, Matthew P L, Chen J J, Gao S, Xing F, Lan H, Chang J W, Ge X H, Yang L, Hu Q, Miao Y, Wang L, Xiao S X, Manosh K B, Zeng W F, Guo F, Cao H B, Yang X M, Xu X W, Cheng Y J, Xu J, Liu J H, Oscar J L, Tang Z H, Guo W W, Kuang H H, Zhang H Y, Mikeal L R, Niranjan N, Deng X X, Ruan Y J. 2013b. The draft genome of sweet orange(Citrus sinensis). Nature genetics, 45 (1):59-66.
doi: 10.1038/ng.2472 URL |
| [34] | Yang Jie, Jiang Li, Zhang Jioajiao, Chen Ziping. 2013. Responses of the drought-resistent vem 1 mutant to NaCl and ABA stress in Arabidopsis. Plant Physiology Journal, 49 (4):337-342. (in Chinese) |
| 杨杰, 江力, 张娇娇, 陈子平. 2013. 拟南芥抗旱突变体vem1对NaCl和ABA胁迫的响应. 植物生理学报, 49 (4):337-342. | |
| [35] | Yang Ning-ning, Sun Wan-cang, Liu Zi-gang, Shi Peng-hui, Fang Yan, Wu Jun-yan, Zeng Xiu-cun, Kong De-jing, Lu Mei-hong, Wang Yue. 2014. Morphological and physiological mechanism of cold resistance in winter rape in northern China. Chinese Agricultural Sciences, 47 (3):452-461. (in Chinese) |
| 杨宁宁, 孙万仓, 刘自刚, 史鹏辉, 方彦, 武军艳, 曾秀存, 孔德晶, 鲁美宏, 王月. 2014. 北方冬油菜抗寒性的形态与生理机制. 中国农业科学, 47 (3):452-461. | |
| [36] | Yao Q M, Bai Y L, Kumar S, Au E, Orfao A, Chim C S. 2021. Minimal residual disease detection by next-generation sequencing in multiple myeloma:a comparison with real-time quantitative PCR. Frontiers in Oncology, 10:611021. |
| [37] | Yoshida K, Komae K. 2006. A rice family 9 glycoside hydrolase isozyme with broad substrate specificity for hemicelluloses in type II cell walls. Plant & Cell Physiology, 47 (11):1541-1554. |
| [38] |
Zhang K, Marina K, Han M, Li W, Yu Z Y, Yang Z L, Li Y, Michael L, Metzker, Rando A, Donald J Z, Laura E K, Pamela S L, Paul W W, Ian M M, Paul A S, David J F, Christopher P A, Robert J G, Radha A, Konstantin P. 2001. A 5-bp deletion in ELOVL 4 is associated with two related forms of autosomal dominant macular dystrophy. Nat Genet, 27 (1):89-93.
pmid: 11138005 |
| [39] | Zhou Jing-wen, Ding Yu-jiao, Zeng Xiao-rong, Ganesh K Jaganathan. 2020. Study on the relationship between freeze tolerance and fatty acid synthesis and metabolism of lettuce seeds after low temperature treatment. Subtropical Plant Science, 49 (1):9-14. (in Chinese) |
| 周婧雯, 丁玉娇, 曾晓蓉, Ganesh K Jaganathan. 2020. 生菜种子低温处理后耐冻性和脂肪酸合成代谢的关系研究. 亚热带植物科学, 49 (1):9-14. |
| [1] | YU Tingting, LI Huan, NING Yuansheng, SONG Jianfei, PENG Lulin, JIA Junqi, ZHANG Weiwei, and YANG Hongqiang. Genome-wide Identification of GRAS Gene Family in Apple and Expression Analysis of Its Response to Auxin [J]. Acta Horticulturae Sinica, 2023, 50(2): 397-409. |
| [2] | ZHAO Xueyan, WANG Qi, WANG Li, WANG Fangyuan, WANG Qing, LI Yan. Comparative Transcriptome Analysis of Differential Expression in Different Tissues of Corydalis yanhusuo [J]. Acta Horticulturae Sinica, 2023, 50(1): 177-187. |
| [3] | HAN Xiaolei, ZHANG Caixia, LIU Kai, YANG An, YAN Jiadi, LI Wuxing, KANG Liqun, and CONG Peihua. A New Mid-ripening Apple Cultivar‘Zhongping Youlei’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 1-2. |
| [4] | YANG Xingwang, WANG Haibo, WANG Yingying, WANG Xiaolong, WANG Zhiqiang, LIU Peipei, LIU Wanchun, and WANG Xiaodi. A New Mid-ripening and Cold Resistant Peach Cultivar‘Zhongnong Ganshuang’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 15-16. |
| [5] | YANG Xingwang, WANG Haibo, WANG Yingying, ZHANG Yican, WANG Baoliang, Liu Peipei, SHI Xiangbin, LIU Wanchun, and WANG Xiaodi. A New Mid-ripening Cold Resistant Peach Cultivar‘Zhongnong Baigan’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 17-18. |
| [6] | YANG Xingwang, LIU Fengzhi, WANG Haibo, WANG Yingying, WANG Zhiqiang, SHI Xiangbin, JI Xiaohao, LIU Wanchun, and WANG Xiaodi. A New Mid-ripening Cold Resistant Peach Cultivar‘Zhongnong Hanshuimi’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 19-20. |
| [7] | YANG Xingwang, LIU Fengzhi, WANG Haibo, WANG Yingying, ZHANG Yican, LI Peng, WANG Xiaolong, LIU Wanchun, and WANG Xiaodi. A New Late-ripening Cold Resistant Peach Cultivar‘Zhongnong Qiuxiang’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 21-22. |
| [8] | SUN Simiao, WANG Kun, GAO Yuan, WANG Dajiang, and LI Lianwen. A New Ornamental Crabapple Cultivar‘Zichen’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 267-268. |
| [9] | HAN Xiaolei, ZHANG Caixia, LIU Kai, YAN Jiadi, LI Wuxing, KANG Liqun, and CONG Peihua. A New Mid-ripening Apple Cultivar‘Pingyou 2’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 1-2. |
| [10] | WANG Qiang, CONG Peihua, and LIU Xiaofeng. A New Late Ripening Apple Cultivar‘Huayou Tianwa’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 3-4. |
| [11] | WANG Qiang, CONG Peihua, and LIU Xiaofeng. A New Mid-ripening Apple Cultivar‘Huayou Baomi’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 5-6. |
| [12] | YANG Ling, CONG Peihua, WANG Qiang, LI Wuxing, and KANG Liqun. A New Mid-ripening Apple Cultivar‘Huafeng’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 7-8. |
| [13] | WANG Yingying, ZHENG Xiaocui, WANG Haibo, SHI Xiangbin, JI Xiaohao, LIU Peipei, LIU Wanchun, and WANG Xiaodi. A New Cold Resistent Peach Cultivar‘Zhongtao Fenyu’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 13-14. |
| [14] | WANG Yingying, WANG Haibo, LIU Peipei, WANG Baoliang, ZHANG Yican, LI Peng, LIU Wanchun, and WANG Xiaodi. A New Cold Resistant Peach Cultivar‘Zhongnong Xiutian’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 15-16. |
| [15] | WANG Yingying, WANG Haibo, LI Peng, WANG Baoliang, SHI Xiangbin, LIU Wanchun, and WANG Xiaodi. A New Cold Resistant Peach Cultivar‘Zhongnong Zhihou’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 17-18. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd