
园艺学报 ›› 2021, Vol. 48 ›› Issue (2): 205-218.doi: 10.16420/j.issn.0513-353x.2020-0272
• 研究论文 • 下一篇
杨亚明, 丁毓端, 陈丽娟, 田雪婷, 殷伟杰, 彭红慧, 杜薇, 梁丽平, 任小林*( )
)
收稿日期:2020-09-07
修回日期:2020-11-11
出版日期:2021-02-25
发布日期:2021-03-09
通讯作者:
任小林
E-mail:renxl@nwsuaf.edu.cn
基金资助:
YANG Yaming, DING Yuduan, CHEN Lijuan, TIAN Xueting, YIN Weijie, PENG Honghui, DU Wei, LIANG Liping, REN Xiaolin*( )
)
Received:2020-09-07
Revised:2020-11-11
Online:2021-02-25
Published:2021-03-09
Contact:
REN Xiaolin
E-mail:renxl@nwsuaf.edu.cn
摘要:
液泡铁离子转运蛋白(vacuolar iron transporter,VIT)参与铁的储存和运输,对植物光合作用、固氮、呼吸、DNA和激素合成有重要作用。本研究中鉴定了苹果全基因组的9个VIT基因,通过系统发育分析其进化关系,表明苹果VIT蛋白与拟南芥进化关系较近。染色体定位显示9个VIT基因分布在7条染色体上,VIT基因的启动子区域富含光反应、植物激素调控和启动子相关顺式作用元件。表达谱分析显示VIT基因在花和果实中的表达量相对较高。此外,对MdVIT家族进行蛋白理化性质、String互作网络和基因结构等分析,表明苹果VIT家族扩增的主要因素是串联重复和片段复制,与苹果VIT蛋白互作的主要蛋白是阳离子共转运蛋白(XP_008372221.1和XP_008383418.1)。qRT-PCR分析结果表明:在2 000 μmol · L-1 FeSO4 · 7H2O(高铁)处理的‘M26’苹果砧木苗中,苹果VIT家族基因在不同组织的表达中存在差异,6 h后,MdVIT4在茎中的相对表达量最高,是对照的237倍。24 h后,MdVIT1和MdVIT2在叶片中的相对表达量与对照相比呈现上调表达,分别是对照的6.6倍和12.0倍。综上,基因MdVIT1、MdVIT2和MdVIT4可能与植物耐铁胁迫有密切关系,为苹果逆境胁迫机制研究提供参考。
中图分类号:
杨亚明, 丁毓端, 陈丽娟, 田雪婷, 殷伟杰, 彭红慧, 杜薇, 梁丽平, 任小林. 苹果液泡铁离子转运蛋白基因家族的鉴定与表达分析[J]. 园艺学报, 2021, 48(2): 205-218.
YANG Yaming, DING Yuduan, CHEN Lijuan, TIAN Xueting, YIN Weijie, PENG Honghui, DU Wei, LIANG Liping, REN Xiaolin. Identification and Expression Analysis of the Vacuolar Iron Transporter Gene Family in Apple[J]. Acta Horticulturae Sinica, 2021, 48(2): 205-218.
| 用途Usage | 基因Gene | 上游引物(5′-3′)Forward primer | 下游引物(5′-3′)Reverse primer |
|---|---|---|---|
| qRT-PCR | MdVIT1 | CTCCAATCCCCGAAGACGA | GCGATGCAACGGAAACCA |
| MdVIT2 | TCCGCTTCTCCCGTTAATATT | CGAGGTCGTTTGTGGTCATT | |
| MdVIT3 | CACCAAACAACGCTAGACCAA | TCACAGCTCCGACTCCCAT | |
| MdVIT4 | TTCGATTACTCCCAGAGGACAC | AGCGATGCAACAGAAACCAG | |
| MdVIT5 | AATCCCAGAAGGCGACGAA | GAGCGATGCAACAGAAACCAGCCCA | |
| MdVIT6 | TCTTACCGTGCCGTTTGCT | ATATCCGCCGAGTCCCATG | |
| MdVIT7 | CAAGGACCGAGCAAGTGCA | CATCAAGGACGTGGTTGACAGT | |
| MdVIT8 | CGACGGTCTCACTGTCCCTT | GGCGGCAACTTCAGCAAT | |
| MdVIT9 | ACCGCGAGAAGCAAGCACT | AGCGAACGGCACGGTAAGA | |
| 克隆基因及 启动子 Clone primers | MD04G1208900 | ATGGCTTCCCAACGTGACCA | TCAGATTCCCAAACCACTAGACC |
| MD05G1023100 | ATGTGTCCTCCACACTTCTCCAA | TCATAGCCCACTTGAGCCAATC | |
| MD12G1223300 | ATGGCTTCCCAACGTGACCA | TCACACCTGCATACCACTAGAC |
表1 苹果VIT基因家族表达分析的实时荧光定量和克隆引物
Table 1 qRT-PCR and clone primers for expression on analysis of VIT in apple
| 用途Usage | 基因Gene | 上游引物(5′-3′)Forward primer | 下游引物(5′-3′)Reverse primer |
|---|---|---|---|
| qRT-PCR | MdVIT1 | CTCCAATCCCCGAAGACGA | GCGATGCAACGGAAACCA |
| MdVIT2 | TCCGCTTCTCCCGTTAATATT | CGAGGTCGTTTGTGGTCATT | |
| MdVIT3 | CACCAAACAACGCTAGACCAA | TCACAGCTCCGACTCCCAT | |
| MdVIT4 | TTCGATTACTCCCAGAGGACAC | AGCGATGCAACAGAAACCAG | |
| MdVIT5 | AATCCCAGAAGGCGACGAA | GAGCGATGCAACAGAAACCAGCCCA | |
| MdVIT6 | TCTTACCGTGCCGTTTGCT | ATATCCGCCGAGTCCCATG | |
| MdVIT7 | CAAGGACCGAGCAAGTGCA | CATCAAGGACGTGGTTGACAGT | |
| MdVIT8 | CGACGGTCTCACTGTCCCTT | GGCGGCAACTTCAGCAAT | |
| MdVIT9 | ACCGCGAGAAGCAAGCACT | AGCGAACGGCACGGTAAGA | |
| 克隆基因及 启动子 Clone primers | MD04G1208900 | ATGGCTTCCCAACGTGACCA | TCAGATTCCCAAACCACTAGACC |
| MD05G1023100 | ATGTGTCCTCCACACTTCTCCAA | TCATAGCCCACTTGAGCCAATC | |
| MD12G1223300 | ATGGCTTCCCAACGTGACCA | TCACACCTGCATACCACTAGAC |
| 基因名 Gene name | 登录号 Gene ID | 序列位置 Sequence position | 染色体 位置 Chr | 开放阅 读区/bp ORF | 氨基酸 数/aa Length of amino acid | 分子量/ kD Molecular weigh | 不稳定 系数 Instability index | 理论等 电点 pI | 蛋白质 疏水性 Hydroph- obicity | 脂肪指数 Aliphatic index | 跨膜结 构域 TMD |
|---|---|---|---|---|---|---|---|---|---|---|---|
| MdVIT1 | MD04G1208900 | 29415642..29416325 | Chr04 | 684 | 227 | 23.71 | 42.66 | 6.13 | 0.29 | 97.14 | 4 |
| MdVIT2 | MD05G1023100 | 3872852..3873643 | Chr05 | 792 | 263 | 27.68 | 41.63 | 6.96 | 0.25 | 97.91 | 5 |
| MdVIT3 | MD10G1024000 | 3007772..3008458 | Chr10 | 687 | 228 | 24.01 | 34.11 | 6.13 | 0.28 | 101.05 | 4 |
| MdVIT4 | MD12G1223300 | 30004855..30005538 | Chr12 | 684 | 227 | 23.87 | 43.44 | 7.78 | 0.30 | 96.26 | 4 |
| MdVIT5 | MD12G1223400 | 30015123..30015806 | Chr12 | 684 | 227 | 23.90 | 45.02 | 6.30 | 0.28 | 96.26 | 4 |
| MdVIT6 | MD13G1018700 | 1174829..1178506 | Chr13 | 744 | 247 | 26.36 | 36.99 | 5.18 | 0.19 | 101.30 | 4 |
| MdVIT7 | MD15G1227000 | 18449646..18451051 | Chr15 | 798 | 265 | 27.99 | 42.77 | 8.34 | 0.01 | 89.51 | 5 |
| MdVIT8 | MD16G1016500 | 1240143..1242202 | Chr16 | 747 | 248 | 26.52 | 36.31 | 5.00 | 0.38 | 105.16 | 4 |
| MdVIT9 | MD16G1016600 | 1243044..1245822 | Chr16 | 744 | 247 | 26.41 | 35.57 | 5.89 | 0.21 | 105.63 | 4 |
表2 苹果中VIT基因的基本信息
Table 2 Detailed information of identified VIT genes in apple genome
| 基因名 Gene name | 登录号 Gene ID | 序列位置 Sequence position | 染色体 位置 Chr | 开放阅 读区/bp ORF | 氨基酸 数/aa Length of amino acid | 分子量/ kD Molecular weigh | 不稳定 系数 Instability index | 理论等 电点 pI | 蛋白质 疏水性 Hydroph- obicity | 脂肪指数 Aliphatic index | 跨膜结 构域 TMD |
|---|---|---|---|---|---|---|---|---|---|---|---|
| MdVIT1 | MD04G1208900 | 29415642..29416325 | Chr04 | 684 | 227 | 23.71 | 42.66 | 6.13 | 0.29 | 97.14 | 4 |
| MdVIT2 | MD05G1023100 | 3872852..3873643 | Chr05 | 792 | 263 | 27.68 | 41.63 | 6.96 | 0.25 | 97.91 | 5 |
| MdVIT3 | MD10G1024000 | 3007772..3008458 | Chr10 | 687 | 228 | 24.01 | 34.11 | 6.13 | 0.28 | 101.05 | 4 |
| MdVIT4 | MD12G1223300 | 30004855..30005538 | Chr12 | 684 | 227 | 23.87 | 43.44 | 7.78 | 0.30 | 96.26 | 4 |
| MdVIT5 | MD12G1223400 | 30015123..30015806 | Chr12 | 684 | 227 | 23.90 | 45.02 | 6.30 | 0.28 | 96.26 | 4 |
| MdVIT6 | MD13G1018700 | 1174829..1178506 | Chr13 | 744 | 247 | 26.36 | 36.99 | 5.18 | 0.19 | 101.30 | 4 |
| MdVIT7 | MD15G1227000 | 18449646..18451051 | Chr15 | 798 | 265 | 27.99 | 42.77 | 8.34 | 0.01 | 89.51 | 5 |
| MdVIT8 | MD16G1016500 | 1240143..1242202 | Chr16 | 747 | 248 | 26.52 | 36.31 | 5.00 | 0.38 | 105.16 | 4 |
| MdVIT9 | MD16G1016600 | 1243044..1245822 | Chr16 | 744 | 247 | 26.41 | 35.57 | 5.89 | 0.21 | 105.63 | 4 |
| 蛋白 | 液泡 | 质膜 | 内质网 | 高尔基体 | 胞外 |
|---|---|---|---|---|---|
| Protein | Vacuolar | Plasma membrane | Endoplasmic reticulum | Golgi apparatus | Extracellular |
| MdVIT1 | 9 | 4 | 1 | ||
| MdVIT2 | 2 | 7 | 1 | ||
| MdVIT3 | 12 | 2 | |||
| MdVIT4 | 4 | 5.5 | 1 | 2 | |
| MdVIT5 | 7 | 5 | 1 | 1 | |
| MdVIT6 | 1 | 12 | |||
| MdVIT7 | 2 | 5 | 6 | 1 | |
| MdVIT8 | 2 | 8 | 3 | 1 | |
| MdVIT9 | 2 | 12 |
表3 苹果MdVIT亚细胞定位预测
Table 3 Subcellular predication of apple MdVIT
| 蛋白 | 液泡 | 质膜 | 内质网 | 高尔基体 | 胞外 |
|---|---|---|---|---|---|
| Protein | Vacuolar | Plasma membrane | Endoplasmic reticulum | Golgi apparatus | Extracellular |
| MdVIT1 | 9 | 4 | 1 | ||
| MdVIT2 | 2 | 7 | 1 | ||
| MdVIT3 | 12 | 2 | |||
| MdVIT4 | 4 | 5.5 | 1 | 2 | |
| MdVIT5 | 7 | 5 | 1 | 1 | |
| MdVIT6 | 1 | 12 | |||
| MdVIT7 | 2 | 5 | 6 | 1 | |
| MdVIT8 | 2 | 8 | 3 | 1 | |
| MdVIT9 | 2 | 12 |
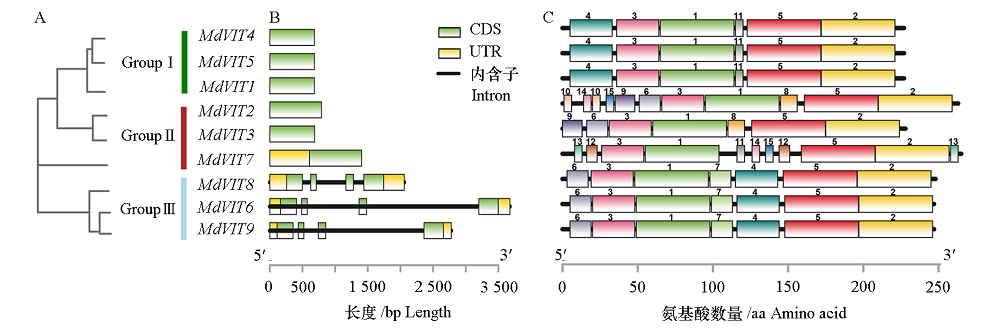
图1 MdVIT家族系统发育树(A)基因的结构分析(B)和MdVIT蛋白的保守基序(C)
Fig. 1 Phylogenetic classification of MdVIT(A),gene structure analyses of MdVIT(B)and conserved motif of MdVIT proteins(C)
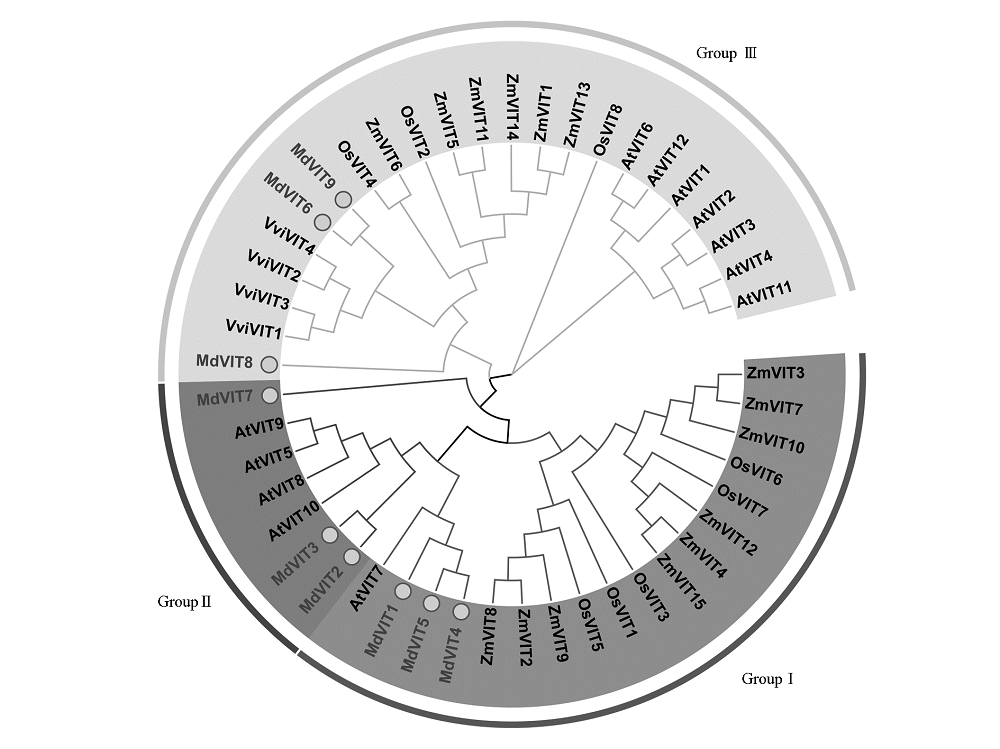
图2 拟南芥(At)、水稻(Os)、玉米(Zm)、葡萄(Vv)和苹果(Md)VIT 基因家族发育树
Fig. 2 The phylogenetic tree of VIT gene family in Arabidopsis thaliana(At),Oryza sativa(Os),Zea mays(Zm), Vitis vinifera(Vv)and Malus × domestica(Md)
| 基因 Gene | 基因 Gene | 同义 替换率 Ka | 非同义 替换率 Ks | 选择性 压力 Ka/Ks | 分离时间/Mya Divergence Time | 复制类型 Duplication Types |
|---|---|---|---|---|---|---|
| MdVIT1(MD04G1208900) | MdVIT5(MD12G1223400) | 0.055 | 0.226 | 0.245 | 7.527 | 片段复制 Segmental duplication |
| MdVIT1(MD04G1208900) | MdVIT4(MD12G1223300) | 0.071 | 0.190 | 0.371 | 6.347 | 片段复制 Segmental duplication |
| MdVIT2(MD05G1023100) | MdVIT3(MD10G1024000) | 1.030 | 0.912 | 1.130 | 30.387 | 串联重复 Tandem dupication |
| MdVIT4(MD12G1223300) | MdVIT5(MD12G1223400) | 0.036 | 0.103 | 0.349 | 3.425 | 片段复制 Segmental duplication |
表4 苹果VIT串联重复基因的分离时间
Table 4 Divergence time of the VIT paralogues in apple
| 基因 Gene | 基因 Gene | 同义 替换率 Ka | 非同义 替换率 Ks | 选择性 压力 Ka/Ks | 分离时间/Mya Divergence Time | 复制类型 Duplication Types |
|---|---|---|---|---|---|---|
| MdVIT1(MD04G1208900) | MdVIT5(MD12G1223400) | 0.055 | 0.226 | 0.245 | 7.527 | 片段复制 Segmental duplication |
| MdVIT1(MD04G1208900) | MdVIT4(MD12G1223300) | 0.071 | 0.190 | 0.371 | 6.347 | 片段复制 Segmental duplication |
| MdVIT2(MD05G1023100) | MdVIT3(MD10G1024000) | 1.030 | 0.912 | 1.130 | 30.387 | 串联重复 Tandem dupication |
| MdVIT4(MD12G1223300) | MdVIT5(MD12G1223400) | 0.036 | 0.103 | 0.349 | 3.425 | 片段复制 Segmental duplication |
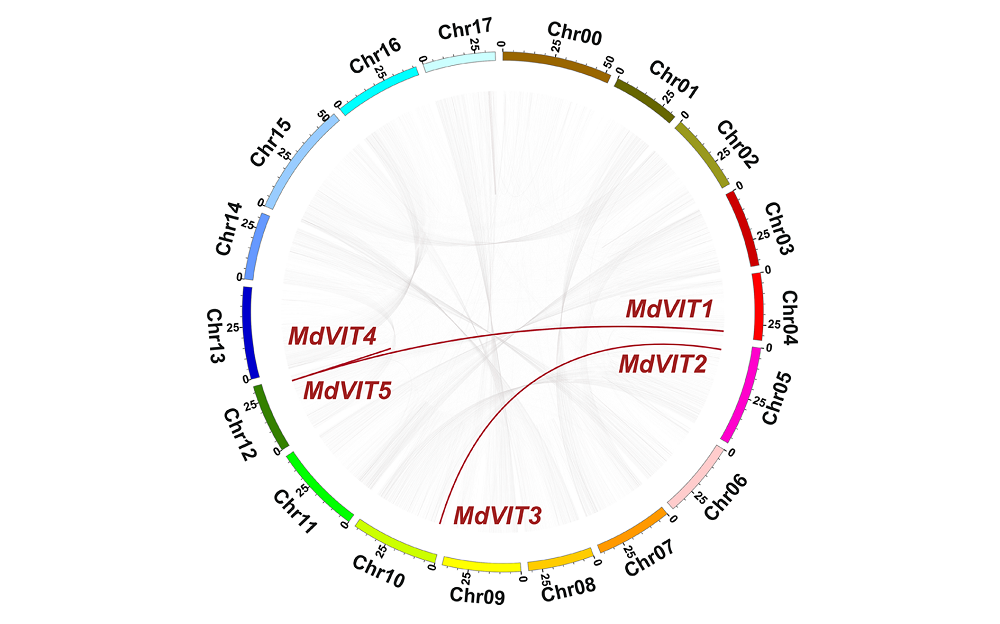
图4 苹果MdVIT复制基因的染色体定位和共线性分析 红色线条表示1对复制基因。
Fig. 4 Chromosome location and collinear distribution of MdVIT genes in apple genome Red lines showed gene duplication.
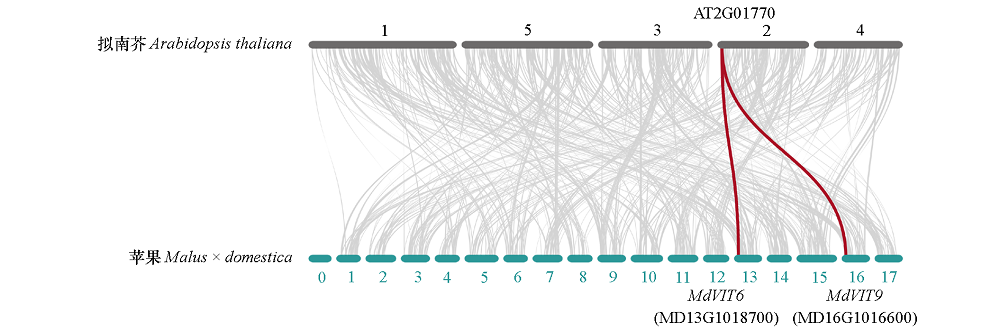
图5 苹果与拟南芥的VIT基因的同源性分析 灰色线条代表苹果和拟南芥基因组中的共线性区域,红色线条突出显示了共线性VIT基因对。
Fig. 5 Synteny analysis of VIT genes between apple and Arabidopsis thaliana Gray lines in the background indicate the collinear blocks with apple and Arabidopsis thaliana,while the red lines highlight the syntenic VIT gene pairs.

图7 MdVIT在不同苹果种质的不同组织中的表达模式 GD表示‘金冠’;M69、M74、M20、X8877、M14、M49、X4102、X4442 × X2596和X3069 × X992为不同苹果杂交种。
Fig. 7 Expression profiles of MdVIT genes in different tissues and different apple varieties GD reference‘Golden Delicious’;M69,M74,M20,X8877,M14,M49,X4102,X4442 × X2596,and X3069 × X992 were various apple hybrids.
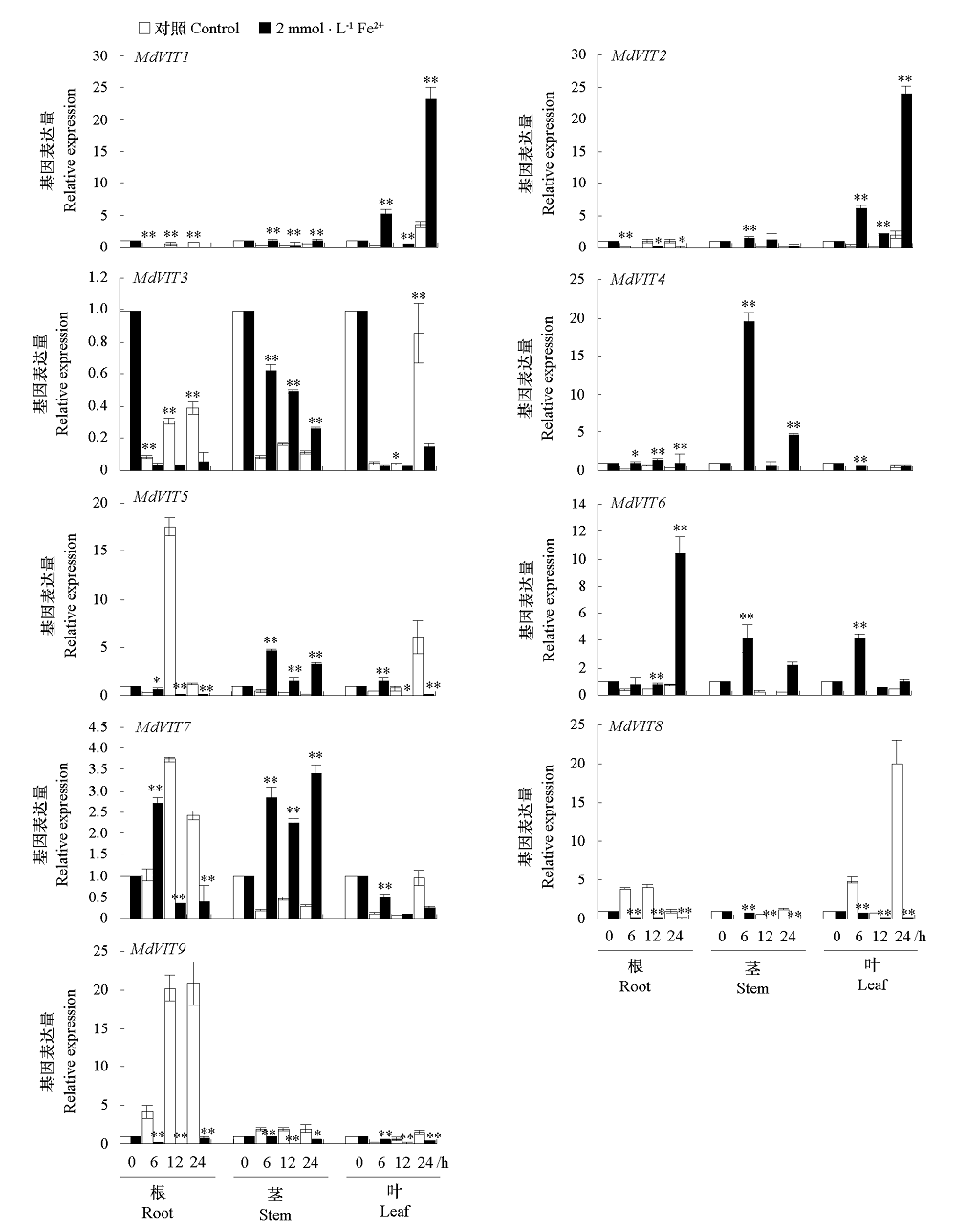
图8 MdVIT在苹果不同组织中的表达分析 处理与对照的差异显著性分析用t-test法统计。* α = 0.05,** α = 0.01.
Fig. 8 Expression profile of MdVIT in different tissues in apple Significance difference between treatment and the control were analyzed by t-test. * α = 0.05,** α = 0.01.
| [1] |
Artimo P, Jonnalagedda M, Arnold K, Baratin D, Csardi G, de Castro E, Duvaud S, Flegel V, Fortier A, Gasteiger E. 2012. ExPASy:SIB bioinformatics resource portal. Nucleic Acids Research, 40(W1):W597-W603.
doi: 10.1093/nar/gks400 URL |
| [2] |
Bailey T L, Boden M, Buske F A, Frith M, Grant C E, Clementi L, Ren J Y, Li W W, Noble W S. 2009. MEME SUITE:tools for motif discovery and searching. Nucleic Acids Research, 37:W202-W208.
doi: 10.1093/nar/gkp335 URL |
| [3] |
Blanc G, Wolfe K H. 2004. Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. The Plant Cell, 16(7):1667-1678.
doi: 10.1105/tpc.021345 URL |
| [4] |
Cao J, Li X. 2015. Identification and phylogenetic analysis of late embryogenesis abundant proteins family in tomato(Solanum lycopersicum). Planta, 241(3):757-772.
doi: 10.1007/s00425-014-2215-y URL |
| [5] |
Cao J. 2019. Molecular evolution of the vacuolar iron transporter(VIT)family genes in 14 plant species. Genes, 10(2):16.
doi: 10.3390/genes10010016 URL |
| [6] |
Colmenero-Flores J M, Martinez G, Gamba G, Vazquez N, Iglesias D J, Brumos J, Talon M. 2007. Identification and functional characterization of cation-chloride cotransporters in plants. Plant Journal, 50(2):278-292.
pmid: 17355435 |
| [7] | Connolly E L, Guerinot M L. 2002. Iron stress in plants. Genome Biology, 3(8):1-4. |
| [8] |
Connorton J M, Jones E R, Rodriguez-Ramiro I, Fairweather-Tait S, Uauy C, Balk J. 2017. Wheat vacuolar iron transporter Ta VIT2 transports Fe and Mn and is effective for biofortification. Plant Physiology, 174(4):2434-2444.
doi: 10.1104/pp.17.00672 pmid: 28684433 |
| [9] |
Crombach A, Hogeweg P. 2007. Chromosome rearrangements and the evolution of genome structuring and adaptability. Molecular Biology and Evolution, 24(5):1130-1139.
doi: 10.1093/molbev/msm033 URL |
| [10] | Grotz N, Guerinot M L. 2006. Molecular aspects of Cu,Fe and Zn homeostasis in plants. Biochimica Et Biophysica Acta- Molecular Cell Research, 1763(7):595-608. |
| [11] |
Gu Z L, Cavalcanti A, Chen F C, Bouman P, Li W H. 2002. Extent of gene duplication in the genomes of Drosophila,nematode,and yeast. Molecular Biology and Evolution, 19(3):256-262.
doi: 10.1093/oxfordjournals.molbev.a004079 URL |
| [12] |
Hell R, Stephan U W. 2003. Iron uptake,trafficking and homeostasis in plants. Planta, 216(4):541-551.
doi: 10.1007/s00425-002-0920-4 URL |
| [13] |
Hu B, Jin J P, Guo A Y, Zhang H, Luo J C, Gao G. 2015. GSDS 2.0:an upgraded gene feature visualization server. Bioinformatics, 31(8):1296-1297.
doi: 10.1093/bioinformatics/btu817 URL |
| [14] |
Imsande J. 1998. Iron,sulfur,and chlorophyll deficiencies:a need for an integrative approach in plant physiology. Physiologia Plantarum, 103(1):139-144.
doi: 10.1034/j.1399-3054.1998.1030117.x URL |
| [15] |
Kim S A, Punshon T, Lanzirotti A, Li L T, Alonso J M, Ecker J R, Kaplan J, Guerinot M L. 2006. Localization of iron in Arabidopsis seed requires the vacuolar membrane transporter VIT1. Science, 314(5803):1295-1298.
doi: 10.1126/science.1132563 URL |
| [16] |
Kong X Q, Gao X H, Sun W, An J, Zhao Y X, Zhang H. 2011. Cloning and functional characterization of a cation-chloride cotransporter gene OsCCC1. Plant Molecular Biology, 75(6):567-578.
doi: 10.1007/s11103-011-9744-6 URL |
| [17] |
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones S J, Marra M A. 2009. Circos: An information aesthetic for comparative genomics. Genome Res, 19(9):1639-1645.
doi: 10.1101/gr.092759.109 URL |
| [18] | Latrasse D, Rodriguez-Granados N Y, Veluchamy A, Mariappan K G, Bevilacqua C, Crapart N, Camps C, Sommard V, Raynaud C, Dogimont C. 2017. The quest for epigenetic regulation underlying unisexual flower development in Cucumis melo. Epigenetics & Chromatin, 10. |
| [19] |
Le Hir H, Nott A, Moore M J. 2003. How introns influence and enhance eukaryotic gene expression. Trends Biochem Sci, 28(4):215-220.
doi: 10.1016/S0968-0004(03)00052-5 URL |
| [20] |
Lee T H, Tang H B, Wang X Y, Paterson A H. 2013. PGDD:a database of gene and genome duplication in plants. Nucleic Acids Research, 41(D1):D1152-D1158.
doi: 10.1093/nar/gks1104 URL |
| [21] |
Li L T, Chen O S, Ward DM, Kaplan J. 2001. CCC1 is a transporter that mediates vacuolar iron storage in yeast. J Biol Chem, 276(31):29515-29519.
pmid: 11390404 |
| [22] | Li S, Wang N, Ji D D, Xue Z Y, Yu Y C, Jiang Y P, Liu J L, Liu Z H, Xiang F N. 2016. Evolutionary and functional analysis of membrane-bound NAC transcription factor genes in soybean. Plant Physiology, 172(3):1804-1820. |
| [23] |
Livak K J, Schmittgen T D. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2-∆∆CT method. Methods, 25(4):402-408.
pmid: 11846609 |
| [24] |
Mattick J S, Gagen M J. 2001. The evolution of controlled multitasked gene networks: The role of introns and other noncoding RNAs in the development of complex organisms. Molecular Biology and Evolution, 18(9):1611-1630.
doi: 10.1093/oxfordjournals.molbev.a003951 URL |
| [25] |
Mistry J, Finn R D, Eddy S R, Bateman A, Punta M. 2013. Challenges in homology search:HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Research, 41(12):e121-e121.
doi: 10.1093/nar/gkt263 URL |
| [26] |
Moes A D, van der Lubbe N, Zietse R, Loffing J, Hoorn E J. 2014. The sodium chloride cotransporter SLC12A3: new roles in sodium,potassium,and blood pressure regulation. Pflugers Archiv-European Journal of Physiology, 466(1):107-118.
doi: 10.1007/s00424-013-1407-9 URL |
| [27] |
Nagasaka S, Takahashi M, Nakanishi-Itai R, Bashir K, Nakanishi H, Mori S, Nishizawa N K. 2009. Time course analysis of gene expression over 24 h in Fe-deficient barley roots. Plant Mol Biol, 69(5):621-631.
doi: 10.1007/s11103-008-9443-0 pmid: 19089316 |
| [28] | Saitou N, Nei M. 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology and Evolution, 4(4):406-425. |
| [29] |
Schmidt W. 2003. Iron solutions: acquisition strategies and signaling pathways in plants. Trends in Plant Science, 8(4):188-193.
doi: 10.1016/S1360-1385(03)00048-7 URL |
| [30] |
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. 2011. MEGA5:molecular evolutionary genetics analysis using maximum likelihood,evolutionary distance,and maximum parsimony methods. Molecular Biology and Evolution, 28(10):2731-2739.
doi: 10.1093/molbev/msr121 URL |
| [31] | Wang Ruonan, Nie Lanchun, Zhang Shuangshuang, Cui Qiang, Jia Mingfei. 2019. Research progress on plant resistance to heavy metal stress. Acta Horticulturae Sinica, 46(1):157-170. (in Chinese) |
| 王若男, 乜兰春, 张双双, 崔强, 贾明飞. 2019. 植物抗重金属胁迫研究进展. 园艺学报, 46(1):157-170. | |
| [32] | Wang Y P, Tang H B, DeBarry J D, Tan X, Li J P, Wang X Y, Lee T H, Jin H Z, Marler B, Guo H. 2012. MCScanX:a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Research, 40(7):14. |
| [33] | Xu Rui-rui, Li Rui, Wang Xiao-fei, Hao Yu-jin, 2018. Identification and expression analysis under abiotic stresses of OFP gene family in apple. Scientia Agricultura Sinica, 51(10):141-152. (in Chinese) |
| 许瑞瑞, 李睿, 王小非, 郝玉金. 2018. 苹果OFP基因家族的全基因组鉴定与非生物逆境表达分析. 中国农业科学, 51(10):141-152. | |
| [34] |
Yang ZF, Zhou Y, Wang X F, Gu S L, Yu J M, Liang G H, Yan C J, Xu C W. 2008. Genomewide comparative phylogenetic and molecular evolutionary analysis of tubby-like protein family in Arabidopsis,rice,and poplar. Genomics, 92(4):246-253.
doi: 10.1016/j.ygeno.2008.06.001 URL |
| [35] |
Zhang Y, Xu Y H, Yi H Y, Gong J M. 2012. Vacuolar membrane transporters OsVIT1 and OsVIT2 modulate iron translocation between flag leaves and seeds in rice. Plant Journal, 72(3):400-410.
doi: 10.1111/tpj.2012.72.issue-3 URL |
| [36] | Zhou Rong-fang, Yuan Wu-zhou, Tong Chao-bo, Huang Jun-yan, Cheng Xiao-Hui, Yu Jing-yin, Dong Cai-hua, Liu Yue-ying, Liu Sheng-yi. 2014. Identification and evolution of VIT gene family between A and C genomes in Brassica. Chinese Journal of Oil Crop Sciences, 36(5):551-561. (in Chinese) |
| 周荣芳, 袁午舟, 童超波, 黄军艳, 程晓晖, 于景印, 董彩华, 刘越英, 刘胜毅. 2014. 芸薹属AC基因组中VIT基因家族的鉴定与进化. 中国油料作物学报, 36(5):551-561. |
| [1] | 王晓晨, 聂子页, 刘先菊, 段 伟, 范培格, 梁振昌, . 脱落酸对‘京香玉’葡萄果实单萜物质合成的影响[J]. 园艺学报, 2023, 50(2): 237-249. |
| [2] | 于婷婷, 李 欢, 宁源生, 宋建飞, 彭璐琳, 贾竣淇, 张玮玮, 杨洪强. 苹果GRAS全基因组鉴定及其对生长素的响应分析[J]. 园艺学报, 2023, 50(2): 397-409. |
| [3] | 翟含含, 翟宇杰, 田义, 张叶, 杨丽, 温陟良, 陈海江. 桃SAUR家族基因分析及PpSAUR5功能鉴定[J]. 园艺学报, 2023, 50(1): 1-14. |
| [4] | 崔建, 钟雄辉, 刘泽慈, 陈登辉, 李海龙, 韩睿, 乐祥庆, 康俊根, 王超. 结球甘蓝染色体片段替换系构建[J]. 园艺学报, 2023, 50(1): 65-78. |
| [5] | 罗海林, 袁雷, 翁华, 闫佳会, 郭青云, 王文清, 马新明. 蚕豆萎蔫病毒2号青海辣椒分离物的鉴定与全基因组序列克隆[J]. 园艺学报, 2023, 50(1): 161-169. |
| [6] | 韩晓蕾, 张彩霞, 刘 锴, 杨 安, 严家帝, 李武兴, 康立群, 丛佩华. 中熟苹果新品种‘中苹优蕾’[J]. 园艺学报, 2022, 49(S2): 1-2. |
| [7] | 韩晓蕾, 张彩霞, 刘 锴, 严家帝, 李武兴, 康立群, 丛佩华. 中熟苹果新品种‘苹优2号’[J]. 园艺学报, 2022, 49(S1): 1-2. |
| [8] | 王 强, 丛佩华, 刘肖烽. 晚熟苹果新品种‘华优甜娃’[J]. 园艺学报, 2022, 49(S1): 3-4. |
| [9] | 王 强, 丛佩华, 刘肖烽. 中熟苹果新品种‘华优宝蜜’[J]. 园艺学报, 2022, 49(S1): 5-6. |
| [10] | 杨 玲, 丛佩华, 王 强, 李武兴, 康立群. 中熟鲜食苹果新品种‘华丰’[J]. 园艺学报, 2022, 49(S1): 7-8. |
| [11] | 丁志杰, 包金波, 柔鲜古丽, 朱甜甜, 李雪丽, 苗浩宇, 田新民. 新疆野苹果与‘元帅’‘金冠’的叶绿体基因组比对研究[J]. 园艺学报, 2022, 49(9): 1977-1990. |
| [12] | 高彦龙, 吴玉霞, 张仲兴, 王双成, 张瑞, 张德, 王延秀. 苹果ELO家族基因鉴定及其在低温胁迫下的表达分析[J]. 园艺学报, 2022, 49(8): 1621-1636. |
| [13] | 郑晓东, 袭祥利, 李玉琪, 孙志娟, 马长青, 韩明三, 李少旋, 田义轲, 王彩虹. 油菜素内酯对盐碱胁迫下平邑甜茶幼苗生长的影响及调控机理研究[J]. 园艺学报, 2022, 49(7): 1401-1414. |
| [14] | 夏炎, 黄松, 武雪莉, 刘一琪, 王苗苗, 宋春晖, 白团辉, 宋尚伟, 庞宏光, 焦健, 郑先波. 基于宏病毒组测序技术的苹果病毒病鉴定与分析[J]. 园艺学报, 2022, 49(7): 1415-1428. |
| [15] | 刘照霞, 张鑫, 王璐, 马玉婷, 陈倩, 朱占玲, 葛顺峰, 姜远茂. 肥料穴施位点对苹果细根生长、15N吸收利用及产量品质的影响[J]. 园艺学报, 2022, 49(7): 1545-1556. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
版权所有 © 2012 《园艺学报》编辑部 京ICP备10030308号-2 国际联网备案号 11010802023439
编辑部地址: 北京市海淀区中关村南大街12号中国农业科学院蔬菜花卉研究所 邮编: 100081
电话: 010-82109523 E-Mail: yuanyixuebao@126.com
技术支持:北京玛格泰克科技发展有限公司