
园艺学报 ›› 2023, Vol. 50 ›› Issue (1): 1-14.doi: 10.16420/j.issn.0513-353x.2021-0932
• 研究论文 • 下一篇
翟含含1,*, 翟宇杰1,*, 田义2( ), 张叶1, 杨丽1, 温陟良1,**(
), 张叶1, 杨丽1, 温陟良1,**( ), 陈海江1,**(
), 陈海江1,**( )
)
收稿日期:2022-07-15
修回日期:2022-11-16
出版日期:2023-01-25
发布日期:2023-01-18
通讯作者:
**(E-mail:作者简介:*共同第一作者
基金资助:
ZHAI Hanhan1,*, ZHAI Yujie1,*, TIAN Yi2( ), ZHANG Ye1, YANG Li1, WEN Zhiliang1,**(
), ZHANG Ye1, YANG Li1, WEN Zhiliang1,**( ), CHEN Haijiang1,**(
), CHEN Haijiang1,**( )
)
Received:2022-07-15
Revised:2022-11-16
Online:2023-01-25
Published:2023-01-18
Contact:
**(E-mail:摘要:
为了探究SAUR(Small Auxin-Up RNA)基因家族成员在桃树(Prunus persica)生长发育中的作用,利用生物信息学方法对桃SAUR家族成员(PpSAUR)进行鉴定,对其染色体定位、基因结构、保守元件、进化关系及基因表达等进行分析,并对PpSAUR5功能进行初步鉴定。结果表明:桃中共有80个SAUR成员,分为12个亚组,不均匀分布在8条染色体上;基因结构显示75个PpSAUR只有1个外显子,余下5个有2 ~ 3个外显子;通过分析不同树势(强、中、弱)桃品种的转录组获得18个与树体长势呈正或负相关的SAUR基因,这些基因对外源植物生长调节物质处理响应不同。PpSAUR5对生长素与赤霉素均有响应,功能验证显示PpSAUR5转基因拟南芥幼苗叶柄、下胚轴及根较野生型长,且对NAA与2,4-D处理敏感性降低。研究结果表明PpSAUR5能够促进器官伸长。
中图分类号:
翟含含, 翟宇杰, 田义, 张叶, 杨丽, 温陟良, 陈海江. 桃SAUR家族基因分析及PpSAUR5功能鉴定[J]. 园艺学报, 2023, 50(1): 1-14.
ZHAI Hanhan, ZHAI Yujie, TIAN Yi, ZHANG Ye, YANG Li, WEN Zhiliang, CHEN Haijiang. Genome-wide Identification of Peach SAUR Gene Family and Characterization of PpSAUR5 Gene[J]. Acta Horticulturae Sinica, 2023, 50(1): 1-14.
| 基因Gene | 登录号Bank number | 引物序列(5'-3')Primer sequence |
|---|---|---|
| PpActin | Prupe.6G163400 | F:GCCCAAGTGCTTCGTATGCT;R:ATCACCGGCTGCAATCCA |
| PpSAUR3 | Prupe.1G221300 | F:TCCGAGACGGGCTAAGAAGA;R:ATTCGGGTACGACAGCCAAG |
| PpSAUR4 | Prupe.1G367800 | F:TTTGGAACTCAGGGCGAGTC;R:CAAGGCCTCCCTTTGGATGT |
| PpSAUR5 | Prupe.1G368100 | F:ACGAAGCAACGGTCGAGAAT;R:CAGTTGCACCCGATTTGTGG |
| PpSAUR10 | Prupe.2G140600 | F:TTTGGAACTCAGGGCGAGTC;R:CAAGGCCTCCCTTTGGATGT |
| PpSAUR20 | Prupe.4G136800 | F:TTTGGAACCGTCACAGGCTT;R:CTCCTTGCAGCAACAAACCC |
| PpSAUR26 | Prupe.6G108500 | F:ATGGAATGGTGATGCCACCC;R:TGGCTATGACGAATGTCCCAA |
| PpSAUR34 | Prupe.7G167000 | F:CTTAAGAACCGCCCGGAGTT;R:AGTACAAGCCGAGGAGGAGT |
| PpSAUR41 | Prupe.8G078600 | F:CAAGGGATTGTGAGGCCCAT;R:CTCAAACCGCAGTGCTCAAG |
| PpSAUR42 | Prupe.8G078700 | F:GCTTCTTTTGGCTCTCCCCA;R:TCCGGTTGCCTGGAATTGTT |
| PpSAUR46 | Prupe.8G079100 | F:AAACTGCGTCTTTTGGCTCC;R:TTGCCCGGAATTGTTAACGC |
| PpSAUR47 | Prupe.8G079200 | F:AGGCATCTTCACTGCATGGG;R:GGAGAGCAAGAAGAAGCGGT |
| PpSAUR56 | Prupe.8G080200 | F:ACATAGGAATGACGACCCGC;R:CCGTCGTCCTAGTAAGCAGC |
| PpSAUR60 | Prupe.8G080600 | F:TGAAACTCTGGATGGCCCAG;R:AGATGTGCCAAAGGGGCATT |
| PpSAUR64 | Prupe.8G081100 | F:TTCTCAAGCTCTCGTGGCTG;R:GTGCCGGAAGATGTGAAGGA |
| PpSAUR73 | Prupe.8G082100 | F:GGCACCCAACTGAGAAGTGA;R:ACGAGGTTCCAAAAGGGCAT |
| PpSAUR74 | Prupe.8G082200 | F:GCGGGAAGTGAGATGAAGGA;R:GGAGCCAAAAGACGCAGTTT |
| PpSAUR77 | Prupe.8G157900 | F:TAATTGCTCTCCATCGCCCC;R:AGAGTTGCTAAGCGAGGCTG |
| PpSAUR80 | Prupe.8G158200 | F:AGACGATCACGTGACAAGGG;R:CGTCGAGCTCCTGAACAAGT |
表1 qRT-PCR验证引物序列
Table 1 Primer used in qRT-PCR Validation
| 基因Gene | 登录号Bank number | 引物序列(5'-3')Primer sequence |
|---|---|---|
| PpActin | Prupe.6G163400 | F:GCCCAAGTGCTTCGTATGCT;R:ATCACCGGCTGCAATCCA |
| PpSAUR3 | Prupe.1G221300 | F:TCCGAGACGGGCTAAGAAGA;R:ATTCGGGTACGACAGCCAAG |
| PpSAUR4 | Prupe.1G367800 | F:TTTGGAACTCAGGGCGAGTC;R:CAAGGCCTCCCTTTGGATGT |
| PpSAUR5 | Prupe.1G368100 | F:ACGAAGCAACGGTCGAGAAT;R:CAGTTGCACCCGATTTGTGG |
| PpSAUR10 | Prupe.2G140600 | F:TTTGGAACTCAGGGCGAGTC;R:CAAGGCCTCCCTTTGGATGT |
| PpSAUR20 | Prupe.4G136800 | F:TTTGGAACCGTCACAGGCTT;R:CTCCTTGCAGCAACAAACCC |
| PpSAUR26 | Prupe.6G108500 | F:ATGGAATGGTGATGCCACCC;R:TGGCTATGACGAATGTCCCAA |
| PpSAUR34 | Prupe.7G167000 | F:CTTAAGAACCGCCCGGAGTT;R:AGTACAAGCCGAGGAGGAGT |
| PpSAUR41 | Prupe.8G078600 | F:CAAGGGATTGTGAGGCCCAT;R:CTCAAACCGCAGTGCTCAAG |
| PpSAUR42 | Prupe.8G078700 | F:GCTTCTTTTGGCTCTCCCCA;R:TCCGGTTGCCTGGAATTGTT |
| PpSAUR46 | Prupe.8G079100 | F:AAACTGCGTCTTTTGGCTCC;R:TTGCCCGGAATTGTTAACGC |
| PpSAUR47 | Prupe.8G079200 | F:AGGCATCTTCACTGCATGGG;R:GGAGAGCAAGAAGAAGCGGT |
| PpSAUR56 | Prupe.8G080200 | F:ACATAGGAATGACGACCCGC;R:CCGTCGTCCTAGTAAGCAGC |
| PpSAUR60 | Prupe.8G080600 | F:TGAAACTCTGGATGGCCCAG;R:AGATGTGCCAAAGGGGCATT |
| PpSAUR64 | Prupe.8G081100 | F:TTCTCAAGCTCTCGTGGCTG;R:GTGCCGGAAGATGTGAAGGA |
| PpSAUR73 | Prupe.8G082100 | F:GGCACCCAACTGAGAAGTGA;R:ACGAGGTTCCAAAAGGGCAT |
| PpSAUR74 | Prupe.8G082200 | F:GCGGGAAGTGAGATGAAGGA;R:GGAGCCAAAAGACGCAGTTT |
| PpSAUR77 | Prupe.8G157900 | F:TAATTGCTCTCCATCGCCCC;R:AGAGTTGCTAAGCGAGGCTG |
| PpSAUR80 | Prupe.8G158200 | F:AGACGATCACGTGACAAGGG;R:CGTCGAGCTCCTGAACAAGT |
| 基因名称 Gene name | 基因登录号 Gene ID | 染色体位置Chromosome location | 蛋白质Protein | |||
|---|---|---|---|---|---|---|
| 起始位点/bp Start position | 终止位点/bp End position | 长度/aa Length | 分子量/D MW | 理论等点 pI | ||
| PpSAUR1 | Prupe.1G000200 | 42 266 | 44 493 | 173 | 19 380.4 | 11.05 |
| PpSAUR2 | Prupe.1G067400 | 4 810 025 | 4 811 236 | 126 | 14 192.5 | 8.63 |
| PpSAUR3 | Prupe.1G221300 | 23 585 167 | 23 586 506 | 127 | 14 725.8 | 8.23 |
| PpSAUR4 | Prupe.1G367800 | 33 658 384 | 33 660 037 | 126 | 14 253.5 | 9.14 |
| PpSAUR5 | Prupe.1G368100 | 33 683 348 | 33 685 088 | 131 | 15 426.6 | 7.38 |
| PpSAUR6 | Prupe.1G455700 | 38 251 559 | 38 252 129 | 127 | 14 616.0 | 8.15 |
| PpSAUR7 | Prupe.2G131400 | 18 964 628 | 18 965 376 | 122 | 13 712.8 | 9.44 |
| PpSAUR8 | Prupe.2G131500 | 18 966 838 | 18 967 203 | 98 | 11 226.7 | 7.22 |
| PpSAUR9 | Prupe.2G131600 | 18 972 827 | 18 973 204 | 91 | 9 998.4 | 7.51 |
| PpSAUR10 | Prupe.2G140600 | 19 729 097 | 19 730 272 | 105 | 11 631.2 | 7.50 |
| PpSAUR11 | Prupe.2G194600 | 23 275 229 | 23 276 493 | 82 | 9 069.3 | 5.02 |
| PpSAUR12 | Prupe.2G211600 | 24 387 357 | 24 388 174 | 84 | 9 269.6 | 9.19 |
| PpSAUR13 | Prupe.2G317100 | 29 700 658 | 29 701 286 | 151 | 16 864.3 | 9.12 |
| PpSAUR14 | Prupe.3G023900 | 1 785 030 | 1 786 044 | 78 | 8 619.9 | 8.98 |
| PpSAUR15 | Prupe.3G024000 | 1 787 115 | 1 787 919 | 102 | 11 216.9 | 8.21 |
| PpSAUR16 | Prupe.3G024100 | 1 789 219 | 1 789 680 | 100 | 11 198.2 | 10.63 |
| PpSAUR17 | Prupe.3G035000 | 2 557 789 | 2 558 556 | 94 | 10 323.8 | 7.51 |
| PpSAUR18 | Prupe.3G035100 | 2 561 778 | 2 562 092 | 101 | 11 069.5 | 5.82 |
| PpSAUR19 | Prupe.3G177500 | 19 444 270 | 19 444 578 | 105 | 11 878.7 | 8.49 |
| PpSAUR20 | Prupe.4G136800 | 7 593 725 | 7 595 443 | 101 | 11 114.6 | 6.51 |
| 基因名称 Gene name | 基因登录号 Gene ID | 染色体位置Chromosome location | 蛋白质Protein | |||
| 起始位点/bp Start position | 终止位点/bp End position | 长度/aa Length | 分子量/D MW | 理论等点 pI | ||
| PpSAUR21 | Prupe.4G140200 | 7 830 216 | 7 831 392 | 100 | 11 186.7 | 9.54 |
| PpSAUR22 | Prupe.5G076200 | 9 038 680 | 9 039 809 | 101 | 11 014.4 | 5.64 |
| PpSAUR23 | Prupe.5G147100 | 13 508 896 | 13 509 615 | 97 | 11 199.2 | 10.16 |
| PpSAUR24 | Prupe.6G108300 | 7 731 068 | 7 732 074 | 101 | 11 239.8 | 8.46 |
| PpSAUR25 | Prupe.6G108400 | 7 734 034 | 7 735 462 | 95 | 10 557.3 | 8.50 |
| PpSAUR26 | Prupe.6G108500 | 7 738 492 | 7 739 254 | 92 | 10 194.6 | 5.64 |
| PpSAUR27 | Prupe.6G232300 | 23 489 981 | 23 490 760 | 95 | 10 594.2 | 9.09 |
| PpSAUR28 | Prupe.6G234900 | 23 623 506 | 23 623 883 | 94 | 10 125.5 | 5.02 |
| PpSAUR29 | Prupe.7G048400 | 8 547 207 | 8 549 249 | 94 | 10 154.6 | 6.51 |
| PpSAUR30 | Prupe.7G048500 | 8 552 726 | 8 553 184 | 91 | 9 912.4 | 6.78 |
| PpSAUR31 | Prupe.7G048600 | 8 555 516 | 8 555 977 | 102 | 11 231.0 | 8.50 |
| PpSAUR32 | Prupe.7G104000 | 13 346 482 | 13 347 250 | 102 | 11 251.0 | 9.22 |
| PpSAUR33 | Prupe.7G120400 | 14 389 996 | 14 390 765 | 99 | 11 347.4 | 10.09 |
| PpSAUR34 | Prupe.7G167000 | 17 051 802 | 17 052 822 | 95 | 10 491.2 | 8.51 |
| PpSAUR35 | Prupe.7G192600 | 18 371 150 | 18 371 953 | 101 | 11 152.6 | 5.82 |
| PpSAUR36 | Prupe.7G192900 | 18 381 958 | 18 384 145 | 102 | 11 092.6 | 7.57 |
| PpSAUR37 | Prupe.8G063100 | 9 167 730 | 9 168 155 | 83 | 9 202.8 | 10.60 |
| PpSAUR38 | Prupe.8G072300 | 10 611 481 | 10 611 777 | 110 | 12 308.0 | 5.68 |
| PpSAUR39 | Prupe.8G072400 | 10 619 587 | 10 619 877 | 98 | 11 109.5 | 5.83 |
| PpSAUR40 | Prupe.8G078300 | 11 247 032 | 11 247 412 | 92 | 10 118.5 | 8.73 |
| PpSAUR41 | Prupe.8G078600 | 11 268 588 | 11 269 189 | 94 | 10 373.9 | 6.51 |
| PpSAUR42 | Prupe.8G078700 | 11 272 704 | 11 273 458 | 100 | 10 956.4 | 5.02 |
| PpSAUR43 | Prupe.8G078800 | 11 274 113 | 11 275 033 | 101 | 11 084.6 | 5.82 |
| PpSAUR44 | Prupe.8G078900 | 11 275 613 | 11 276 700 | 94 | 10 170.6 | 6.50 |
| PpSAUR45 | Prupe.8G079000 | 11 277 974 | 11 278 668 | 101 | 11 105.7 | 8.21 |
| PpSAUR46 | Prupe.8G079100 | 11 279 560 | 11 280 268 | 75 | 8 307.4 | 9.72 |
| PpSAUR47 | Prupe.8G079200 | 11 283 706 | 11 284 527 | 100 | 11 156.7 | 6.96 |
| PpSAUR48 | Prupe.8G079300 | 11 285 739 | 11 286 129 | 142 | 16 708.7 | 11.69 |
| PpSAUR49 | Prupe.8G079400 | 11 317 660 | 11 318 049 | 91 | 10 108.4 | 5.65 |
| PpSAUR50 | Prupe.8G079500 | 11 319 064 | 11 319 345 | 130 | 14 432.6 | 9.50 |
| PpSAUR51 | Prupe.8G079600 | 11 320 009 | 11 320 901 | 83 | 9 461.8 | 4.25 |
| PpSAUR52 | Prupe.8G079700 | 11 321 405 | 11 322 307 | 143 | 16 113.0 | 6.78 |
| PpSAUR53 | Prupe.8G079900 | 11 334 888 | 11 335 121 | 126 | 14 374.5 | 8.41 |
| PpSAUR54 | Prupe.8G080000 | 11 347 844 | 11 348 229 | 141 | 15 995.3 | 8.26 |
| PpSAUR55 | Prupe.8G080100 | 11 365 948 | 11 367 159 | 139 | 15 191.0 | 6.26 |
| PpSAUR56 | Prupe.8G080200 | 11 368 586 | 11 368 888 | 129 | 14 373.4 | 6.92 |
| PpSAUR57 | Prupe.8G080300 | 11 401 448 | 11 402 420 | 140 | 15 993.1 | 9.61 |
| PpSAUR58 | Prupe.8G080400 | 11 404 019 | 11 404 243 | 136 | 15 427.5 | 8.71 |
| PpSAUR59 | Prupe.8G080500 | 11 439 488 | 11 439 769 | 176 | 19 727.6 | 9.92 |
| PpSAUR60 | Prupe.8G080600 | 11 445 368 | 11 447 102 | 167 | 19 436.6 | 11.24 |
| PpSAUR61 | Prupe.8G080700 | 11 447 103 | 11 448 991 | 153 | 17 140.7 | 8.96 |
| PpSAUR62 | Prupe.8G080800 | 11 463 264 | 11 463 545 | 154 | 17 115.6 | 10.24 |
| PpSAUR63 | Prupe.8G081000 | 11 469 135 | 11 470 199 | 133 | 14 817.0 | 8.35 |
| PpSAUR64 | Prupe.8G081100 | 11 471 700 | 11 473 503 | 208 | 23 314.0 | 10.40 |
| PpSAUR65 | Prupe.8G081200 | 11 473 808 | 11 474 113 | 107 | 12 082.8 | 8.71 |
| PpSAUR66 | Prupe.8G081300 | 11 475 669 | 11 476 574 | 193 | 21 340.7 | 6.79 |
| PpSAUR67 | Prupe.8G081400 | 11 478 089 | 11 478 731 | 126 | 14 033.7 | 5.19 |
| PpSAUR68 | Prupe.8G081500 | 11 480 644 | 11 480 943 | 130 | 14 265.3 | 7.44 |
| PpSAUR69 | Prupe.8G081600 | 11 482 304 | 11 483 354 | 154 | 17 300.3 | 10.11 |
| PpSAUR70 | Prupe.8G081700 | 11 483 436 | 11 485 084 | 118 | 13 160.0 | 6.51 |
| PpSAUR71 | Prupe.8G081800 | 11 485 961 | 11 486 206 | 146 | 16 597.0 | 8.50 |
| PpSAUR72 | Prupe.8G081900 | 11 487 552 | 11 488 319 | 189 | 21 234.2 | 9.62 |
| PpSAUR73 | Prupe.8G082100 | 11 514 115 | 11 514 913 | 103 | 11 608.7 | 10.19 |
| PpSAUR74 | Prupe.8G082200 | 11 528 634 | 11 529 256 | 105 | 11 938.9 | 9.86 |
| PpSAUR75 | Prupe.8G157700 | 16 792 611 | 16 792 886 | 151 | 17 033.3 | 10.23 |
| PpSAUR76 | Prupe.8G157800 | 16 794 545 | 16 795 015 | 106 | 12 089.9 | 8.24 |
| PpSAUR77 | Prupe.8G157900 | 16 797 713 | 16 798 249 | 141 | 15 509.0 | 8.33 |
| PpSAUR78 | Prupe.8G158000 | 16 799 338 | 16 800 140 | 167 | 18 583.6 | 10.43 |
| PpSAUR79 | Prupe.8G158100 | 16 802 092 | 16 802 385 | 165 | 18 522.6 | 9.92 |
| PpSAUR80 | Prupe.8G158200 | 16 803 867 | 16 804 613 | 189 | 21 198.9 | 8.38 |
表2 桃树PpSAUR基因家族成员的基本信息
Table 2 Basic information of PpSAUR gene family members in peach
| 基因名称 Gene name | 基因登录号 Gene ID | 染色体位置Chromosome location | 蛋白质Protein | |||
|---|---|---|---|---|---|---|
| 起始位点/bp Start position | 终止位点/bp End position | 长度/aa Length | 分子量/D MW | 理论等点 pI | ||
| PpSAUR1 | Prupe.1G000200 | 42 266 | 44 493 | 173 | 19 380.4 | 11.05 |
| PpSAUR2 | Prupe.1G067400 | 4 810 025 | 4 811 236 | 126 | 14 192.5 | 8.63 |
| PpSAUR3 | Prupe.1G221300 | 23 585 167 | 23 586 506 | 127 | 14 725.8 | 8.23 |
| PpSAUR4 | Prupe.1G367800 | 33 658 384 | 33 660 037 | 126 | 14 253.5 | 9.14 |
| PpSAUR5 | Prupe.1G368100 | 33 683 348 | 33 685 088 | 131 | 15 426.6 | 7.38 |
| PpSAUR6 | Prupe.1G455700 | 38 251 559 | 38 252 129 | 127 | 14 616.0 | 8.15 |
| PpSAUR7 | Prupe.2G131400 | 18 964 628 | 18 965 376 | 122 | 13 712.8 | 9.44 |
| PpSAUR8 | Prupe.2G131500 | 18 966 838 | 18 967 203 | 98 | 11 226.7 | 7.22 |
| PpSAUR9 | Prupe.2G131600 | 18 972 827 | 18 973 204 | 91 | 9 998.4 | 7.51 |
| PpSAUR10 | Prupe.2G140600 | 19 729 097 | 19 730 272 | 105 | 11 631.2 | 7.50 |
| PpSAUR11 | Prupe.2G194600 | 23 275 229 | 23 276 493 | 82 | 9 069.3 | 5.02 |
| PpSAUR12 | Prupe.2G211600 | 24 387 357 | 24 388 174 | 84 | 9 269.6 | 9.19 |
| PpSAUR13 | Prupe.2G317100 | 29 700 658 | 29 701 286 | 151 | 16 864.3 | 9.12 |
| PpSAUR14 | Prupe.3G023900 | 1 785 030 | 1 786 044 | 78 | 8 619.9 | 8.98 |
| PpSAUR15 | Prupe.3G024000 | 1 787 115 | 1 787 919 | 102 | 11 216.9 | 8.21 |
| PpSAUR16 | Prupe.3G024100 | 1 789 219 | 1 789 680 | 100 | 11 198.2 | 10.63 |
| PpSAUR17 | Prupe.3G035000 | 2 557 789 | 2 558 556 | 94 | 10 323.8 | 7.51 |
| PpSAUR18 | Prupe.3G035100 | 2 561 778 | 2 562 092 | 101 | 11 069.5 | 5.82 |
| PpSAUR19 | Prupe.3G177500 | 19 444 270 | 19 444 578 | 105 | 11 878.7 | 8.49 |
| PpSAUR20 | Prupe.4G136800 | 7 593 725 | 7 595 443 | 101 | 11 114.6 | 6.51 |
| 基因名称 Gene name | 基因登录号 Gene ID | 染色体位置Chromosome location | 蛋白质Protein | |||
| 起始位点/bp Start position | 终止位点/bp End position | 长度/aa Length | 分子量/D MW | 理论等点 pI | ||
| PpSAUR21 | Prupe.4G140200 | 7 830 216 | 7 831 392 | 100 | 11 186.7 | 9.54 |
| PpSAUR22 | Prupe.5G076200 | 9 038 680 | 9 039 809 | 101 | 11 014.4 | 5.64 |
| PpSAUR23 | Prupe.5G147100 | 13 508 896 | 13 509 615 | 97 | 11 199.2 | 10.16 |
| PpSAUR24 | Prupe.6G108300 | 7 731 068 | 7 732 074 | 101 | 11 239.8 | 8.46 |
| PpSAUR25 | Prupe.6G108400 | 7 734 034 | 7 735 462 | 95 | 10 557.3 | 8.50 |
| PpSAUR26 | Prupe.6G108500 | 7 738 492 | 7 739 254 | 92 | 10 194.6 | 5.64 |
| PpSAUR27 | Prupe.6G232300 | 23 489 981 | 23 490 760 | 95 | 10 594.2 | 9.09 |
| PpSAUR28 | Prupe.6G234900 | 23 623 506 | 23 623 883 | 94 | 10 125.5 | 5.02 |
| PpSAUR29 | Prupe.7G048400 | 8 547 207 | 8 549 249 | 94 | 10 154.6 | 6.51 |
| PpSAUR30 | Prupe.7G048500 | 8 552 726 | 8 553 184 | 91 | 9 912.4 | 6.78 |
| PpSAUR31 | Prupe.7G048600 | 8 555 516 | 8 555 977 | 102 | 11 231.0 | 8.50 |
| PpSAUR32 | Prupe.7G104000 | 13 346 482 | 13 347 250 | 102 | 11 251.0 | 9.22 |
| PpSAUR33 | Prupe.7G120400 | 14 389 996 | 14 390 765 | 99 | 11 347.4 | 10.09 |
| PpSAUR34 | Prupe.7G167000 | 17 051 802 | 17 052 822 | 95 | 10 491.2 | 8.51 |
| PpSAUR35 | Prupe.7G192600 | 18 371 150 | 18 371 953 | 101 | 11 152.6 | 5.82 |
| PpSAUR36 | Prupe.7G192900 | 18 381 958 | 18 384 145 | 102 | 11 092.6 | 7.57 |
| PpSAUR37 | Prupe.8G063100 | 9 167 730 | 9 168 155 | 83 | 9 202.8 | 10.60 |
| PpSAUR38 | Prupe.8G072300 | 10 611 481 | 10 611 777 | 110 | 12 308.0 | 5.68 |
| PpSAUR39 | Prupe.8G072400 | 10 619 587 | 10 619 877 | 98 | 11 109.5 | 5.83 |
| PpSAUR40 | Prupe.8G078300 | 11 247 032 | 11 247 412 | 92 | 10 118.5 | 8.73 |
| PpSAUR41 | Prupe.8G078600 | 11 268 588 | 11 269 189 | 94 | 10 373.9 | 6.51 |
| PpSAUR42 | Prupe.8G078700 | 11 272 704 | 11 273 458 | 100 | 10 956.4 | 5.02 |
| PpSAUR43 | Prupe.8G078800 | 11 274 113 | 11 275 033 | 101 | 11 084.6 | 5.82 |
| PpSAUR44 | Prupe.8G078900 | 11 275 613 | 11 276 700 | 94 | 10 170.6 | 6.50 |
| PpSAUR45 | Prupe.8G079000 | 11 277 974 | 11 278 668 | 101 | 11 105.7 | 8.21 |
| PpSAUR46 | Prupe.8G079100 | 11 279 560 | 11 280 268 | 75 | 8 307.4 | 9.72 |
| PpSAUR47 | Prupe.8G079200 | 11 283 706 | 11 284 527 | 100 | 11 156.7 | 6.96 |
| PpSAUR48 | Prupe.8G079300 | 11 285 739 | 11 286 129 | 142 | 16 708.7 | 11.69 |
| PpSAUR49 | Prupe.8G079400 | 11 317 660 | 11 318 049 | 91 | 10 108.4 | 5.65 |
| PpSAUR50 | Prupe.8G079500 | 11 319 064 | 11 319 345 | 130 | 14 432.6 | 9.50 |
| PpSAUR51 | Prupe.8G079600 | 11 320 009 | 11 320 901 | 83 | 9 461.8 | 4.25 |
| PpSAUR52 | Prupe.8G079700 | 11 321 405 | 11 322 307 | 143 | 16 113.0 | 6.78 |
| PpSAUR53 | Prupe.8G079900 | 11 334 888 | 11 335 121 | 126 | 14 374.5 | 8.41 |
| PpSAUR54 | Prupe.8G080000 | 11 347 844 | 11 348 229 | 141 | 15 995.3 | 8.26 |
| PpSAUR55 | Prupe.8G080100 | 11 365 948 | 11 367 159 | 139 | 15 191.0 | 6.26 |
| PpSAUR56 | Prupe.8G080200 | 11 368 586 | 11 368 888 | 129 | 14 373.4 | 6.92 |
| PpSAUR57 | Prupe.8G080300 | 11 401 448 | 11 402 420 | 140 | 15 993.1 | 9.61 |
| PpSAUR58 | Prupe.8G080400 | 11 404 019 | 11 404 243 | 136 | 15 427.5 | 8.71 |
| PpSAUR59 | Prupe.8G080500 | 11 439 488 | 11 439 769 | 176 | 19 727.6 | 9.92 |
| PpSAUR60 | Prupe.8G080600 | 11 445 368 | 11 447 102 | 167 | 19 436.6 | 11.24 |
| PpSAUR61 | Prupe.8G080700 | 11 447 103 | 11 448 991 | 153 | 17 140.7 | 8.96 |
| PpSAUR62 | Prupe.8G080800 | 11 463 264 | 11 463 545 | 154 | 17 115.6 | 10.24 |
| PpSAUR63 | Prupe.8G081000 | 11 469 135 | 11 470 199 | 133 | 14 817.0 | 8.35 |
| PpSAUR64 | Prupe.8G081100 | 11 471 700 | 11 473 503 | 208 | 23 314.0 | 10.40 |
| PpSAUR65 | Prupe.8G081200 | 11 473 808 | 11 474 113 | 107 | 12 082.8 | 8.71 |
| PpSAUR66 | Prupe.8G081300 | 11 475 669 | 11 476 574 | 193 | 21 340.7 | 6.79 |
| PpSAUR67 | Prupe.8G081400 | 11 478 089 | 11 478 731 | 126 | 14 033.7 | 5.19 |
| PpSAUR68 | Prupe.8G081500 | 11 480 644 | 11 480 943 | 130 | 14 265.3 | 7.44 |
| PpSAUR69 | Prupe.8G081600 | 11 482 304 | 11 483 354 | 154 | 17 300.3 | 10.11 |
| PpSAUR70 | Prupe.8G081700 | 11 483 436 | 11 485 084 | 118 | 13 160.0 | 6.51 |
| PpSAUR71 | Prupe.8G081800 | 11 485 961 | 11 486 206 | 146 | 16 597.0 | 8.50 |
| PpSAUR72 | Prupe.8G081900 | 11 487 552 | 11 488 319 | 189 | 21 234.2 | 9.62 |
| PpSAUR73 | Prupe.8G082100 | 11 514 115 | 11 514 913 | 103 | 11 608.7 | 10.19 |
| PpSAUR74 | Prupe.8G082200 | 11 528 634 | 11 529 256 | 105 | 11 938.9 | 9.86 |
| PpSAUR75 | Prupe.8G157700 | 16 792 611 | 16 792 886 | 151 | 17 033.3 | 10.23 |
| PpSAUR76 | Prupe.8G157800 | 16 794 545 | 16 795 015 | 106 | 12 089.9 | 8.24 |
| PpSAUR77 | Prupe.8G157900 | 16 797 713 | 16 798 249 | 141 | 15 509.0 | 8.33 |
| PpSAUR78 | Prupe.8G158000 | 16 799 338 | 16 800 140 | 167 | 18 583.6 | 10.43 |
| PpSAUR79 | Prupe.8G158100 | 16 802 092 | 16 802 385 | 165 | 18 522.6 | 9.92 |
| PpSAUR80 | Prupe.8G158200 | 16 803 867 | 16 804 613 | 189 | 21 198.9 | 8.38 |
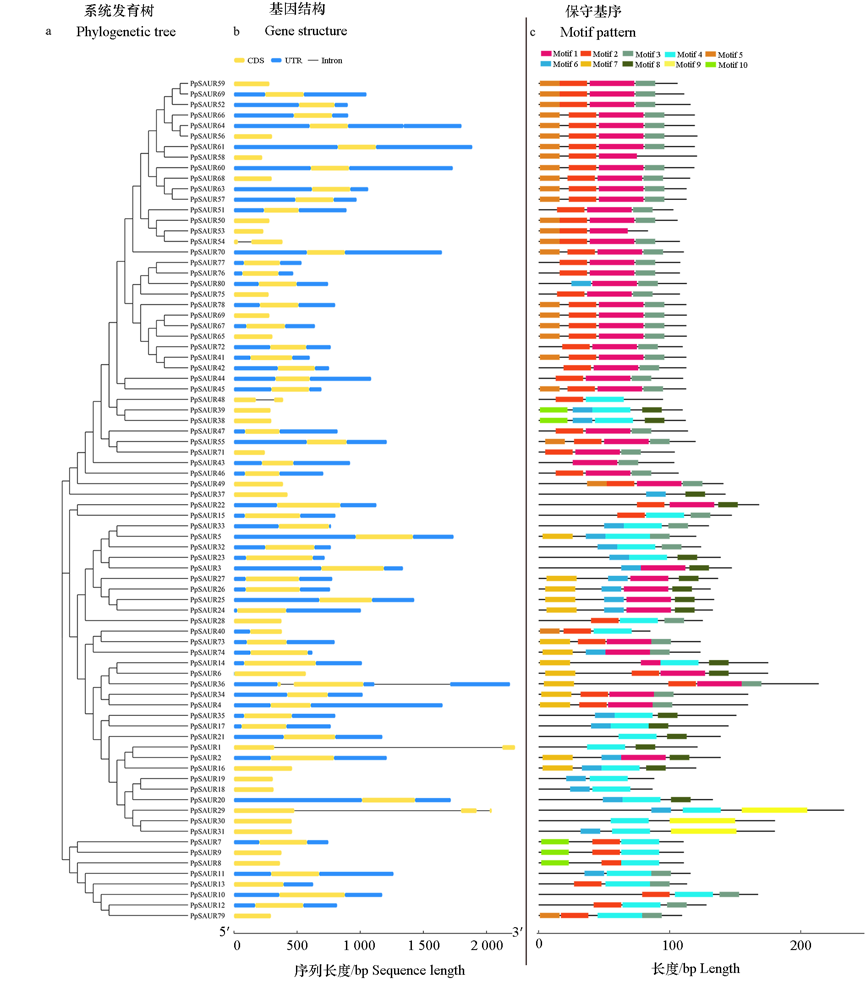
图2 桃PpSAUR蛋白的系统发育关系(a),基因结构(b)和保守基序分布(c)
Fig. 2 Phylogenetic relationship(a),gene structure(b)and distribution of conserved motifs(c)of PpSAUR proteins in peach
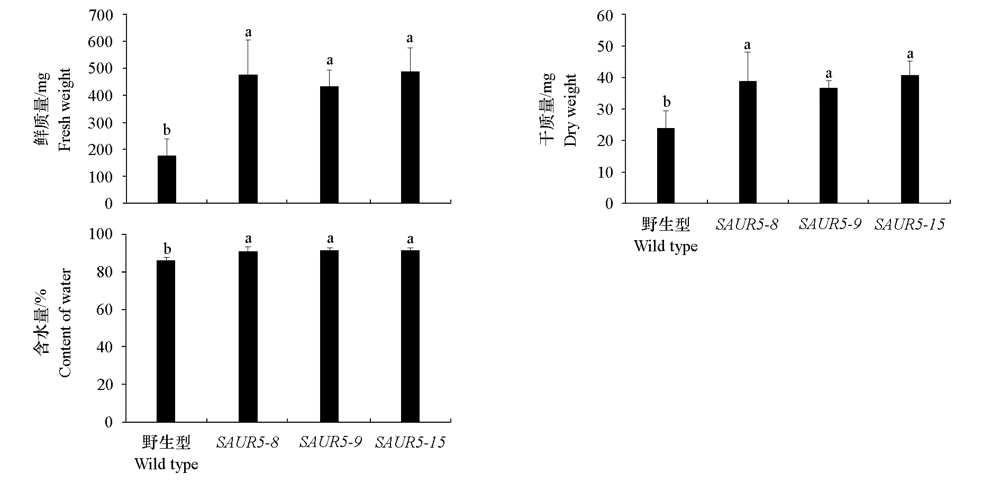
图5 在MS培养基生长20 d的拟南芥植株的鲜质量、干质量和含水量分析
Fig. 5 Analysis of fresh weight,dry weight and water content of Arabidopsis plants growing on MS medium for 20 days n = 20.
| [1] | Altschul S F, Madden T L, Schffer A A, Zhang J, Zhang Z, Miller W, Lipman D J. 1997. Gapped BLAST and PSI-BLAST:a new generation of protein database search programs. Nucleic Acids Research, 25 (17):3389-3402. |
| [2] | Bailey T. 2009. Challenge and opportunity:rethinking the role and function of developmental education in community college. New Directions for Community Colleges,(145):11-30. |
| [3] |
Bemer M, van Mourik H, Muiño J M, Ferrándiz C, Kaufmann K, Angenent G C. 2017. Fruitfull controls SAUR10 expression and regulates Arabidopsis growth and architecture. Journal of Experimental Botany, 68 (13):3391-3403.
doi: 10.1093/jxb/erx184 URL |
| [4] |
Cao K, Yang X W, Li Y, Zhu G R, Fang W C, Chen C W, Wang X W, Wu J L, Wang L R. 2021. New high-quality peach(Prunus persica L. Batsch)genome assembly to analyze the molecular evolutionary mechanism of volatile compounds in peach fruit. The Plant Journal, 108 (1):281-295.
doi: 10.1111/tpj.15439 pmid: 34309935 |
| [5] |
Chen Y, Hao X, Cao J. 2014. Small auxin upregulated RNA(SAUR)gene family in maize:identification,evolution,and its phylogenetic comparison with Arabidopsis,rice,and sorghum. Journal of Integrative Plant Biology, 56 (2):133-150.
doi: 10.1111/jipb.12127 URL |
| [6] |
Cheng J, Zhang M M, Tan B, Jiang Y J, Zheng X B, Ye X, Guo Z J, Xiong T T, Wang W, Li J D, Feng J C. 2019. A single nucleotide mutation in GID1c disrupts its interaction with DELLA1 and causes a GA-insensitive dwarf phenotype in peach. Plant Biotechnology Journal, 17 (9):1723-1735.
doi: 10.1111/pbi.13094 pmid: 30776191 |
| [7] |
Deng G, Huang X, Xie L, Tan S, Gbokie T J, Bao Y, Xie Z, Yi K. 2019. Identification and expression of SAUR genes in the CAM plant agave. Genes, 10 (7):555.
doi: 10.3390/genes10070555 URL |
| [8] |
Gil P, Liu Y, Orbović V, Verkamp E, Poff K L, Green P J. 1994. Characterization of the auxin-inducible SAUR-AC1 gene for use as a molecular genetic tool in Arabidopsis. Plant Physiology, 104 (2):777-784.
doi: 10.1104/pp.104.2.777 pmid: 8159792 |
| [9] |
Hagen G, Guilfoyle T. 2002. Auxin-responsive gene expression:genes,promoters and regulatory factors. Plant Molecular Biology, 49 (3-4):373-385.
doi: 10.1023/A:1015207114117 URL |
| [10] |
Hou K, Wu W, Gan S S. 2013. SAUR36,a small auxin up RNA gene,is involved in the promotion of leaf senescence in Arabidopsis. Plant Physiology, 161 (2):1002-1009.
doi: 10.1104/pp.112.212787 URL |
| [11] |
Hu W F, Yan H W, Luo S S, Pan F, Wang Y, Xiang Y. 2018. Genome-wide analysis of poplar SAUR gene family and expression profiles under cold,polyethylene glycol and indole-3-acetic acid treatments. Plant Physiology and Biochemistry, 128:50-65.
doi: 10.1016/j.plaphy.2018.04.021 URL |
| [12] | Jain M, Kaur N, Tyagi A K, Khurana J P. 2006a. The auxin-responsive GH 3 gene family in rice (Oryza sativa). Functional & Integrative Genomics, 6 (1):36-46. |
| [13] |
Jain M, Tyagi A K, Khurana J P. 2006b. Genome-wide analysis,evolutionary expansion,and expression of early auxin-respon-sive SAUR gene family in rice(Oryza sativa). Genomics, 88 (3):360-371.
doi: 10.1016/j.ygeno.2006.04.008 URL |
| [14] |
Kant S, Bi Y M, Zhu T, Rothstein S J. 2009. SAUR39,a Small Auxin-Up RNA gene,acts as a negative regulator of auxin synthesis and transport in rice. Plant Physiology, 151 (2):691-701.
doi: 10.1104/pp.109.143875 URL |
| [15] |
Knauss S, Rohrmeier T, Lehle L. 2003. The auxin-induced maize gene ZmSAUR2 encodes a short-lived nuclear protein expressed in elongating tissues. Journal of Biological Chemistry, 278 (26):23936-23943.
doi: 10.1074/jbc.M212585200 URL |
| [16] |
Laloum T, Guiomar M, Duque P. 2017. Alternative splicing control of abiotic stress responses. Trends in Plant Science, 23 (2):140-150.
doi: 10.1016/j.tplants.2017.09.019 URL |
| [17] | Li Ao, Cui Meng-jie, Chen Ke. 2018. Grape SAUR gene family identification and bioinformatics analysis. Plant Genetic Resources, 19 (2):326-337. (in Chinese) |
| 李傲, 崔梦杰, 陈珂. 2018. 葡萄SAUR基因家族鉴定与生物信息学分析. 植物遗传资源学报, 19 (2):326-337. | |
| [18] | Li Jiao. 2013. Analysis of alternative splicing in whole transcriptome and its regulator SR protein under drought stress in seed of Zea mays L [M. D. Dissertation]. Zhengzhou: Zhengzhou University. (in Chinese) |
| 李娇. 2013. 玉米苗期干旱胁迫下可变剪接及其调控因子的研究[硕士论文]. 郑州: 郑州大学. | |
| [19] |
Li X, Liu G, Geng Y, Wu M, Pei W, Zhai H, Zang X, Li X, Zhang J, Yu S, Yu J. 2017. A genome-wide analysis of the small auxin-up RNA(SAUR)gene family in cotton. BMC Genomics, 18 (1):815-837.
doi: 10.1186/s12864-017-4224-2 URL |
| [20] |
Livak K J, Schmittgen T D. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2-∆∆Ctmethod. Methods, 25 (4):402-408.
doi: 10.1006/meth.2001.1262 pmid: 11846609 |
| [21] | Liu Yan. 2016. Functional exploration of OsSAUR45 gene in rice growth and development[M. D. Dissertation]. Hangzhou: Zhejiang University. (in Chinese) |
| 刘嫣. 2016. OsSAUR45基因在水稻生长发育中的功能探索[硕士论文]. 杭州: 浙江大学. | |
| [22] |
Ma P A, Chen X, Liu C, Meng Y, Xia Z, Zeng C, Lu C, Wang W. 2017. MeSAUR1,encoded by a Small Auxin-Up RNA gene,acts as a transcription regulator to positively regulate ADP-glucose pyrophosphorylase small subunit1a gene in cassava. Frontiers in Plant Science, 8 (21):1315-1326.
doi: 10.3389/fpls.2017.01315 URL |
| [23] |
McClure B A, Guilfoyle T. 1987. Characterization of a class of smallauxin-inducible soybean polyadenylated RNAs. Plant Molecular Biology, 9 (6):611-623.
doi: 10.1007/BF00020537 pmid: 24277197 |
| [24] | Newman T C, Ohme T M, Taylor C B, Green P J. 1993. DST sequences,highly conserved among plant SAUR genes,target reporter transcripts for rapid decay in tobacco. The Plant Cell, 5 (6):701-714. |
| [25] |
Niek S, Marian B. 2018. The SAUR gene family:the plant’s toolbox for adaptation of growth and development. Journal of Experimental Botany, 70 (1):17-27.
doi: 10.1093/jxb/ery332 URL |
| [26] |
Park J E, Kim Y S, Yoon H K, Park C M. 2007. Functional characterization of a Small Auxin-Up RNA gene in apical hook development in Arabidopsis. Plant Science, 172 (1):150-157.
doi: 10.1016/j.plantsci.2006.08.005 URL |
| [27] |
Praveen K K, Sunethra D, Nihal D. 2018. SAUR53 regulates organ elongation and apical hook development in Arabidopsis. Plant Signaling & Behavior, 13 (10). doi:10.1080/15592324.
doi: 10.1080/15592324 URL |
| [28] |
Ren H, Gray W M. 2015. SAUR proteins as effectors of hormonal and environmental signals in plant growth. Molecular Plant, 8 (8):1153-1164.
doi: 10.1016/j.molp.2015.05.003 pmid: 25983207 |
| [29] |
Spartz A K, Lee S H, Wenger J P, Gonzalez N, Itoh H, Inzé D, Peer W A, Murphy A S, Overvoorde P J, Gray W M. 2012. The SAUR19 subfamily of Small Auxin-Up RNA genes promote cell expansion. Plant Journal, 70 (6):978-990.
doi: 10.1111/j.1365-313X.2012.04946.x URL |
| [30] |
Spartz A K, Lor V S, Ren H, Olszewski N E, Miller N D, Wu G S, Spalding E P, Gray William M. 2017. Constitutive expression of Arabidopsis Small Auxin-Up RNA19(SAUR19)in tomato confers auxin-independent hypocotyl elongation. Plant Physiology, 173 (2):1453-1462.
doi: 10.1104/pp.16.01514 URL |
| [31] |
Spartz A K, Ren H, Park M Y, Grandt K N, Lee S H, Murphy A S, Sussman M R, Overvoorde P J, Gray W M. 2014. SAUR inhibition of PP2C-D phosphatases activates plasma membrane H+-ATPases to promote cell expansion in Arabidopsis. The Plant Cell, 26 (5):2129-2142.
doi: 10.1105/tpc.114.126037 URL |
| [32] |
Stamm P, Kumar P P. 2013. Auxin and gibberellin responsive arabidopsis Small Auxin Up RNA36 regulates hypocotyl elongation in the light. Plant Cell Reports, 32:759-769.
doi: 10.1007/s00299-013-1406-5 pmid: 23503980 |
| [33] | Sun N, Wang J, Gao Z, Dong J, He H, Terzaghi W, Wei N, Deng X W, Chen H. 2016. Arabidopsis SAURs are critical for differential light regulation of the development of various organs. Proceedings of the National Academy of Sciences, 113 (21):6071-6076. |
| [34] | Wang Fusheng, Yu Hong, Hu Zhou, Guan Delong, Zhang Pan, Zhu Shiping, Zhao Xiaochun. 2020. Genome-wide identification and expression analysis of the citrus SAUR gene family. Acta Horticulturae Sinica, 47 (1):23-40. (in Chinese) |
| 王福生, 余洪, 胡洲, 管德龙, 张盼, 朱世平, 赵晓春. 2020. 柑橘属SAUR基因家族的全基因组鉴定及表达分析. 园艺学报, 47 (1):23-40. | |
| [35] | Wang Hongfei, Shang Qingmao. 2019. Entification and expression analysis of cucumber SAUR gene family. Acta Horticulturae Sinica, 46 (6):1093-1111. (in Chinese) |
| 王红飞, 尚庆茂. 2019. 黄瓜SAUR基因家族的鉴定与表达分析. 园艺学报, 46 (6):1093-1111. | |
| [36] | Wang S, Bai Y, Shen C, Wu Y, Zhang S, Jiang D, Guilfoyle T J, Chen M, Qi Y. 2010. Auxin-related gene families in abiotic stress response in Sorghum bicolor. Functional & Integrative Genomics, 10 (4):533-546. |
| [37] | Wang Wen-ran, Fan Xiu-cei, Zhang Wen-ying, Liu Chong-huai, Fang Jing-gui, Wang Chen. 2017. Research progress in fruit tree erythromycin metabolism and signal pathway. Biotech Bulletin, 33 (11):1-7. (in Chinese) |
|
王文然, 樊秀彩, 张文颖, 刘崇怀, 房经贵, 王晨. 2017. 果树赤霉素代谢与信号途径研究进展. 生物技术通报, 33 (11):1-7.
doi: 10.13560/j.cnki.biotech.bull.1985.2017-0650 |
|
| [38] |
Wong J H, Spartz A K, Park M Y, Du M M, Gray W M. 2019. Mutation of a conserved motif of PP2C.D phosphatases confers SAUR immunity and constitutive activity. Plant Physiology, 181 (1):353-366.
doi: 10.1104/pp.19.00496 pmid: 31311832 |
| [39] |
Wu J, Liu S, He Y J, Guan X Y, Zhu X F, Cheng L, Wang J, Lu G. 2012. Genome-wide analysis of SAUR gene family in Solanaceae species. Gene, 509 (1):38-50.
doi: 10.1016/j.gene.2012.08.002 pmid: 22903030 |
| [40] | Xie R J, Dong C C, Ma Y Y, Deng L, He S L, Yi S L, Lv Q, Zheng Y Q. 2015. Comprehensive analysis of SAUR gene family in citrus and its transcriptional correlation with fruitlet drop from abscission zone A. Functional & Integrative Genomics, 15 (6):729-740. |
| [41] | Zhai Yu-jie. 2020. Growth potential difference and expression and regulation of related genes in different peach varieties[M. D. Dissertation]. Baoding: Hebei Agricultural University. (in Chinese) |
| 翟宇杰. 2020. 不同品种桃树生长势的差异及相关基因的表达与调控[硕士论文]. 保定: 河北农业大学. | |
| [42] |
Zhang C J, Wang J, Long M Y, Fan C Z. 2013. gKaKs:the pipeline for genome-level Ka/Ks calculation. Bioinformatics, 29 (5):645-646.
doi: 10.1093/bioinformatics/btt009 URL |
| [43] |
Zhang T, Jiang B, Xie R, Xu M, Liu K. 2020. Genome-wide identification and analysis of medicago truncatula Small Auxin-Up RNA(SAUR) gene family uncover their roles in nodule formation. Journal of Plant Biochemistry and Biotechnology, 30 (1):126-137.
doi: 10.1007/s13562-020-00576-7 URL |
| [44] | Zhou Xiao-ya. 2020. Effect of paclobutrazol on the growth of peach shoots and related molecular mechanism research[M. D. Dissertation]. Baoding: Hebei Agricultural University. (in Chinese) |
| 周晓雅. 2020. 多效唑抑制桃新梢生长的效应及相关分子机理研究[硕士论文]. 保定: 河北农业大学. | |
| [45] |
Zhu Yu-bin, Kong Ying-ying, Wang Jun-hui. 2014. Research advances in auxin-responsive SAUR genes. Life Sciences, 26 (4):407-413. (in Chinese)
doi: 10.1016/0024-3205(80)90158-7 URL |
| 朱宇斌, 孔莹莹, 王君晖. 2014. 植物生长素响应基因SAUR的研究进展. 生命科学, 26 (4):407-413. |
| [1] | 王晓晨, 聂子页, 刘先菊, 段 伟, 范培格, 梁振昌, . 脱落酸对‘京香玉’葡萄果实单萜物质合成的影响[J]. 园艺学报, 2023, 50(2): 237-249. |
| [2] | 王 瑞, 洪文娟, 罗 华, 赵丽娜, 陈 颖, 王 君, . 石榴品种SSR指纹图谱构建及杂种父本鉴定[J]. 园艺学报, 2023, 50(2): 265-278. |
| [3] | 任海龙, 许东林, 张 晶, 邹集文, 李光光, 周贤玉, 肖婉钰, 孙艺嘉. 菜薹KASP-SNP指纹图谱构建及品种鉴定[J]. 园艺学报, 2023, 50(2): 307-318. |
| [4] | 梁嘉莉, 吴启松, 陈广全, 张 荣, 徐春香, 冯淑杰, . 香蕉叶斑病病原菌芭蕉新拟盘多毛孢的鉴定[J]. 园艺学报, 2023, 50(2): 410-420. |
| [5] | 蒋靖东, 韦壮敏, 王楠, 朱晨桥, 叶俊丽, 谢宗周, 邓秀新, 柴利军. 山金柑四倍体资源的发掘与鉴定[J]. 园艺学报, 2023, 50(1): 27-35. |
| [6] | 袁馨, 徐云鹤, 张雨培, 单楠, 陈楚英, 万春鹏, 开文斌, 翟夏琬, 陈金印, 甘增宇. 猕猴桃后熟过程中ABA响应结合因子AcAREB1调控AcGH3.1的表达[J]. 园艺学报, 2023, 50(1): 53-64. |
| [7] | 邢柱东, 吕福堂, 郭尚敬, 张演义. 新品种‘聊大红金’桃[J]. 园艺学报, 2023, 50(1): 225-226. |
| [8] | 杨兴旺, 王海波, 王莹莹, 王小龙, 王志强, 刘培培, 刘万春, 王孝娣. 中熟抗寒桃新品种‘中农甘爽’[J]. 园艺学报, 2022, 49(S2): 15-16. |
| [9] | 杨兴旺, 王海波, 王莹莹, 张艺灿, 王宝亮, 刘培培, 史祥宾, 刘万春, 王孝娣. 中熟抗寒桃新品种‘中农白干’[J]. 园艺学报, 2022, 49(S2): 17-18. |
| [10] | 杨兴旺, 刘凤之, 王海波, 王莹莹, 王志强, 史祥宾, 冀晓昊, 刘万春, 王孝娣. 中熟抗寒桃新品种‘中农寒水蜜’[J]. 园艺学报, 2022, 49(S2): 19-20. |
| [11] | 杨兴旺, 刘凤之, 王海波, 王莹莹, 张艺灿, 李 鹏, 王小龙, 刘万春, 王孝娣. 晚熟抗寒桃新品种‘中农秋香’[J]. 园艺学报, 2022, 49(S2): 21-22. |
| [12] | 王莹莹, 刘立常, 刘志伍, 杨兴旺, 刘万春, 王孝娣, . 极晚熟桃新品种‘中农冬蜜’[J]. 园艺学报, 2022, 49(S2): 23-24. |
| [13] | 王莹莹, 刘立常, 刘志伍, 杨兴旺, 刘万春, 王孝娣, . 小果油桃新品种‘中农珍珠’[J]. 园艺学报, 2022, 49(S2): 25-26. |
| [14] | 吴延军, 刘庆忠, 陈鸿才, 戚行江, 朱东姿, 郑家祥, 曹学敏, 方丹燕. 甜樱桃新品种‘江南锦’[J]. 园艺学报, 2022, 49(S2): 29-30. |
| [15] | 张晓明, 闫国华, 周 宇, 王 晶, 段续伟, 吴传宝, 张开春. 甜樱桃砧木新品种‘京春2号’[J]. 园艺学报, 2022, 49(S2): 31-32. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
版权所有 © 2012 《园艺学报》编辑部 京ICP备10030308号-2 国际联网备案号 11010802023439
编辑部地址: 北京市海淀区中关村南大街12号中国农业科学院蔬菜花卉研究所 邮编: 100081
电话: 010-82109523 E-Mail: yuanyixuebao@126.com
技术支持:北京玛格泰克科技发展有限公司