
Acta Horticulturae Sinica ›› 2021, Vol. 48 ›› Issue (3): 465-476.doi: 10.16420/j.issn.0513-353x.2020-0468
• Research Papers • Previous Articles Next Articles
ZHU Ziguo1, ZHANG Qingtian1, HAN Zhen1, LI Bo1, LI Guodong2, LI Xiujie1,*( )
)
Received:2020-11-24
Online:2021-03-25
Published:2021-04-02
Contact:
LI Xiujie
E-mail:lixiujie-2007@163.com
CLC Number:
ZHU Ziguo, ZHANG Qingtian, HAN Zhen, LI Bo, LI Guodong, LI Xiujie. VvMYB6,an R2R3-MYB Transcription Factor,is Involved in Anthocyanin Biosynthesis of Grapevine[J]. Acta Horticulturae Sinica, 2021, 48(3): 465-476.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2020-0468
| 基因名称 Gene name | 登录号 Accession number | 正向引物序列(5′-3′) Forward primer | 反向引物序列(3′-5′) Reverse primer | ||||
|---|---|---|---|---|---|---|---|
| VvMYB6 | MN125488 | CTTATCATCCGGCTTCGTTCCCT | TTTGCCCGTGTGCTTCAATCTCT | ||||
| VvActin | XP_002282516 | CCTCAACCCCAAGGCCAACAGA | ACCATCACCAGAATCCAGCACA | ||||
| NtCHI | X75963 | GTCAGGCCATTGAAAAGCTC | CTAATCGTCAATGCCCCAAC | ||||
| NtCHS | AB066274 | TTGTTCGAGCTTGTCTCTGC | AGCCCAGGAACATCTTTGAG | ||||
| NtFLS | AB086055 | GAACTTGAAGGGAAAAGGGG | TCCCTGTAGGAGGGAGGATT | ||||
| NtDFR | NM_001281215 | AACCAACAGTCAGGGGAATG | TTGGACATCGACAGTTCCAG | ||||
| NtANR1 | NM_001280956 | CATTTGACTTTCCCAAACGC | ATTGGGCTTTTGAGTTGTGC | ||||
| NtANR2 | XP_016512400 | TGTTCCCACTTGGGATGATA | TGCACCTATACTCTGTTAGTGGC | ||||
| NtANS | JQ866631 | TGGCGTTGAAGCTCATACTG | GGAATTAGGCACACACTTTGC | ||||
| NtUFGT | AF000371 | GAGTGCATTGGATGCCTTTT | CCAGCTCCATTAGGTCCTTG | ||||
| NtPAL | EF192469 | CCTTTATCCTACATTGCTGGTTT | TCGGGCTTTCCGTTCATTACTTC | ||||
| NtActin | XM_016618072 | AATGGAACTGGAATGGTCAAGGC | TGCCAGATCTTCTCCATGTCATCCCA | ||||
Table 1 Sequence of the primers in qPCR experiment
| 基因名称 Gene name | 登录号 Accession number | 正向引物序列(5′-3′) Forward primer | 反向引物序列(3′-5′) Reverse primer | ||||
|---|---|---|---|---|---|---|---|
| VvMYB6 | MN125488 | CTTATCATCCGGCTTCGTTCCCT | TTTGCCCGTGTGCTTCAATCTCT | ||||
| VvActin | XP_002282516 | CCTCAACCCCAAGGCCAACAGA | ACCATCACCAGAATCCAGCACA | ||||
| NtCHI | X75963 | GTCAGGCCATTGAAAAGCTC | CTAATCGTCAATGCCCCAAC | ||||
| NtCHS | AB066274 | TTGTTCGAGCTTGTCTCTGC | AGCCCAGGAACATCTTTGAG | ||||
| NtFLS | AB086055 | GAACTTGAAGGGAAAAGGGG | TCCCTGTAGGAGGGAGGATT | ||||
| NtDFR | NM_001281215 | AACCAACAGTCAGGGGAATG | TTGGACATCGACAGTTCCAG | ||||
| NtANR1 | NM_001280956 | CATTTGACTTTCCCAAACGC | ATTGGGCTTTTGAGTTGTGC | ||||
| NtANR2 | XP_016512400 | TGTTCCCACTTGGGATGATA | TGCACCTATACTCTGTTAGTGGC | ||||
| NtANS | JQ866631 | TGGCGTTGAAGCTCATACTG | GGAATTAGGCACACACTTTGC | ||||
| NtUFGT | AF000371 | GAGTGCATTGGATGCCTTTT | CCAGCTCCATTAGGTCCTTG | ||||
| NtPAL | EF192469 | CCTTTATCCTACATTGCTGGTTT | TCGGGCTTTCCGTTCATTACTTC | ||||
| NtActin | XM_016618072 | AATGGAACTGGAATGGTCAAGGC | TGCCAGATCTTCTCCATGTCATCCCA | ||||
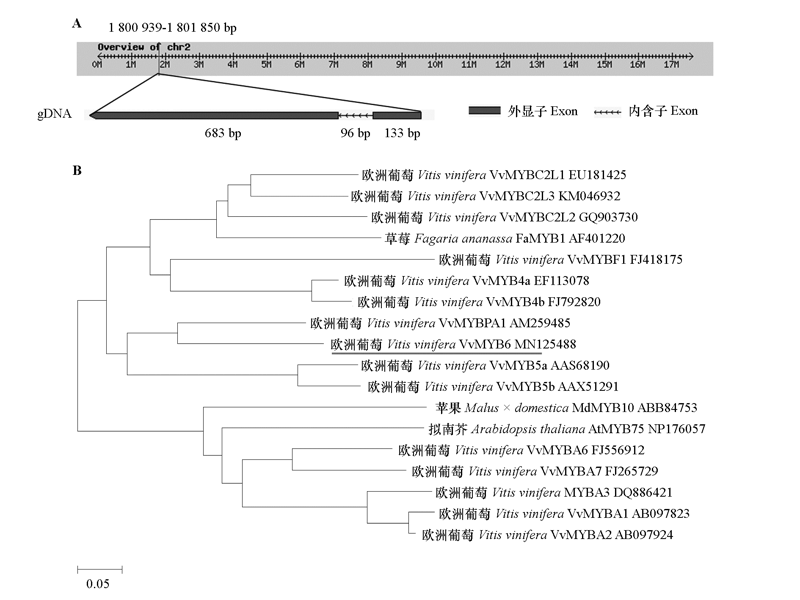
Fig. 1 The genomic sequence of VvMYB6 which is mapped to Vitis vinifera‘Pinot Noir’clone P40024 genome(A)and phylogenetic tree(B)of R2R3-MYB transcription factors from grapevine and other species
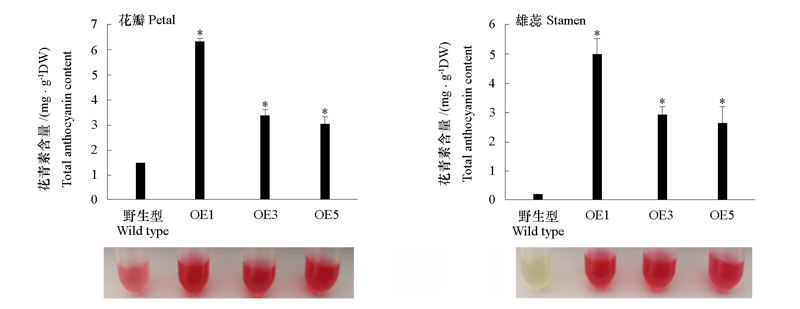
Fig. 8 Anthocyanin accumulation in VvMYB6 transgenic plants Significant difference from the wild type was confirmed by ANOVA and Tukey’s test. *α = 0.05.
| 编号 ID | 代谢物 Metabolites | 相对含量 Relative concentration | OE1/WT | |
|---|---|---|---|---|
| OE1 | WT | |||
| M466T139 | 飞燕草色素 Delphinidin | 36 999.82 ± 1 938.03 | 16 293.89 ± 822.94 | 2.27 |
| M488T142 | 飞燕草色素 Delphinidin | 7 152.78 ± 666.11 | 1 809.76 ± 235.23 | 3.95 |
| M552T143 | 飞燕草色素 Delphinidin | 2 705.68 ± 253.64 | 964.45 ± 107.88 | 2.81 |
| M597T60 | 飞燕草色素 Delphinidin | 6 892.49 ± 643.20 | 2 622.56 ± 352.73 | 2.63 |
| M604T134 | 矢车菊色素 Cyanidin | 25 610.01 ± 955.04 | 8 350.96 ± 311.87 | 3.07 |
| M612T139 | 矢车菊色素 Cyanidin | 121 588.30 ± 14 228.53 | 55 138.26 ± 3 964.54 | 2.21 |
| M628T137 | 飞燕草色素 Delphinidin | 8 934.53 ± 350.41 | 3 835.04 ± 346.59 | 2.33 |
| M637T55 | 矢车菊色素 Cyanidin | 625.84 ± 97.10 | 261.51 ± 34.42 | 2.39 |
| M774T133 | 飞燕草色素 Delphinidin | 15 256.59 ± 410.53 | 4 467.30 ± 148.95 | 3.42 |
| M790T151 | 飞燕草色素 Delphinidin | 1 220.32 ± 72.70 | 437.53 ± 62.86 | 2.79 |
| M796T133 | 飞燕草色素 Delphinidin | 10 307.65 ± 399.36 | 3 850.49 ± 247.85 | 2.68 |
Table 2 Relative concentration and fold-changes in the levels of major metabolites in flowers of transgenic OE1 line
| 编号 ID | 代谢物 Metabolites | 相对含量 Relative concentration | OE1/WT | |
|---|---|---|---|---|
| OE1 | WT | |||
| M466T139 | 飞燕草色素 Delphinidin | 36 999.82 ± 1 938.03 | 16 293.89 ± 822.94 | 2.27 |
| M488T142 | 飞燕草色素 Delphinidin | 7 152.78 ± 666.11 | 1 809.76 ± 235.23 | 3.95 |
| M552T143 | 飞燕草色素 Delphinidin | 2 705.68 ± 253.64 | 964.45 ± 107.88 | 2.81 |
| M597T60 | 飞燕草色素 Delphinidin | 6 892.49 ± 643.20 | 2 622.56 ± 352.73 | 2.63 |
| M604T134 | 矢车菊色素 Cyanidin | 25 610.01 ± 955.04 | 8 350.96 ± 311.87 | 3.07 |
| M612T139 | 矢车菊色素 Cyanidin | 121 588.30 ± 14 228.53 | 55 138.26 ± 3 964.54 | 2.21 |
| M628T137 | 飞燕草色素 Delphinidin | 8 934.53 ± 350.41 | 3 835.04 ± 346.59 | 2.33 |
| M637T55 | 矢车菊色素 Cyanidin | 625.84 ± 97.10 | 261.51 ± 34.42 | 2.39 |
| M774T133 | 飞燕草色素 Delphinidin | 15 256.59 ± 410.53 | 4 467.30 ± 148.95 | 3.42 |
| M790T151 | 飞燕草色素 Delphinidin | 1 220.32 ± 72.70 | 437.53 ± 62.86 | 2.79 |
| M796T133 | 飞燕草色素 Delphinidin | 10 307.65 ± 399.36 | 3 850.49 ± 247.85 | 2.68 |
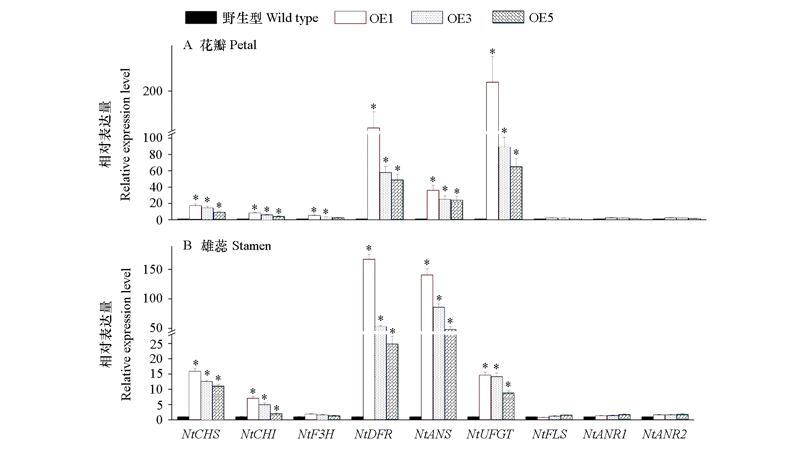
Fig. 9 Expression analysis of anthocyanin biosynthetic genes in petal(A)and stamen(B)from three VvMYB6 transgenic lines and wild-type lines Significant difference from the wild type was confirmed by ANOVA and Tukey’s test. *α = 0.05.
| [1] |
Aharoni A, De Vos C H, Wein M, Sun Z, Greco R, Kroon A, Mol J N, O'Connell A P. 2001. The strawberry FaMYB1 transcription factor suppresses anthocyanin and flavonol accumulation in transgenic tobacco. Plant J, 28:319-332.
pmid: 11722774 |
| [2] |
An J P, Wang X F, Zhang X W, Xu H F, Bi S Q, You C X, Hao Y J. 2020. An apple MYB transcription factor regulates cold tolerance and anthocyanin accumulation and undergoes MIEL1-mediated degradation. Plant Biotechnol J, 18:337-353.
doi: 10.1111/pbi.v18.2 URL |
| [3] |
Bogs J, Downey M O, Harvey J S, Ashton A R, Tanner G J, Robinson S P. 2005. Proanthocyanidin synthesis and expression of genes encoding leucoanthocyanidin reductase and anthocyanidin reductase in developing grape berries and grapevine leaves. Plant Physiology, 139:652-663.
doi: 10.1104/pp.105.064238 URL |
| [4] |
Bogs J, Jaffe F W, Takos A M, Walker A R, Robinson S P. 2007. The grapevine transcription factor VvMYBPA1 regulates proanthocyanidin synthesis during fruit development. Plant Physiology, 143:1347-1361.
doi: 10.1104/pp.106.093203 URL |
| [5] |
Cavallini E, Matus J T, Finezzo L, Zenoni S, Loyola R, Guzzo F, Schlechter R, Ageorges A, Arce-Johnson P, Tornielli G B. 2015. The phenylpropanoid pathway is controlled at different branches by a set of R2R3-MYB C 2 repressors in grapevine. Plant Physiology, 167 (4):1448-1470.
doi: 10.1104/pp.114.256172 pmid: 25659381 |
| [6] |
Chen K, Liu H, Lou Q, Liu Y. 2017. Ectopic expression of the grape Hyacinth Muscari armeniacum R2R3-MYB transcription factor gene, MaAN2, induces anthocyanin accumulation in tobacco. Front Plant Sci, 8:965.
doi: 10.3389/fpls.2017.00965 URL |
| [7] |
Cutanda-Perez M C, Ageorges A, Gomez C, Vialet S, Terrier N, Romieu C, Torregrosa L. 2009. Ectopic expression of VlmybA1 in grapevine activates a narrow set of genes involved in anthocyanin synthesis and transport. Plant Molecular Biology, 69:633-648.
doi: 10.1007/s11103-008-9446-x URL |
| [8] | Czemmel S, Heppel S C, Bogs J. 2012. R2R3 MYB transcription factors:key regulators of the flavonoid biosynthetic pathway in grapevine. Protoplasma, 249 (Suppl 2):S109-18. |
| [9] |
Czemmel S, Stracke R, Weisshaar B, Cordon N, Harris N N, Walker A R, Robinson S P, Bogs J. 2009. The grapevine R2R3-MYB transcription factor VvMYBF1 regulates flavonol synthesis in developing grape berries. Plant Physiology, 151:1513-1530.
doi: 10.1104/pp.109.142059 pmid: 19741049 |
| [10] |
Deluc L, Barrieu F, Marchive C, Lauvergeat V, Decendit A, Richard T, Carde J P, Merillon J M, Hamdi S. 2006. Characterization of a grapevine R2R3-MYB transcription factor that regulates the phenylpropanoid pathway. Plant Physiology, 140:499-511.
doi: 10.1104/pp.105.067231 URL |
| [11] |
Deluc L, Bogs J, Walker A R, Ferrier T, Decendit A, Merillon J M, Robinson S P, Barrieu F. 2008. The transcription factor VvMYB5b contributes to the regulation of anthocyanin and proanthocyanidin biosynthesis in developing grape berries. Plant Physiology, 147:2041-2053.
doi: 10.1104/pp.108.118919 URL |
| [12] |
Espley R V, Hellens R P, Putterill J, Stevenson D E, Kutty-Amma S, Allan A C. 2007. Red colouration in apple fruit is due to the activity of the MYB transcription factor,MdMYB10. Plant J, 49:414-427.
doi: 10.1111/tpj.2007.49.issue-3 URL |
| [13] |
Feng S, Wang Y, Yang S, Xu Y, Chen X. 2010. Anthocyanin biosynthesis in pears is regulated by a R2R3-MYB transcription factor PyMYB10. Planta, 232:245-255.
doi: 10.1007/s00425-010-1170-5 URL |
| [14] |
Fu Z Z, Shang H Q, Jiang H, Gao J, Dong X Y, Wang H J, Li Y M, Wang L M, Zhang J, Shu Q Y, Chao Y C, Xu M L, Wang R, Wang L S, Zhang H C. 2020. Systematic identification of the light-quality responding anthocyanin synthesis-related transcripts in Petunia petals. Horticultural Plant Journal, 6 (6):428-438.
doi: 10.1016/j.hpj.2020.11.006 URL |
| [15] |
Gonzalez A, Zhao M, Leavitt J M, Lloyd A M. 2008. Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J, 53:814-827.
doi: 10.1111/tpj.2008.53.issue-5 URL |
| [16] |
Hermanns A S, Zhou X S, Xu Q, Tadmor Y, Li L. 2020. Carotenoid pigment accumulation in horticultural plants. Horticultural Plant Journal, 6 (6):343-360.
doi: 10.1016/j.hpj.2020.10.002 URL |
| [17] | Hong Y, Li M, Dai S. 2019. Ectopic expression of multiple chrysanthemum(Chrysanthemum × morifolium)R2R3-MYB transcription factor genes regulates anthocyanin accumulation in tobacco. Genes,(10). DOI: 10.3390/genes10100777. |
| [18] |
Horsch R B, Fry J E, Hoffmann N L, Eichholtz D, Rogers S G, Farley R T. 1985. A simple and general method for transferring genes into plants. Science, 227:1229-1231.
doi: 10.1126/science.227.4691.1229 URL |
| [19] | Huang W, Khaldun A B, Chen J, Zhang C, Lv H, Yuan L, Wang Y. 2016. A R2R3-MYB transcription factor regulates the flavonol biosynthetic pathway in a traditional Chinese medicinal plant, Epimedium sagittatum. Front Plant Sci, 7:1089. |
| [20] |
Huang Z A, Zhao T, Fan H J, Wang N, Zheng S S, Ling H Q. 2012. The upregulation of NtAN2 expression at low temperature is required for anthocyanin accumulation in juvenile leaves of Lc-transgenic tobacco Nicotiana tabacum L. J Genet Genomics, 39:149-156.
doi: 10.1016/j.jgg.2012.01.007 URL |
| [21] |
Liu Y H, Zhang J L, Yu B, Wang J, Wang D. 2017. The MYB transcription factor StMYBA1 from potato requires light to activate anthocyanin biosynthesis in transgenic tobacco. J Plant Biol, 60:93-101.
doi: 10.1007/s12374-016-0199-9 URL |
| [22] |
Livak K J, Schmittgen T D. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2-Delta Delta CT Method. Methods, 25:402-408.
pmid: 11846609 |
| [23] | Man Yu-ping, Li Gang, Liu Hong, Wang Yan-chang, Qin Rui. 2012. Cloning and expression analysis of MVB in Actinidia chinensis‘Hongyang’. Journal of Huazhong Agricultural University, 31 (6):679-685. (in Chinese) |
| 满玉萍, 李刚, 刘虹, 王彦昌, 覃瑞. 2012. ‘红阳’猕猴桃MYB基因的克隆与表达. 华中农业大学学报, 31 (6):679-685. | |
| [24] |
Matsui K, Umemura Y, Ohme-Takagi M. 2008. AtMYBL2,a protein with a single MYB domain,acts as a negative regulator of anthocyanin biosynthesis in Arabidopsis. Plant J, 55:954-967.
doi: 10.1111/tpj.2008.55.issue-6 URL |
| [25] |
Matus J T, Cavallini E, Loyola R, Holl J, Finezzo L, Dal Santo S, Vialet S, Commisso M, Roman F, Schubert A, Alcalde J A, Bogs J, Ageorges A, Tornielli G B, Arce-Johnson P. 2017. A group of grapevine MYBA transcription factors located in chromosome 14 control anthocyanin synthesis in vegetative organs with different specificities compared with the berry color locus. Plant J, 91 (2):220-236.
doi: 10.1111/tpj.2017.91.issue-2 URL |
| [26] |
Meng Jiaxin, Gao Yan, Han Meiling, Liu Pengyuan, Yang Chen, Shen Ting, Houhua Li. 2020. In vitro anthocyanin induction and metabolite analysis in Malus spectabilis leaves under low nitrogen conditions. Horticultural Plant Journal, 6 (5):284-292.
doi: 10.1016/j.hpj.2020.06.004 URL |
| [27] |
Mehrtens F, Kranz H, Bednarek P, Weisshaar B. 2005. The Arabidopsis transcription factor MYB12 is a flavonol-specific regulator of phenylpropanoid biosynthesis. Plant Physiology, 138:1083-1096.
doi: 10.1104/pp.104.058032 URL |
| [28] |
Park J S, Kim J B, Cho K J, Cheon C I, Sung M K, Choung M G, Roh K H. 2008. Arabidopsis R2R3-MYB transcription factor AtMYB60 functions as a transcriptional repressor of anthocyanin biosynthesis in lettuce Lactuca sativa. Plant Cell Rep, 27:985-994.
doi: 10.1007/s00299-008-0521-1 URL |
| [29] |
Perez-Diaz J R, Perez-Diaz J, Madrid-Espinoza J, Gonzalez-Villanueva E, Moreno Y, Ruiz-Lara S. 2016. New member of the R2R3-MYB transcription factors family in grapevine suppresses the anthocyanin accumulation in the flowers of transgenic tobacco. Plant Molecular Biology, 90:63-76.
doi: 10.1007/s11103-015-0394-y URL |
| [30] |
Rinaldo A R, Cavallini E, Jia Y, Moss S M A, McDavid D A J, Hooper L C, Robinson S P, Tornielli G B, Zenoni S, Ford C M. 2015. A grapevine anthocyanin acyltransferase,transcriptionally regulated by VvMYBA,can produce most acylated anthocyanins present in grape skins. Plant Physiology, 169:1897-1916.
doi: 10.1104/pp.15.01255 pmid: 26395841 |
| [31] |
Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B. 2007. Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J, 50:660-677.
pmid: 17419845 |
| [32] |
Stracke R, Jahns O, Keck M, Tohge T, Niehaus K, Fernie A R, Weisshaar B. 2010. Analysis of PRODUCTION OF FLAVONOL GLYCOSIDES- dependent flavonol glycoside accumulation in Arabidopsis thaliana plants reveals MYB11-,MYB12-and MYB111-independent flavonol glycoside accumulation. New Phytol, 188:985-1000.
doi: 10.1111/nph.2010.188.issue-4 URL |
| [33] | Sun Shasha. 2016. The functional characterization of PyMYB10.1 regulating anthocyanin biosythesis in pear[M. D. Dissertation]. Tai'an:Shandong Agricultural University. (in Chinese) |
| 孙莎莎. 2016. 梨花青苷调控基因 PyMYB10.1功能鉴定[硕士论文]. 泰安:山东农业大学. | |
| [34] |
Teng S, Keurentjes J, Bentsink L, Koornneef M, Smeekens S. 2005. Sucrose-specific induction of anthocyanin biosynthesis in Arabidopsis requires the MYB75/PAP1 gene. Plant Physiology, 139:1840-1852.
doi: 10.1104/pp.105.066688 URL |
| [35] |
Walker A R, Lee E, Bogs J, McDavid D A, Thomas M R, Robinson S P. 2007. White grapes arose through the mutation of two similar and adjacent regulatory genes. Plant J, 49:772-785.
doi: 10.1111/tpj.2007.49.issue-5 URL |
| [36] |
Xi W, Feng J, Liu Y, Zhang S, Zhao G. 2019. The R2R3-MYB transcription factor PaMYB10 is involved in anthocyanin biosynthesis in apricots and determines red blushed skin. BMC Plant Biology, 19:287.
doi: 10.1186/s12870-019-1898-4 URL |
| [37] |
Xu Qing, He Jie, Dong Jianhui, Hou Xiaojin, Zhang Xian. 2018. Genomic survey and expression profiling of the MYB gene family in watermelon. Horticultural Plant Journal, 6 (5):284-292.
doi: 10.1016/j.hpj.2020.06.004 URL |
| [38] |
Yoo S D, Cho Y H, Sheen J. 2007. Arabidopsis mesophyll protoplasts:a versatile cell system for transient gene expression analysis. Nat Protoc, 2:1565-1572.
doi: 10.1038/nprot.2007.199 URL |
| [39] |
Zhang C M, Hao Y J. 2020. Advances in genomic,transcriptomic,and metabolomic analyses of fruit quality in fruit crops. Horticultural Plant Journal, 6 (6):361-371.
doi: 10.1016/j.hpj.2020.11.001 URL |
| [40] |
Zhu Z, Li G, Liu L, Zhang Q, Han Z, Chen X, Li B. 2019. A R2R3-MYB transcription factor, VvMYBC2L2,functions as a transcriptional repressor of anthocyanin biosynthesis in grapevine( Vitis vinifera L.). Molecules, 24 (92):2-13.
doi: 10.3390/molecules24010002 URL |
| [41] |
Zimmermann I M, Heim M A, Weisshaar B, Uhrig J F. 2004. Comprehensive identification of Arabidopsis thaliana MYB transcription factors interacting with R/B-like BHLH proteins. Plant J, 40:22-34.
pmid: 15361138 |
| [1] | YE Zimao, SHEN Wanxia, LIU Mengyu, WANG Tong, ZHANG Xiaonan, YU Xin, LIU Xiaofeng, and ZHAO Xiaochun, . Effect of R2R3-MYB Transcription Factor CitMYB21 on Flavonoids Biosynthesis in Citrus [J]. Acta Horticulturae Sinica, 2023, 50(2): 250-264. |
| [2] | ZOU Xue, DING Fan, LIU Lifang, YU Hankaizong, CHEN Nianwei, and RAO Liping. A New Purple Potato Cultivar‘Mianziyu 1’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 93-94. |
| [3] | WANG Sha, ZHANG Xinhui, ZHAO Yujie, LI Bianbian, ZHAO Xueqing, SHEN Yu, DONG Jianmei, YUAN Zhaohe. Cloning and Functional Analysis of PgMYB111 Related to Anthocyanin Synthesis in Pomegranate [J]. Acta Horticulturae Sinica, 2022, 49(9): 1883-1894. |
| [4] | HUANG Ling, HU Xianmei, LIANG Zehui, WANG Yanping, CHAN Zhulong, XIANG Lin. Cloning and Function Identification of Anthocyanidin Synthase Gene TgANS in Tulipa gesneriana [J]. Acta Horticulturae Sinica, 2022, 49(9): 1935-1944. |
| [5] | LI Maofu, YANG Yuan, WANG Hua, FAN Youwei, SUN Pei, JIN Wanmei. Analysis the Function of R2R3 MYB Transcription Factor RhMYB113c on Regulating Anthocyanin Synthesis in Rosa hybrida [J]. Acta Horticulturae Sinica, 2022, 49(9): 1957-1966. |
| [6] | XU Haifeng, WANG Zhongtang, CHEN Xin, LIU Zhiguo, WANG Lihu, LIU Ping, LIU Mengjun, ZHANG Qiong. The Analyses of Target Metabolomics in Flavonoid and Its Potential MYB Regulation Factors During Coloring Period of Winter Jujube [J]. Acta Horticulturae Sinica, 2022, 49(8): 1761-1771. |
| [7] | QIAN Jieyu, JIANG Lingli, ZHENG Gang, CHEN Jiahong, LAI Wuhao, XU Menghan, FU Jianxin, ZHANG Chao. Identification and Expression Analysis of MYB Transcription Factors Regulating the Anthocyanin Biosynthesis in Zinnia elegans and Function Research of ZeMYB9 [J]. Acta Horticulturae Sinica, 2022, 49(7): 1505-1518. |
| [8] | ZHANG Lugang, LU Qianqian, HE Qiong, XUE Yihua, MA Xiaomin, MA Shuai, NIE Shanshan, YANG Wenjing. Creation of Novel Germplasm of Purple-orange Heading Chinese Cabbage [J]. Acta Horticulturae Sinica, 2022, 49(7): 1582-1588. |
| [9] | WEI Xiaoyu, WANG Yuejin. Correlation Between Anatomical Structure and Resistance to Powdery Mildew in Chinese Wild Vitis Species [J]. Acta Horticulturae Sinica, 2022, 49(6): 1200-1212. |
| [10] | CHEN Daozong, LIU Yi, SHEN Wenjie, ZHU Bo, TAN Chen. Identification and Analysis of PAP1/2 Homologous Genes in Brassica rapa,B. oleracea and B. napus [J]. Acta Horticulturae Sinica, 2022, 49(6): 1301-1312. |
| [11] | LI Xiaoming, YU Junchi, WANG Chunxia. Comparison of Growth and Secondary Metabolites of Purple and White Flower Dracocephalum moldavica Under Field,Greenhouse and Greenhouse Shading Conditions [J]. Acta Horticulturae Sinica, 2022, 49(6): 1363-1370. |
| [12] | LI Lixian, WANG Shuo, CHEN Ying, WU Yingtao, WANG Yaqian, FANG Yue, CHEN Xuesen, TIAN Changping, FENG Shouqian. PavMYB10.1 Promotes Anthocyanin Accumulation by Positively Regulating PavRiant in Sweet Cherry [J]. Acta Horticulturae Sinica, 2022, 49(5): 1023-1030. |
| [13] | DENG Jiao, SU Mengyue, LIU Xuelian, OU Kefang, HU Zhengrong, YANG Pingfang. Transcriptome Analysis Revealed the Formation Mechanism of Floral Color of Lotus‘Dasajin’with Bicolor Petal [J]. Acta Horticulturae Sinica, 2022, 49(2): 365-377. |
| [14] | WANG Zhiyu, CHANG Beibei, LIU Qi, CHENG Xiaofan, DU Xiaoyun, YU Xiaoli, SONG Laiqing, ZHAO Lingling. Study on Expression and Anthocyanin Accumulation of Solute Carrier Gene MdSLC35F2-like in Apple [J]. Acta Horticulturae Sinica, 2022, 49(11): 2293-2303. |
| [15] | SUN Wei, SUN Shiyu, CHEN Yiran, WANG Yuhan, ZHANG Yan, JU Zhigang, YI Yin. Cloning and Function Analysis of Chalcone Isomerase Gene RdCHI1 in Rhododendron delavayi [J]. Acta Horticulturae Sinica, 2022, 49(11): 2407-2418. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd