
Acta Horticulturae Sinica ›› 2026, Vol. 53 ›› Issue (4): 1125-1142.doi: 10.16420/j.issn.0513-353x.2024-1054
• Genetic & Breeding · Germplasm Resources · Molecular Biology • Previous Articles Next Articles
BAI Xueying, TANG Manzhi, QU Tianhui, ZHANG Liting, XIAO Dong, LI Ying, HOU Xilin, LIU Tongkun*( )
)
Received:2025-11-23
Revised:2025-12-31
Online:2026-04-25
Published:2026-04-20
Contact:
LIU Tongkun
BAI Xueying, TANG Manzhi, QU Tianhui, ZHANG Liting, XIAO Dong, LI Ying, HOU Xilin, LIU Tongkun. Identification of FRL Gene Family,Expression Analysis and Functional Study of BrcFRI1 in Non-Heading Chinese Cabbage[J]. Acta Horticulturae Sinica, 2026, 53(4): 1125-1142.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2024-1054
| 基因 Gene | 正向引物序列(5′-3′) Forward primer | 反向引物序列(5′-3′) Reverse primer |
|---|---|---|
| ACTIN2 | GTTCCAGCCCTCGTTTGTG | CAAGTGCTGTGATTTCTTTGCTC |
| At-FRI | GGTGCTTTGAAGCGGTCACAGT | GCCCTCTCAAATGACTCCTTGC |
| At-FLC | TACAGCTTCTCCTCCGGCGATA | TCCCACAAGCTTGCTATCCACA |
| At-FT | GGTTGGTGACTGATATCCC | GGAAGGCCGAGATTGTATAG |
| Brc-Actin | CTCAGTCCAAAAGAGGTATTCT | GTAGAATGTGTGATGCCAGATC |
| BrcFRI1 | ATGGCCGTCCGTAATGGTTC | AAGACCGCATTGGAGGAATG |
| BrcFRI2 | GGCCTTTCGTAATGGTTCTC | CAAAATCAGGTCTCCTCGTA |
| BrcFRL1 | ATCACACCGCTCTCTCCTCT | ACCTAACTCAAACGCTGCGA |
| BrcFRL3-1 | GGATATGGACTCAGCAGGGC | GACAATCCAAGAGACCGGCA |
| BrcFRL3-2 | GCCTGGACAGCAACTCTCTT | GGAGTGGCTCAACTGGGTAC |
| BrcFRL3-3 | GTGGAAGGAACTGGAGGAGC | GGTGGTGGTGACGACTCAAT |
| BrcFRL4-1 | TCACCACGCTTGAGGAGAAC | AACGCTCGGAGTAACAAGCA |
| BrcFRL4-2 | AGGAAGTCGCGGTTGATTGT | GCTGGCTCTGGTTCTCTTGT |
| BrcFLC | CCGCAAGCCTATCTCTAACC | CAGAGGAGTTTACAGAAAGCAGA |
| BrcFT | ACGAGAGTCCAAGGCCCAAC | TGGCGCCATCCTGGTTCATA |
Table 1 Primer sequences used for qRT-PCR analysis in this study
| 基因 Gene | 正向引物序列(5′-3′) Forward primer | 反向引物序列(5′-3′) Reverse primer |
|---|---|---|
| ACTIN2 | GTTCCAGCCCTCGTTTGTG | CAAGTGCTGTGATTTCTTTGCTC |
| At-FRI | GGTGCTTTGAAGCGGTCACAGT | GCCCTCTCAAATGACTCCTTGC |
| At-FLC | TACAGCTTCTCCTCCGGCGATA | TCCCACAAGCTTGCTATCCACA |
| At-FT | GGTTGGTGACTGATATCCC | GGAAGGCCGAGATTGTATAG |
| Brc-Actin | CTCAGTCCAAAAGAGGTATTCT | GTAGAATGTGTGATGCCAGATC |
| BrcFRI1 | ATGGCCGTCCGTAATGGTTC | AAGACCGCATTGGAGGAATG |
| BrcFRI2 | GGCCTTTCGTAATGGTTCTC | CAAAATCAGGTCTCCTCGTA |
| BrcFRL1 | ATCACACCGCTCTCTCCTCT | ACCTAACTCAAACGCTGCGA |
| BrcFRL3-1 | GGATATGGACTCAGCAGGGC | GACAATCCAAGAGACCGGCA |
| BrcFRL3-2 | GCCTGGACAGCAACTCTCTT | GGAGTGGCTCAACTGGGTAC |
| BrcFRL3-3 | GTGGAAGGAACTGGAGGAGC | GGTGGTGGTGACGACTCAAT |
| BrcFRL4-1 | TCACCACGCTTGAGGAGAAC | AACGCTCGGAGTAACAAGCA |
| BrcFRL4-2 | AGGAAGTCGCGGTTGATTGT | GCTGGCTCTGGTTCTCTTGT |
| BrcFLC | CCGCAAGCCTATCTCTAACC | CAGAGGAGTTTACAGAAAGCAGA |
| BrcFT | ACGAGAGTCCAAGGCCCAAC | TGGCGCCATCCTGGTTCATA |
| 基因名称 Gene | 基因号 Gene number | CDS长度/bp CDS length | 氨基酸长度 Protein length | 分子量 Molecular weigh | 等电点 Isoelectric point | 亲水性 Gravy | 不稳定性指数 Instability Index |
|---|---|---|---|---|---|---|---|
| BrcFRI1 | BraC03g015970.1 | 1 791 | 596 | 66 693.58 | 7.64 | -0.48 | 70.28 |
| BrcFRI2 | BraC10g009270.1 | 1 710 | 569 | 63 390.88 | 9.14 | -0.42 | 67.68 |
| BrcFRL1 | BraC10g024850.1 | 1 374 | 457 | 51 253.90 | 7.62 | -0.50 | 52.23 |
| BrcFRL2 | BraC09g038630.1 | 1 386 | 461 | 51 418.23 | 9.04 | -0.32 | 58.87 |
| BrcFRL3-1 | BraC02g041830.1 | 1 659 | 552 | 61 499.29 | 5.77 | -0.41 | 52.56 |
| BrcFRL3-2 | BraC06g042450.1 | 1 140 | 379 | 41 594.13 | 6.42 | -0.21 | 60.23 |
| BrcFRL3-3 | BraC09g003460.1 | 1 563 | 520 | 58 366.95 | 7.14 | -0.56 | 58.92 |
| BrcFRL4-1 | BraC01g035560.1 | 1 605 | 534 | 59 035.71 | 6.53 | -0.44 | 57.25 |
| BrcFRL4-2 | BraC05g030410.1 | 1 380 | 459 | 51 512.32 | 6.08 | -0.40 | 52.40 |
| BrcFRL5-1 | BraC06g040730.1 | 2 595 | 864 | 96 375.21 | 7.73 | -0.43 | 39.23 |
| BrcFRL5-2 | BraC09g004180.1 | 2 523 | 840 | 95 082.19 | 6.86 | -0.48 | 37.41 |
| BrcFRL6 | BraC06g040740.1 | 3 471 | 1 156 | 131 268.90 | 5.65 | -0.59 | 47.73 |
| BrcFRL7 | BraC09g004160.1 | 3 420 | 1 139 | 130 221.60 | 5.21 | -0.57 | 49.36 |
Table 2 Physicochemical properties of non-heading Chinese cabbage FRL gene family members
| 基因名称 Gene | 基因号 Gene number | CDS长度/bp CDS length | 氨基酸长度 Protein length | 分子量 Molecular weigh | 等电点 Isoelectric point | 亲水性 Gravy | 不稳定性指数 Instability Index |
|---|---|---|---|---|---|---|---|
| BrcFRI1 | BraC03g015970.1 | 1 791 | 596 | 66 693.58 | 7.64 | -0.48 | 70.28 |
| BrcFRI2 | BraC10g009270.1 | 1 710 | 569 | 63 390.88 | 9.14 | -0.42 | 67.68 |
| BrcFRL1 | BraC10g024850.1 | 1 374 | 457 | 51 253.90 | 7.62 | -0.50 | 52.23 |
| BrcFRL2 | BraC09g038630.1 | 1 386 | 461 | 51 418.23 | 9.04 | -0.32 | 58.87 |
| BrcFRL3-1 | BraC02g041830.1 | 1 659 | 552 | 61 499.29 | 5.77 | -0.41 | 52.56 |
| BrcFRL3-2 | BraC06g042450.1 | 1 140 | 379 | 41 594.13 | 6.42 | -0.21 | 60.23 |
| BrcFRL3-3 | BraC09g003460.1 | 1 563 | 520 | 58 366.95 | 7.14 | -0.56 | 58.92 |
| BrcFRL4-1 | BraC01g035560.1 | 1 605 | 534 | 59 035.71 | 6.53 | -0.44 | 57.25 |
| BrcFRL4-2 | BraC05g030410.1 | 1 380 | 459 | 51 512.32 | 6.08 | -0.40 | 52.40 |
| BrcFRL5-1 | BraC06g040730.1 | 2 595 | 864 | 96 375.21 | 7.73 | -0.43 | 39.23 |
| BrcFRL5-2 | BraC09g004180.1 | 2 523 | 840 | 95 082.19 | 6.86 | -0.48 | 37.41 |
| BrcFRL6 | BraC06g040740.1 | 3 471 | 1 156 | 131 268.90 | 5.65 | -0.59 | 47.73 |
| BrcFRL7 | BraC09g004160.1 | 3 420 | 1 139 | 130 221.60 | 5.21 | -0.57 | 49.36 |
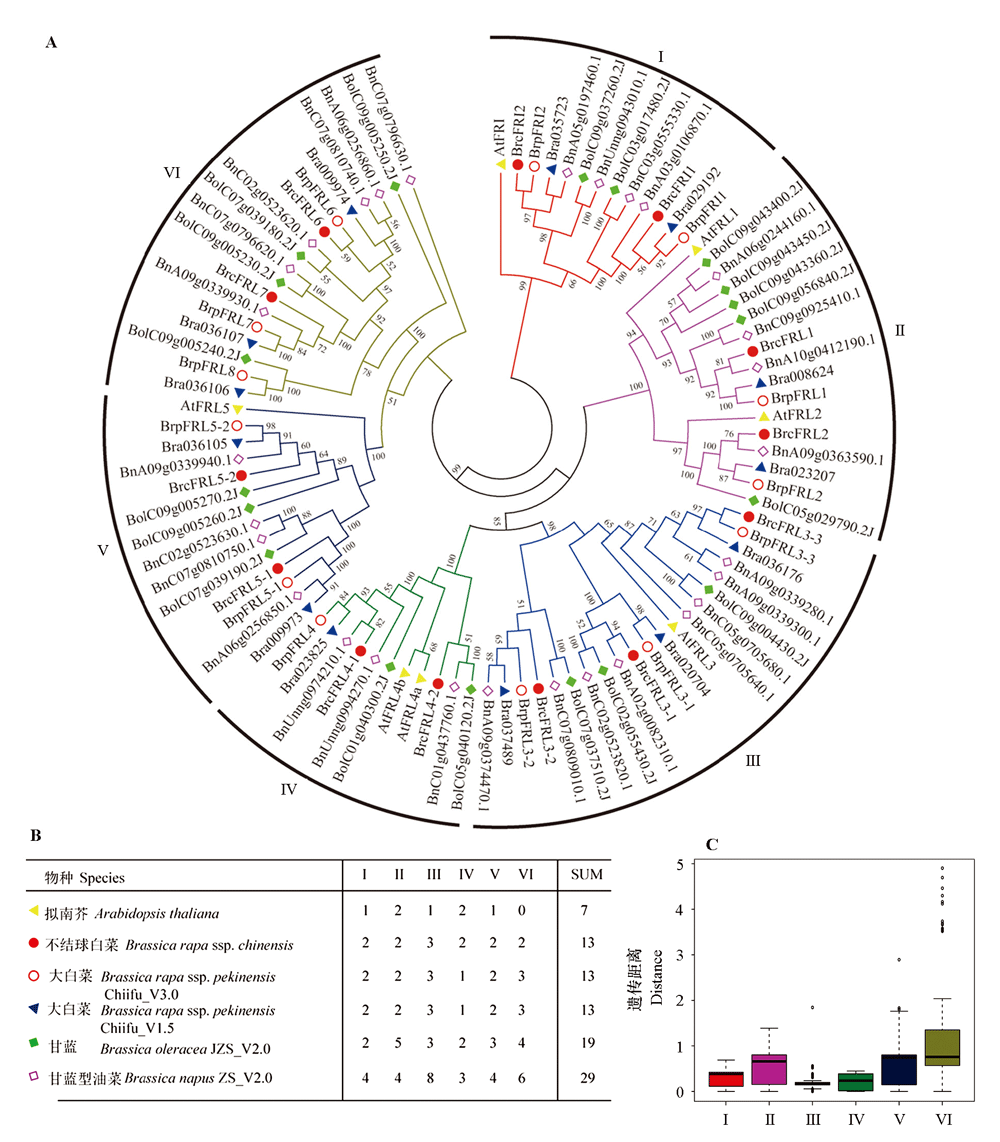
Fig. 1 Systematic evolution and diversity analysis of the FRL gene family in cruciferous species A:Phylogenetic tree of FRL proteins constructed based on the maximum likelihood method. The species include Arabidopsis thaliana,Brassica rapa ssp. chinensis,B. rapa ssp. pekinensis,B. oleracea,and B. napus;B:Distribution of FRL genes in different subgroups(Ⅰ- Ⅵ) among the five species;C:Analysis of nucleotide differences between different subgroups
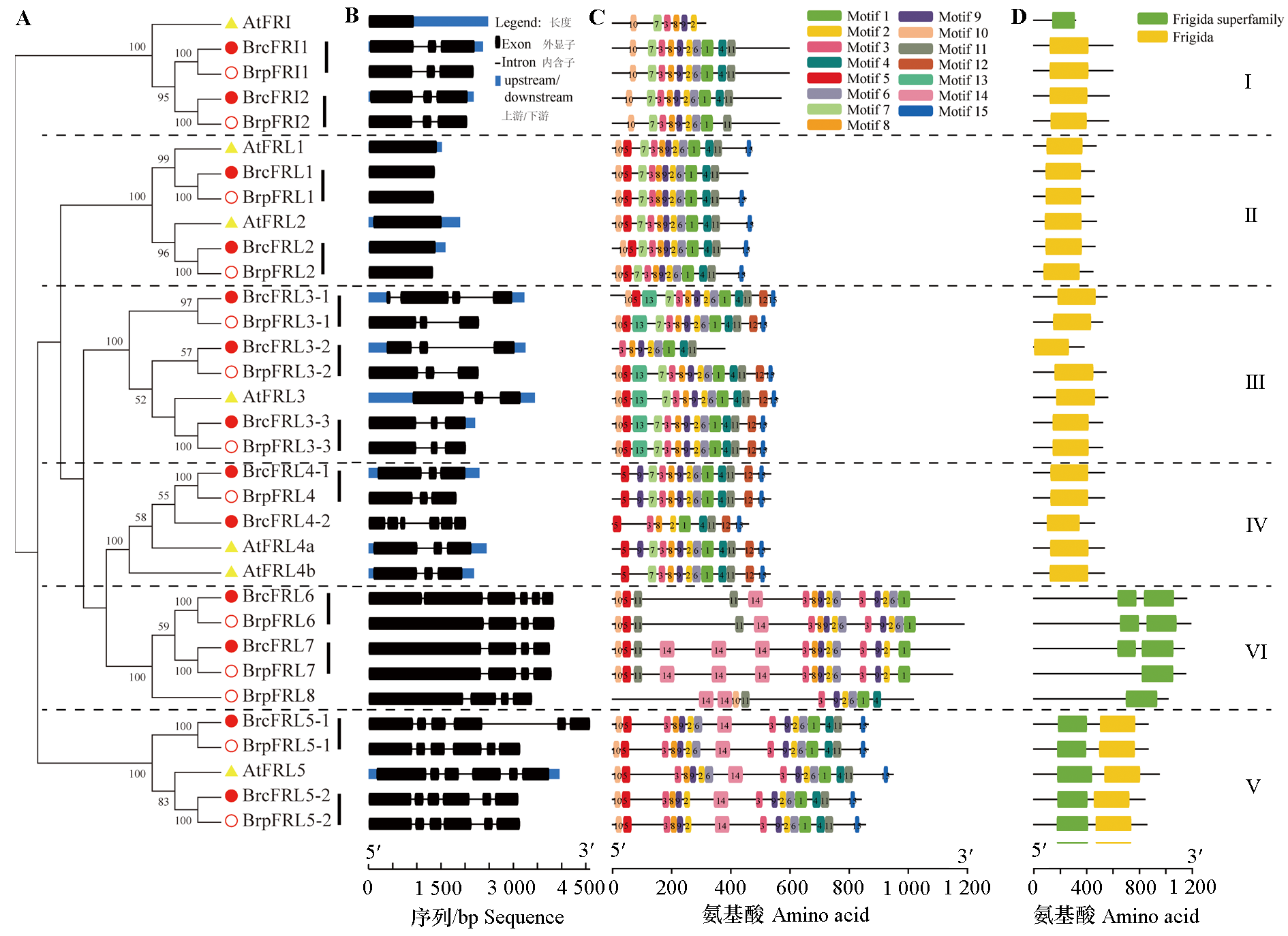
Fig. 4 Comprehensive analysis of the FRL gene family in Arabidopsis,non-heading Chinese cabbage,and Chinese cabbage A:Phylogenetic tree;B:Gene structure;C:Distribution of conserved motifs;D:Characteristics of the FRIGIDA domain composition
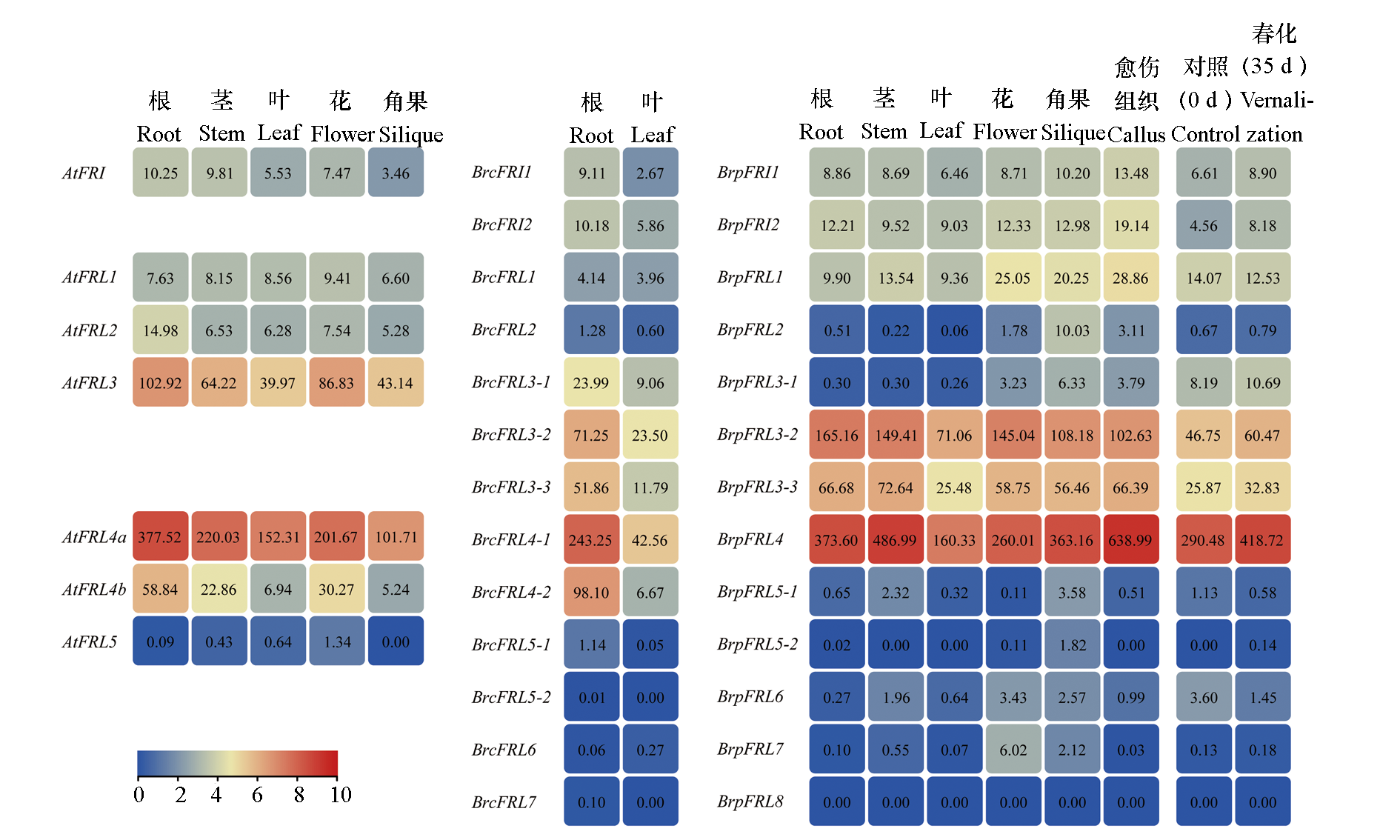
Fig. 6 Expression analysis of FRL genes in Arabidopsis thaliana,non-heading Chinese cabbage and Chinese cabbage in different tissues and before and after vernalization in Chinese cabbage
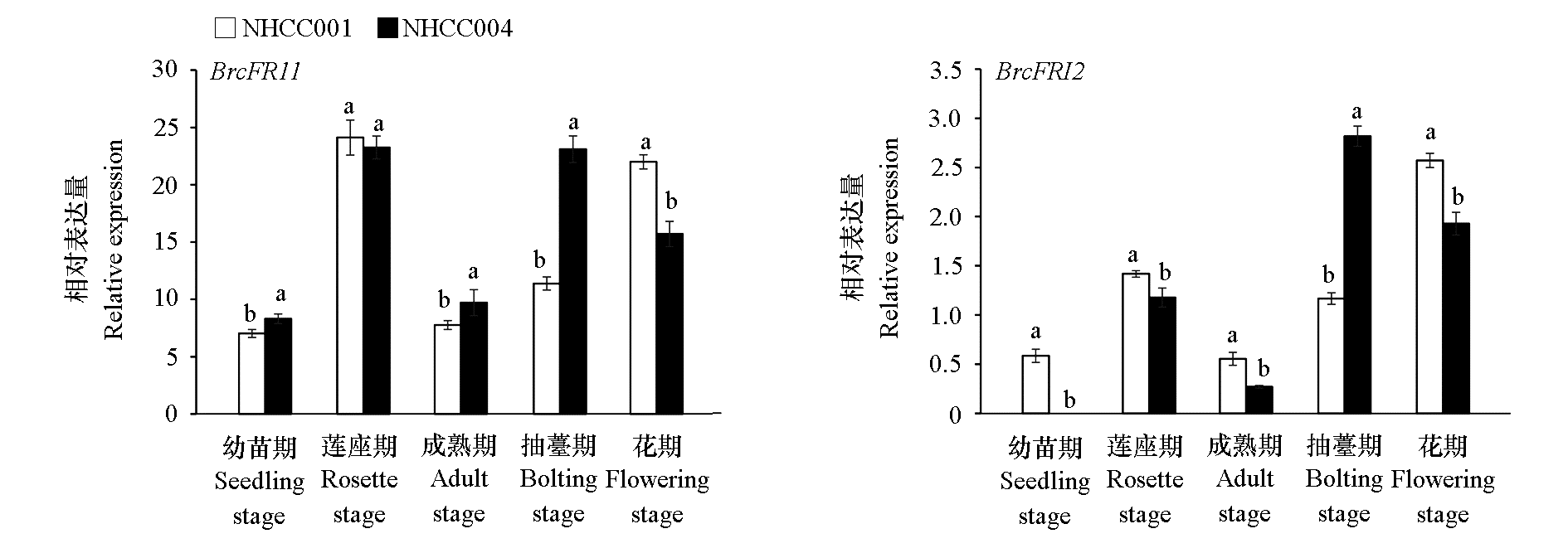
Fig. 7 Expression patterns of BrcFRI1 and BrcFRI2 at different developmental stages in early-and late-flowering non-heading cabbage varieties The different lowercase letters indicate significant differences(P < 0.05). The same below
| [1] |
doi: 10.1093/pcp/pcm176 pmid: 18156133 |
| [2] |
doi: 10.1007/s11033-012-2266-8 URL |
| [3] |
doi: 10.1016/j.molp.2020.06.009 URL |
| [4] |
doi: 10.1093/plphys/kiac264 pmid: 35670724 |
| [5] |
doi: 10.1016/j.bbrc.2017.11.149 URL |
| [6] |
|
|
段文优. 2020. 甘蓝型油菜开花基因BnaFRI的表达模式分析及突变体创建[硕士论文]. 武汉: 华中农业大学
|
|
| [7] |
|
|
高华北. 2020. 月季成花相关基因对环境变化的响应[硕士论文]. 北京: 北京林业大学.
|
|
| [8] |
doi: 10.1126/science.1068275 URL |
| [9] |
|
|
侯喜林, 李英, 刘同坤. 2022. 不结球白菜遗传育种与分子生物学研究进展. 南京农业大学学报, 45 (5):864-873.
|
|
| [10] |
|
|
胡思凡. 2018. 枳PtFRI基因在成花过程中的功能分析[硕士论文]. 武汉: 华中农业大学.
|
|
| [11] |
doi: 10.1105/tpc.114.132738 URL |
| [12] |
|
|
黄方, 罗聪, 余海霞, 范志毅, 谢小杰, 刘源, 莫啸, 何新华. 2021. 芒果MiFRI1和MiFRI2基因的克隆与表达分析. 热带作物学报, 42 (1):17-24.
doi: 10.3969/j.issn.1000-2561.2021.01.003 |
|
| [13] |
|
| [14] |
doi: 10.1105/tpc.106.045179 URL |
| [15] |
doi: 10.1093/jxb/erz287 URL |
| [16] |
doi: 10.1007/s13580-021-00357-8 |
| [17] |
|
|
李涵韬. 2021. 梅PmFRL3基因克隆及其与RGL2和ABI5蛋白互作分析[硕士论文]. 南京: 南京农业大学.
|
|
| [18] |
doi: 10.1016/j.hpj.2024.01.009 URL |
| [19] |
doi: 10.1105/tpc.11.5.949 pmid: 10330478 |
| [20] |
|
|
邱美琪. 2021. 柑橘CiFRI基因在干旱胁迫中的作用[硕士论文]. 武汉: 华中农业大学.
|
|
| [21] |
doi: 10.1007/s11103-010-9635-2 pmid: 20405310 |
| [22] |
doi: 10.1104/pp.105.061309 URL |
| [23] |
|
| [24] |
doi: 10.1073/pnas.0306401101 URL |
| [25] |
|
| [26] |
|
| [27] |
doi: 10.1038/s41598-019-44666-6 |
| [28] |
doi: 10.1038/ng.919 |
| [29] |
doi: 10.16420/j.issn.0513-353x.2023-0946 |
|
文宋琴, 李佳霖, 池卓恒, 夏燕, 王淑明, 吴頔, 郝雅雯, 郭启高, 梁国鲁, 景丹龙. 2024. 光和温度对开花时间基因可变剪接的影响研究进展. 园艺学报, 51 (8):1949-1963.
|
|
| [30] |
doi: 10.1111/tpj.v111.1 URL |
| [31] |
|
|
易丽聪. 2018. 甘蓝型油菜开花调控因子BnaA3.FRI的功能分析和基于染色体代换系的花期QTL定位[博士论文]. 武汉: 华中农业大学.
|
|
| [32] |
doi: 10.1038/s41598-023-45722-y |
| [33] |
|
|
翟江园. 2020. 蜡梅开花相关的FRIGIDA-LIKE基因家族成员功能的初步探究[硕士论文]. 武汉: 华中农业大学.
|
|
| [34] |
doi: 10.16420/j.issn.0513-353x.2025-0549 |
|
张昌伟, 厉恩铜, 王文龙, 侯喜林. 2026. 不结球白菜的起源、进化、研究及产业发展. 园艺学报, 53 (3):892-918.
doi: 10.16420/j.issn.0513-353x.2025-0549 |
|
| [35] |
|
| [1] | WANG Ronghua, LIU Shuantao, ZHAO Zhizhong, LI Qiaoyun, XU Nianfang, ZHANG Zhigang, WANG Shubin. Identification of the WSD Family Gene in Chinese Cabbage and Expression Analysis Under Drought Stress in Waxy Near-Isogenic Lines [J]. Acta Horticulturae Sinica, 2026, 53(4): 1143-1156. |
| [2] | DAI Yuru, SUN Pingping, WANG Qiong. Genome-Wide Identification of GAD Gene Family in Tomato and Expression Profiling after Ralstonia solanacearum Infection [J]. Acta Horticulturae Sinica, 2026, 53(4): 1157-1174. |
| [3] | REN Jiaxuan, ZHANG Fan, WANG Chenbing. Identification of the Peach SAP Gene Family and Alleviative Response in Rootstocks Under Abiotic Stress [J]. Acta Horticulturae Sinica, 2026, 53(3): 635-652. |
| [4] | CHEN Yushan, GAO Huan, WU Junkang, WANG Limei, HUANG Chunhui, XU Xiaobiao, LIAO Guanglian. Identification and Expression Analysis of AeBAM Gene Family in Actinidia eriantha [J]. Acta Horticulturae Sinica, 2026, 53(3): 683-696. |
| [5] | ZHANG Changwei, LI Entong, WANG Wenlong, HOU Xilin. The Origin,Evolution,Research,and Industrial Development of Non- Heading Chinese Cabbage [J]. Acta Horticulturae Sinica, 2026, 53(3): 892-918. |
| [6] | CHEN Yiquan, FAN Ronghui, LIN Bing, CHEN Yan, WU Jianshe, ZHONG Huaiqin. Screening and Validation of Reference Genes for qRT-PCR Analysis of Genes Related to Floral Scent Biosynthesis in Camellia japonica [J]. Acta Horticulturae Sinica, 2026, 53(2): 598-612. |
| [7] | ZHU Xuanyi, ZHAO Yuqing, TANG Feihong, WU Haiyan, ZHANG Chun, ZHAO Kai, PENG Donghui, LAN Siren, ZHOU Yuzhen. Whole-Genome Identification of the CNGC Gene Family in Cymbidium ensifolium and Transcriptional Reponses Under Abiotic Stresses [J]. Acta Horticulturae Sinica, 2026, 53(1): 131-148. |
| [8] | WANG Yanqiu, BAN Jingjie, ZENG Yuhan, LAI Chunwang, HUANG Xiaoling, ZHOU Jianjin, WAN Kui, LAI Zhongxiong, LIN Yuling. Identification of the DREB Gene Family in Polygonatum cyrtonema and Analysis of Its Response to Salt Stress [J]. Acta Horticulturae Sinica, 2026, 53(1): 159-173. |
| [9] | WU Jue, SHI Yinghong, TANG Yuying, WANG Xinman, ZHAN Xiuping. Analysis and Evaluation of Agronomic Traits and Nutritional Quality in Different Types of Non-Heading Chinese Cabbage for Autumn-Winter Cultivation in Shanghai [J]. Acta Horticulturae Sinica, 2026, 53(1): 219-228. |
| [10] | ZHU Caishuo, LI Jianzhong, ZHANG Qiannan, XU Lijie, ZHANG Shujiang, ZHANG Shifan, LI Fei, ZHANG Hui, XU Jin, SONG Zihan, LI Guoliang. Identification of the TCP Gene Family in Brassica campestris and Analysis of Their Effects on Axillary Bud Traits [J]. Acta Horticulturae Sinica, 2026, 53(1): 82-98. |
| [11] | LI Peirong, ZHAO Xiuyun, ZHANG Fenglan, YU Yangjun, ZHANG Deshuang, YU Shuancang, WANG Weihong, XIN Xiaoyun, ZHANG Bin, SU Tongbing. A New Heat-Resistant Non-Heading Chinese Cabbage Cultivar‘Guoxia 3’ [J]. Acta Horticulturae Sinica, 2025, 52(S2): 97-98. |
| [12] | ZHANG Wenqi, YANG Qinyu, YANG Xiao, ZHANG Li, HUANG Tao, OUYANG Lin, XU Hai, CHENG Longzheng, JING Zange, SONG Bo, YANG Qichang, LI Yuejian. A New Non-Heading Chinese Cabbage Cultivar‘Zhongshangqing 2’ [J]. Acta Horticulturae Sinica, 2025, 52(S2): 99-100. |
| [13] | LIU Zhaokun, WANG Huan, HAN Jianjun, WANG Yingying, ZHOU Hongzhang. A New Non-Heading Chinese Cabbage Cultivar‘Suqing 1’ [J]. Acta Horticulturae Sinica, 2025, 52(S2): 101-102. |
| [14] | ZHOU Jie, ZHANG Lei, WEI Lijun, WANG Qingming, WANG Yue’e, CAI Huaqing, LIU Daomin. A New Non-Heading Chinese Cabbage Cultivar‘Wan Cuilü’ [J]. Acta Horticulturae Sinica, 2025, 52(S2): 103-104. |
| [15] | LIU Chong, YI Dandan, ZOU Danrong, ZHOU Lixue, WANG Hui, SHENG Lili, CHEN Xiaofeng, ZHANG Yaming, GUI Hang. A New Non-Heading Chinese Cabbage Cultivar‘Qingmei 1’of High-Yield and High-Quality [J]. Acta Horticulturae Sinica, 2025, 52(S1): 87-88. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd