
Acta Horticulturae Sinica ›› 2026, Vol. 53 ›› Issue (1): 82-98.doi: 10.16420/j.issn.0513-353x.2025-0094
• Genetic & Breeding·Germplasm Resources·Molecular Biology • Previous Articles Next Articles
ZHU Caishuo1,2, LI Jianzhong2,3, ZHANG Qiannan2,4, XU Lijie2,5, ZHANG Shujiang2, ZHANG Shifan2, LI Fei2, ZHANG Hui2, XU Jin1, SONG Zihan2,**( ), LI Guoliang2,**(
), LI Guoliang2,**( )
)
Received:2025-04-17
Revised:2025-09-19
Online:2026-01-25
Published:2026-01-26
Contact:
SONG Zihan, LI Guoliang
ZHU Caishuo, LI Jianzhong, ZHANG Qiannan, XU Lijie, ZHANG Shujiang, ZHANG Shifan, LI Fei, ZHANG Hui, XU Jin, SONG Zihan, LI Guoliang. Identification of the TCP Gene Family in Brassica campestris and Analysis of Their Effects on Axillary Bud Traits[J]. Acta Horticulturae Sinica, 2026, 53(1): 82-98.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2025-0094
| 基因编号 Gene ID | 正向引物序列(5′-3′) Forward primer | 反向引物序列(5′-3′) Reverse primer |
|---|---|---|
| BraA01g009870.3C | GCTTCCTTCCCTCAACAACACGAAT | GGCTGTTCTCTCTCTAGCTCTCTCC |
| BraA01g034360.3C | ATTATTGCTCACAACCATTACTCCC | GATGGTCAAGATGATTGTTTGATGT |
| BraA01g036950.3C | ACCGTCCGAGCAAAGCCGTTGACTG | GGCGACGGAGAGATTTTAGTTTTGG |
| BraA02g002690.3C | CGCCGTCAAAAAACCACCGACCAAG | GTGGCGGCTATGATGGAAGGCTCGG |
| BraA02g015610.3C | CCCAAACAAGCAACAGGAAGTCGGG | TCCTCCGACCACGTCCCTCTACTTT |
| BraA02g017550.3C | TAATCCCTACAATCATCAGTCAACA | TCATAACCTTTCTGATTCACAGCAC |
| BraA02g018710.3C | TCTTTCACTACTCATATCCTAAGCC | CCAGTTCTTCTTGGTGAAACCTTTT |
| BraA02g019350.3C | TGGCTCATCCTCATCAGGTGCGTCC | ATCCAGCCGTTGATTGAACTAAATA |
| BraA02g020650.3C | CCGCTAGAGTTTTCCAGTTGACGAG | GCGTAAAGTAGCGTTTAAAGTCGAG |
| BraA02g042560.3C | CTGAGCCTTCCATCATAGCCGCAAC | GGTATAGCACCTCCGAAACCCTCTC |
| BraA02g042770.3C | CAGAATCAGCCAAGTTCAGTACCAA | TTTGCTGTGTCTGTCTTTGCCACCA |
| BraA03g003300.3C | CGATTCAGCTCTACGACCTTCAAGA | TTCAAACCTTCTCTCTGGATCGTAT |
| BraA03g003460.3C | CACAATGGGGGCTTCTTTGATGAGT | GCCAGAAGATTGAGATGATGACCCG |
| BraA03g023810.3C | CAATAGGACCGCCACTGAAGAGAGC | GGCGTTCTCCAACAACCACCGAATC |
| BraA03g030780.3C | CTTTCACTCTTTTCAACCATCATCA | CACCTCTCTCATTGTTGTTTTCTTC |
| BraA03g036760.3C | TTTCTCCATCTCCTCCGCCGCAGCA | TGGTTATTATCATTACTCCGACGTT |
| BraA03g038400.3C | AACTCCAACATATCAAGCAATCCTC | GTTCAAGGGAAGAAGTGATGGATGA |
| BraA03g043950.3C | AGGTTGTCCGTTCCAACAGCAATTC | GGAAATTGTAATGGAGGGAGCTTGT |
| BraA03g048240.3C | CTAACTGGTGCTTCTCCTACCGCTG | CTCTCCTTGGTCTCTTTAGCTGTTC |
| BraA04g013610.3C | AATCAAACCCGCCACCGACTCCGTT | ACCAAACCCAGTCTCATTTTCCCAA |
| BraA05g005380.3C | CCGACGAACCAGCCTCAGATGTTAG | CTCCTGTTTCTCGTAAATCTCCAGC |
| BraA05g013580.3C | ATCTTCTTCTTCCGAGCCCAACTTG | ACCGCCGCTTGTAGTTGTTGTCGCC |
| BraA05g032060.3C | GAAACCGAGAAGAACCGCCGCTAAA | CTCGTTCTCGTTCCCCCGATGCTCC |
| BraA05g041050.3C | AAAATTAATGGATGTGTAGAAGAGT | GTTGATGCTGTTCCATGTATTGATA |
| BraA06g019370.3C | ACTTCACTTCTCTCAACATCTCTCT | GTTGCTGTTGAAGTAGCGATGGAGA |
| BraA06g030710.3C | AAGCTCCACCACTATTTCCACTTCC | GGCGTAGCAAAAGATCCCATGTCGT |
| BraA06g037920.3C | CATCTTCAGGGTGGTGGGTCTCTCA | CTCCATCTCCCCCGCCAACACCCCA |
| BraA07g030260.3C | AGTCACATTCTTCTTCTTCTTCGCA | CGTATTATCACCACTGCTCCAAAAC |
| BraA07g031420.3C | TTTCTCGCCAATTCTTCGATCTCCA | CCATCGTTTCCAAAATCTTCAACAT |
| BraA07g032600.3C | TCAAGAAGGCGGTGAAGAAAGATAG | GTCTAATGTTTTACTGGCTTTGTCA |
| BraA07g034590.3C | CTCTCTCTCTGCTGCTCATCTTCGT | AGAAGAAGAACAAGAATCCGACTCA |
| BraA07g036490.3C | ATGTCGTTTCAGGGTAGTGGAATGT | GGATTTCTCTCTGCCGACATTATAG |
| BraA08g001700.3C | GGCTATTCAGTTCTACGATGTTCAA | TTTTGGGTCTTTTGGGTTTGTTTGC |
| BraA08g023460.3C | AGCCGTCGTTGAATCATAGTAATAG | GAGAGGTAGTGAATTGGACTGAAGG |
| BraA09g003050.3C | AAGACGACAATCATCATCACAACAG | CTTTGGTGTGTCTGTCTTTGTTTGA |
| BraA09g006660.3C | TTACAACAACAGCAGCAACAAGCAG | GCTCCACTAGATAAAGAAGCAAGCA |
| BraA09g018190.3C | CACCATCAGTACGAGGAGCAAGGAG | TGTTGATGATGTTGCTGTTGTTGAT |
| BraA09g034850.3C | TCTTCCTCTTCTCTACATCATTACA | GAGAATGTGAATTGGACTGAAGGGT |
| BraA10g030640.3C | GTCAAGCCGAGCCTTCTATCATAGC | TGTGTCCAACCGATAATTCACCACC |
Table 1 Primer sequences for genes related to the Brassica campestris TCP gene family
| 基因编号 Gene ID | 正向引物序列(5′-3′) Forward primer | 反向引物序列(5′-3′) Reverse primer |
|---|---|---|
| BraA01g009870.3C | GCTTCCTTCCCTCAACAACACGAAT | GGCTGTTCTCTCTCTAGCTCTCTCC |
| BraA01g034360.3C | ATTATTGCTCACAACCATTACTCCC | GATGGTCAAGATGATTGTTTGATGT |
| BraA01g036950.3C | ACCGTCCGAGCAAAGCCGTTGACTG | GGCGACGGAGAGATTTTAGTTTTGG |
| BraA02g002690.3C | CGCCGTCAAAAAACCACCGACCAAG | GTGGCGGCTATGATGGAAGGCTCGG |
| BraA02g015610.3C | CCCAAACAAGCAACAGGAAGTCGGG | TCCTCCGACCACGTCCCTCTACTTT |
| BraA02g017550.3C | TAATCCCTACAATCATCAGTCAACA | TCATAACCTTTCTGATTCACAGCAC |
| BraA02g018710.3C | TCTTTCACTACTCATATCCTAAGCC | CCAGTTCTTCTTGGTGAAACCTTTT |
| BraA02g019350.3C | TGGCTCATCCTCATCAGGTGCGTCC | ATCCAGCCGTTGATTGAACTAAATA |
| BraA02g020650.3C | CCGCTAGAGTTTTCCAGTTGACGAG | GCGTAAAGTAGCGTTTAAAGTCGAG |
| BraA02g042560.3C | CTGAGCCTTCCATCATAGCCGCAAC | GGTATAGCACCTCCGAAACCCTCTC |
| BraA02g042770.3C | CAGAATCAGCCAAGTTCAGTACCAA | TTTGCTGTGTCTGTCTTTGCCACCA |
| BraA03g003300.3C | CGATTCAGCTCTACGACCTTCAAGA | TTCAAACCTTCTCTCTGGATCGTAT |
| BraA03g003460.3C | CACAATGGGGGCTTCTTTGATGAGT | GCCAGAAGATTGAGATGATGACCCG |
| BraA03g023810.3C | CAATAGGACCGCCACTGAAGAGAGC | GGCGTTCTCCAACAACCACCGAATC |
| BraA03g030780.3C | CTTTCACTCTTTTCAACCATCATCA | CACCTCTCTCATTGTTGTTTTCTTC |
| BraA03g036760.3C | TTTCTCCATCTCCTCCGCCGCAGCA | TGGTTATTATCATTACTCCGACGTT |
| BraA03g038400.3C | AACTCCAACATATCAAGCAATCCTC | GTTCAAGGGAAGAAGTGATGGATGA |
| BraA03g043950.3C | AGGTTGTCCGTTCCAACAGCAATTC | GGAAATTGTAATGGAGGGAGCTTGT |
| BraA03g048240.3C | CTAACTGGTGCTTCTCCTACCGCTG | CTCTCCTTGGTCTCTTTAGCTGTTC |
| BraA04g013610.3C | AATCAAACCCGCCACCGACTCCGTT | ACCAAACCCAGTCTCATTTTCCCAA |
| BraA05g005380.3C | CCGACGAACCAGCCTCAGATGTTAG | CTCCTGTTTCTCGTAAATCTCCAGC |
| BraA05g013580.3C | ATCTTCTTCTTCCGAGCCCAACTTG | ACCGCCGCTTGTAGTTGTTGTCGCC |
| BraA05g032060.3C | GAAACCGAGAAGAACCGCCGCTAAA | CTCGTTCTCGTTCCCCCGATGCTCC |
| BraA05g041050.3C | AAAATTAATGGATGTGTAGAAGAGT | GTTGATGCTGTTCCATGTATTGATA |
| BraA06g019370.3C | ACTTCACTTCTCTCAACATCTCTCT | GTTGCTGTTGAAGTAGCGATGGAGA |
| BraA06g030710.3C | AAGCTCCACCACTATTTCCACTTCC | GGCGTAGCAAAAGATCCCATGTCGT |
| BraA06g037920.3C | CATCTTCAGGGTGGTGGGTCTCTCA | CTCCATCTCCCCCGCCAACACCCCA |
| BraA07g030260.3C | AGTCACATTCTTCTTCTTCTTCGCA | CGTATTATCACCACTGCTCCAAAAC |
| BraA07g031420.3C | TTTCTCGCCAATTCTTCGATCTCCA | CCATCGTTTCCAAAATCTTCAACAT |
| BraA07g032600.3C | TCAAGAAGGCGGTGAAGAAAGATAG | GTCTAATGTTTTACTGGCTTTGTCA |
| BraA07g034590.3C | CTCTCTCTCTGCTGCTCATCTTCGT | AGAAGAAGAACAAGAATCCGACTCA |
| BraA07g036490.3C | ATGTCGTTTCAGGGTAGTGGAATGT | GGATTTCTCTCTGCCGACATTATAG |
| BraA08g001700.3C | GGCTATTCAGTTCTACGATGTTCAA | TTTTGGGTCTTTTGGGTTTGTTTGC |
| BraA08g023460.3C | AGCCGTCGTTGAATCATAGTAATAG | GAGAGGTAGTGAATTGGACTGAAGG |
| BraA09g003050.3C | AAGACGACAATCATCATCACAACAG | CTTTGGTGTGTCTGTCTTTGTTTGA |
| BraA09g006660.3C | TTACAACAACAGCAGCAACAAGCAG | GCTCCACTAGATAAAGAAGCAAGCA |
| BraA09g018190.3C | CACCATCAGTACGAGGAGCAAGGAG | TGTTGATGATGTTGCTGTTGTTGAT |
| BraA09g034850.3C | TCTTCCTCTTCTCTACATCATTACA | GAGAATGTGAATTGGACTGAAGGGT |
| BraA10g030640.3C | GTCAAGCCGAGCCTTCTATCATAGC | TGTGTCCAACCGATAATTCACCACC |
| 基因编号 Gene ID | 分子量/kD Molecular weight | 等电点 Isoelectric point | 不稳定性系数 Instability coefficient | 平均亲水性系数 Hydrophilicity average coefficient |
|---|---|---|---|---|
| BraA01g009870.3C | 39 219.70 | 7.91 | 42.74 | -0.871 |
| BraA01g034360.3C | 47 318.52 | 6.77 | 49.63 | -0.867 |
| BraA01g036950.3C | 38 203.22 | 6.81 | 75.02 | -0.719 |
| BraA02g002690.3C | 23 842.65 | 9.57 | 49.44 | -0.388 |
| BraA02g015610.3C | 30 182.49 | 5.50 | 56.51 | -0.651 |
| BraA02g017550.3C | 39 308.67 | 5.50 | 48.92 | -0.823 |
| BraA02g018710.3C | 33 911.64 | 6.03 | 58.48 | -0.818 |
| BraA02g019350.3C | 33 870.94 | 7.12 | 59.49 | -0.696 |
| BraA02g020650.3C | 39 098.21 | 8.63 | 43.51 | -0.667 |
| BraA02g042560.3C | 18 600.96 | 7.92 | 54.69 | -0.162 |
| BraA02g042770.3C | 33 796.16 | 6.50 | 47.54 | -0.791 |
| BraA03g003300.3C | 27 367.45 | 6.49 | 51.36 | -0.518 |
| BraA03g003460.3C | 24 263.25 | 10.18 | 37.89 | -0.311 |
| BraA03g023810.3C | 33 174.90 | 9.67 | 55.70 | -0.237 |
| BraA03g030780.3C | 31 309.72 | 6.50 | 52.25 | -0.782 |
| BraA03g036760.3C | 44 499.00 | 7.49 | 67.30 | -0.804 |
| BraA03g038400.3C | 45 530.80 | 6.76 | 56.68 | -0.787 |
| BraA03g043950.3C | 40 698.22 | 6.18 | 56.00 | -0.904 |
| BraA03g048240.3C | 42 262.93 | 7.12 | 45.39 | -0.979 |
| BraA04g013610.3C | 24 724.61 | 7.20 | 36.43 | -0.845 |
| BraA05g005380.3C | 34 510.13 | 9.58 | 57.82 | -0.274 |
| BraA05g013580.3C | 38 727.41 | 6.34 | 50.44 | -0.870 |
| BraA05g032060.3C | 44 173.53 | 7.00 | 63.12 | -0.797 |
| BraA05g041050.3C | 34 451.32 | 7.21 | 56.70 | -0.706 |
| BraA06g019370.3C | 50 238.37 | 6.59 | 67.54 | -0.927 |
| BraA06g030710.3C | 26 991.12 | 9.69 | 57.09 | -0.589 |
| BraA06g037920.3C | 32 446.81 | 7.25 | 55.21 | -0.720 |
| BraA07g030260.3C | 33 733.76 | 7.43 | 56.69 | -0.715 |
| BraA07g031420.3C | 39 407.14 | 6.28 | 49.11 | -0.784 |
| BraA07g032600.3C | 38 800.34 | 6.36 | 57.19 | -0.875 |
| BraA07g034590.3C | 34 066.09 | 6.91 | 60.33 | -0.733 |
| BraA07g036490.3C | 37 206.26 | 6.84 | 45.51 | -0.605 |
| BraA08g001700.3C | 37 595.34 | 6.74 | 66.60 | -0.786 |
| BraA08g023460.3C | 34 410.52 | 6.61 | 64.29 | -0.921 |
| BraA09g003050.3C | 29 671.62 | 6.09 | 41.97 | -0.704 |
| BraA09g006660.3C | 23 116.85 | 9.38 | 44.70 | -0.491 |
| BraA09g018190.3C | 41 385.31 | 6.09 | 54.72 | -0.682 |
| BraA09g034850.3C | 35 327.53 | 7.80 | 51.70 | -0.929 |
| BraA10g030640.3C | 24 733.42 | 8.89 | 45.27 | -0.484 |
Table 2 Basic characterisation table of TCP gene families
| 基因编号 Gene ID | 分子量/kD Molecular weight | 等电点 Isoelectric point | 不稳定性系数 Instability coefficient | 平均亲水性系数 Hydrophilicity average coefficient |
|---|---|---|---|---|
| BraA01g009870.3C | 39 219.70 | 7.91 | 42.74 | -0.871 |
| BraA01g034360.3C | 47 318.52 | 6.77 | 49.63 | -0.867 |
| BraA01g036950.3C | 38 203.22 | 6.81 | 75.02 | -0.719 |
| BraA02g002690.3C | 23 842.65 | 9.57 | 49.44 | -0.388 |
| BraA02g015610.3C | 30 182.49 | 5.50 | 56.51 | -0.651 |
| BraA02g017550.3C | 39 308.67 | 5.50 | 48.92 | -0.823 |
| BraA02g018710.3C | 33 911.64 | 6.03 | 58.48 | -0.818 |
| BraA02g019350.3C | 33 870.94 | 7.12 | 59.49 | -0.696 |
| BraA02g020650.3C | 39 098.21 | 8.63 | 43.51 | -0.667 |
| BraA02g042560.3C | 18 600.96 | 7.92 | 54.69 | -0.162 |
| BraA02g042770.3C | 33 796.16 | 6.50 | 47.54 | -0.791 |
| BraA03g003300.3C | 27 367.45 | 6.49 | 51.36 | -0.518 |
| BraA03g003460.3C | 24 263.25 | 10.18 | 37.89 | -0.311 |
| BraA03g023810.3C | 33 174.90 | 9.67 | 55.70 | -0.237 |
| BraA03g030780.3C | 31 309.72 | 6.50 | 52.25 | -0.782 |
| BraA03g036760.3C | 44 499.00 | 7.49 | 67.30 | -0.804 |
| BraA03g038400.3C | 45 530.80 | 6.76 | 56.68 | -0.787 |
| BraA03g043950.3C | 40 698.22 | 6.18 | 56.00 | -0.904 |
| BraA03g048240.3C | 42 262.93 | 7.12 | 45.39 | -0.979 |
| BraA04g013610.3C | 24 724.61 | 7.20 | 36.43 | -0.845 |
| BraA05g005380.3C | 34 510.13 | 9.58 | 57.82 | -0.274 |
| BraA05g013580.3C | 38 727.41 | 6.34 | 50.44 | -0.870 |
| BraA05g032060.3C | 44 173.53 | 7.00 | 63.12 | -0.797 |
| BraA05g041050.3C | 34 451.32 | 7.21 | 56.70 | -0.706 |
| BraA06g019370.3C | 50 238.37 | 6.59 | 67.54 | -0.927 |
| BraA06g030710.3C | 26 991.12 | 9.69 | 57.09 | -0.589 |
| BraA06g037920.3C | 32 446.81 | 7.25 | 55.21 | -0.720 |
| BraA07g030260.3C | 33 733.76 | 7.43 | 56.69 | -0.715 |
| BraA07g031420.3C | 39 407.14 | 6.28 | 49.11 | -0.784 |
| BraA07g032600.3C | 38 800.34 | 6.36 | 57.19 | -0.875 |
| BraA07g034590.3C | 34 066.09 | 6.91 | 60.33 | -0.733 |
| BraA07g036490.3C | 37 206.26 | 6.84 | 45.51 | -0.605 |
| BraA08g001700.3C | 37 595.34 | 6.74 | 66.60 | -0.786 |
| BraA08g023460.3C | 34 410.52 | 6.61 | 64.29 | -0.921 |
| BraA09g003050.3C | 29 671.62 | 6.09 | 41.97 | -0.704 |
| BraA09g006660.3C | 23 116.85 | 9.38 | 44.70 | -0.491 |
| BraA09g018190.3C | 41 385.31 | 6.09 | 54.72 | -0.682 |
| BraA09g034850.3C | 35 327.53 | 7.80 | 51.70 | -0.929 |
| BraA10g030640.3C | 24 733.42 | 8.89 | 45.27 | -0.484 |
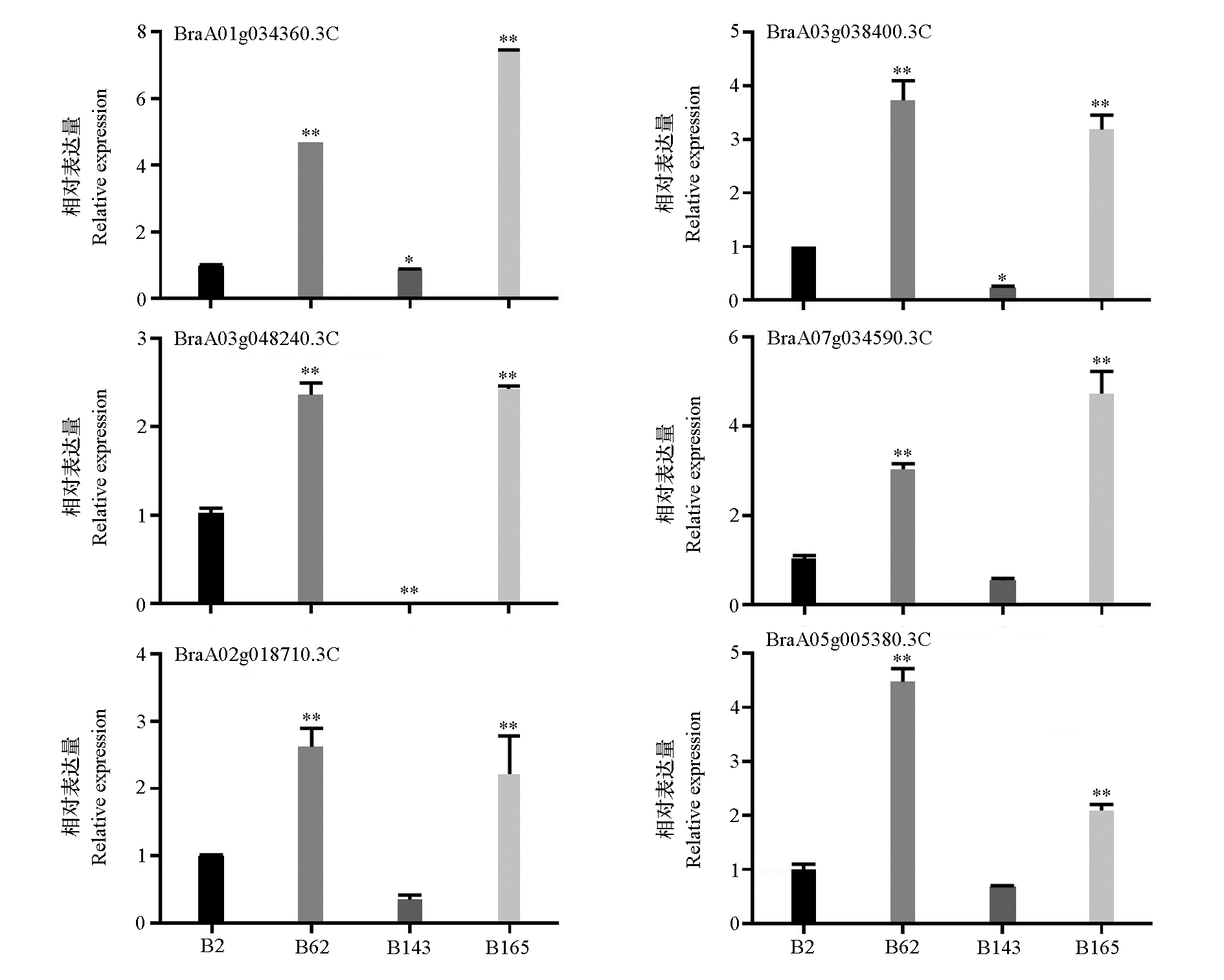
Analysis of different expression of TCP gene in various Brassica campestris materials Results were the mean of three independent biological replicates with quantitative real-time PCR(qRT-PCR)technology. Error bars represent the standard deviation of replicates. Using B2 as the control. Significance tests were performed using one-way analysis of variance(ANOVA). * indicates a significance level of 0.05,** indicates a significance level of 0.01
| [1] |
doi: 10.1105/tpc.106.048934 URL |
| [2] |
doi: 10.1104/pp.111.182725 pmid: 22045922 |
| [3] |
doi: 10.1007/s12298-017-0476-1 pmid: 29158628 |
| [4] |
doi: 10.1016/j.molp.2020.06.009 URL |
| [5] |
|
| [6] |
doi: 10.13560/j.cnki.biotech.bull.1985.2024-0115 |
|
陈智华, 乔振升, 李嘉其, 张晓琳, 马少杰, 何承忠, 纵丹. 2024. 滇杨TCP基因家族的全基因组鉴定与分析. 生物技术通报, 40 (11):214-226.
doi: 10.13560/j.cnki.biotech.bull.1985.2024-0115 |
|
| [7] |
|
| [8] |
doi: 10.16420/j.issn.0513-353x.2024-0791 |
|
董炫克, 李映旸, 马豫皖, 蔡艳芳, 丁启涵, 陈海霞, 李玉帆, 陈己任. 2025. 月季‘月月红’RcTCP9基因在盐和干旱胁迫中的功能分析. 园艺学报, 52 (7):1789-1802.
doi: 10.16420/j.issn.0513-353x.2024-0791 |
|
| [9] |
|
|
杜姗姗, 周绍礼, 赵宝林, 赵维月, 陈江华. 2022. 豌豆TCP基因家族的鉴定与分析. 植物生理学报, 58 (8):1575-1586.
|
|
| [10] |
|
|
段龙飞, 李文燕, 慕小倩, 张园园, 郭邦利, 陈国爱. 2016. 茉莉酸信号受体COI蛋白家族的分子进化与基因表达分析. 生物信息学, 14 (3):146-155.
|
|
| [11] |
|
|
冯献君, 吴英华, 史艳, 侯雷平, 李梅兰. 2022. 菜心酸性蔗糖转化酶基因家族的鉴定及表达分析. 江苏农业科学, 50 (19):44-50.
|
|
| [12] |
|
| [13] |
|
| [14] |
|
| [15] |
|
|
侯喜林, 李英, 黄菲艺. 2020. 不结球白菜(Brassica campestris ssp. chinensis)主要性状及育种技术的分子生物学研究新进展. 园艺学报, 47 (9):1663-1677.
|
|
| [16] |
|
|
黄依琳, 许玉富, 李荣华, 李光光, 张华, 郭培国, 夏岩石. 2023. 菜心群体中AUF2基因的遗传变异及其与农艺性状的关联分析. 广东农业科学, 50 (4):33-41.
|
|
| [17] |
doi: 10.1105/tpc.106.044792 URL |
| [18] |
|
|
李坤杰, 谭杉杉, 孙勃, 饶丹, 何琦, 闵爱玲, 李梦瑶. 2019. 芥菜TCP转录因子家族全基因组鉴定及表达分析. 四川农业大学学报, 37 (4):459-468.
|
|
| [19] |
doi: 10.1093/mp/ssr086 URL |
| [20] |
doi: 10.1186/s12870-022-03709-3 |
| [21] |
doi: 10.3390/ijms20143591 URL |
| [22] |
|
|
刘璇, 陈虞超, 郭生虎, 钟楠, 石磊, 甘晓燕. 2024. 芦竹TCP家族的基因鉴定与盐胁迫下的表达分析. 草业科学, 41 (10):2316-2329.
|
|
| [23] |
doi: 10.1006/meth.2001.1262 pmid: 11846609 |
| [24] |
doi: 10.1105/tpc.112.109090 pmid: 23613197 |
| [25] |
|
|
彭莹. 2014. 菜心(Brassica campestris L. ssp. chinensis var. utilis Tsen et Lee)Eru细胞质雄性不育系的研究与利用[硕士论文]. 武汉: 华中农业大学.
|
|
| [26] |
doi: 10.4161/psb.2.6.4811 pmid: 19704556 |
| [27] |
|
| [28] |
|
|
饶莉萍, 苏文瑾, 刘意, 宋天晓,
|
|
| [29] |
|
| [30] |
|
| [31] |
|
| [32] |
|
|
史楚然. 2022. 不结球白菜开花相关基因BcSPL15的功能分析及BcTCP家族鉴定[硕士论文]. 南京: 南京农业大学.
|
|
| [33] |
|
| [34] |
doi: 10.1016/j.cj.2021.01.007 URL |
| [35] |
|
|
孙启雪, 陈焯婷, 陈荣荣, 张兵. 2024. 狗牙根TCP转录因子基因家族鉴定与分析. 草业科学, 41 (10):2330-2344.
|
|
| [36] |
|
|
苏甜, 张应华, 吕霞, 彭梦玲, 倪雪梅, 许俊强. 2021. 植物侧枝发育的分子调控机理研究进展. 植物生理学报, 57 (8):1609-1616.
|
|
| [37] |
|
|
王崇, 兰孟焦, 肖满秋, 潘皓, 邓佳祺, 吴问胜. 2025. 甘薯PAL基因家族的全基因组鉴定及生物信息学分析. 植物遗传资源学报, 26 (6):1091-1105.
|
|
| [38] |
doi: 10.13345/j.cjb.230345 pmid: 38258643 |
|
王世泽, 李云, 韩玉翠, 余世洲, 王爽, 刘勇, 林小虎. 2024. 烟草TCP家族成员鉴定及表达分析. 生物工程学报, 40 (1):226-238.
pmid: 38258643 |
|
| [39] |
doi: 10.16420/j.issn.0513-353x.2025-0036 |
|
王文龙, 刘照坤, 鲁文君, 王耀龙, 李晓锋, 朱红芳, 刘同坤, 李英, 侯喜林, 张昌伟. 2025. 利用基因编辑技术提高不结球白菜抗坏血酸的含量. 园艺学报, 52 (5):1317-1325.
doi: 10.16420/j.issn.0513-353x.2025-0036 |
|
| [40] |
doi: 10.1038/ng.919 pmid: 21873998 |
| [41] |
|
|
吴栋雄. 2017. 白菜TCP基因家族的鉴定及利用CRISPR-Cas9系统进行编辑的初步探索[硕士论文]. 福建: 福建农林大学.
|
|
| [42] |
|
|
闫敏, 王晗, 许晔, 刘少华, 刘皓琦. 2024. 大白菜BrRopGEF基因家族全基因组鉴定及表达分析. 分子植物育种, 22 (22):7347-7354.
|
|
| [43] |
doi: 10.13345/j.cjb.220774 pmid: 36847088 |
|
阳文龙, 徐楚, 韩嘉琪, 张晓辉, 宋江萍, 贾会霞, 王海平. 2023. 芥菜WOX基因家族的全基因组鉴定与分析. 生物工程学报, 39 (2):537-551.
pmid: 36847088 |
|
| [44] |
|
| [45] |
|
| [46] |
|
|
张新, 李敏敏, 赵杨, 孙允超, 冀传允, 程倩倩, 张兰迎. 2024. 六倍体栽培小麦AP2/ERF基因家族的全基因组鉴定及表达分析. 江苏农业科学, 52 (16):58-68.
|
|
| [47] |
|
|
张莹, 李海俊, 高富成, 穆晓国, 高虎, 吴雪, 叶林, 曹凯. 2023. 菜心耐盐品种的筛选及其耐盐机理. 华北农学报, 38 (S1):170-179.
doi: 10.7668/hbnxb.20194042 |
|
| [48] |
|
|
郑玲, 白雪婷, 李会云. 2019. 高粱TCP基因家族全基因组鉴定及表达分析. 河南农业科学, 48 (10):30-36.
|
|
| [49] |
|
|
周而勋, 杨媚, 张华, 刘自珠. 2002. 菜心炭疽病菌菌丝生长、产孢和孢子萌发的影响因素. 南京农业大学学报, 25 (2):47-51.
|
|
| [50] |
doi: 10.3390/cimb45080401 URL |
| [1] | WANG Yanqiu, BAN Jingjie, ZENG Yuhan, LAI Chunwang, HUANG Xiaoling, ZHOU Jianjin, WAN Kui, LAI Zhongxiong, LIN Yuling. Identification of the DREB Gene Family in Polygonatum cyrtonema and Analysis of Its Response to Salt Stress [J]. Acta Horticulturae Sinica, 2026, 53(1): 159-173. |
| [2] | WANG Xinru, BAI Shuaishuai, SUN Yule, WU Xinyu, ZENG Feng, LIU Chunying, ZHANG Yuxi, GAI Shupeng, YUAN Yanchao. Genome-Wide Identification of PoSnRK Gene Family in Paeonia ostii and Expression Analysis During Flower Senescence [J]. Acta Horticulturae Sinica, 2025, 52(9): 2363-2376. |
| [3] | DONG Xuanke, LI Yingyang, MA Yuwan, CAI Yanfang, DING Qihan, CHEN Haixia, LI Yufan, and CHEN Jiren. The Functional Analysis of RcTCP9 Gene in Response to Salt and Drought Stress in Rosa chinensis‘Yueyuehong’ [J]. Acta Horticulturae Sinica, 2025, 52(7): 1789-1802. |
| [4] | YANG Chunmei, YU Yang, DING Yuge, XIA Jing, ZHOU Ling, and PENG Lei. Joint Transcription-Metabolism Analysis of Starch and Sucrose Metabolic Pathways in the Transformation of Axillary Buds into Flower Buds of ‘Guifei’Mango [J]. Acta Horticulturae Sinica, 2025, 52(6): 1412-1426. |
| [5] | CHEN Mindong, WANG Bin, BAI Changhui, QIU Boyin, LIU Jianting, WEN Qingfang, ZHU Haisheng, and LI Yongping. Identification of Expansin Family Gene and Its Expression Analysis During Fruit Enlargement in Sponge Gourd [J]. Acta Horticulturae Sinica, 2025, 52(6): 1488-1504. |
| [6] | SUN Tingzhen, SHU Qin, MA Wei, SHI Yuzi, ZHANG Meng, XIANG Chenggang, BO Kailiang, DUAN Ying, WANG Changlin. Identification and Phylogenetic Analysis of Gibberellin Oxidase Gene Family and Its Expression Pattern in Main Vine Growth in Cucurbita maxima [J]. Acta Horticulturae Sinica, 2025, 52(3): 603-622. |
| [7] | REN Xiliang, WANG Yuhong, GAO Tianyi, WANG Jie, HUANG Yunping. A New High-Quality Non-Heading Chinese Cabbage Cultivar‘Yongqing 8’ [J]. Acta Horticulturae Sinica, 2025, 52(2): 541-542. |
| [8] | HAN Hongwei, CHANG Yanan, LIU Huifang, ZHUANG Hongmei, XING Jiayi, ZHANG Xiaoda, ZULIPIYE · Abudumilike, WANG Hao, WANG Qiang. Identification of Cytochrome P450 Family Genes in Melon and Expression Analysis in Response to Light Intensity [J]. Acta Horticulturae Sinica, 2025, 52(12): 3271-3287. |
| [9] | SU Yanyan, WANG Yutong, JING Yanfu, XU Yaoguang, YU Yang, and XIE Hua. Identification of PpTRM Gene Family Members and Screening the Interacting PpTRMs with the Fruit Shape Regulator PpOFP1 in Peach [J]. Acta Horticulturae Sinica, 2025, 52(11): 2835-2845. |
| [10] | XIA Yan, HU Ruoqian, YI Guangquan, SUN Hongwen, HUANG Shuxin, CHI Zhuoheng, SHI Min, LIANG Guolu, JING Danlong, GUO Qigao. Characterization of Loquat bHLH Gene Family and Functional Analysis of EjbHLH104 on Flowering Regulation in Triploid Loquat [J]. Acta Horticulturae Sinica, 2025, 52(10): 2582-2596. |
| [11] | BIAN Shuanling, DENG Changrong, FAN Luze, WU Yanxun, REN Yanjing. Construction of Yeast Library and Identification of the BrrCDPK Gene Family of PSY Potential Interacting Proteins in Turnip [J]. Acta Horticulturae Sinica, 2025, 52(10): 2625-2641. |
| [12] | LIU Lu, WANG Meng, ZHENG Xiaomei, WANG Keqing, MIAO Ruyi, ZANG Qiaolu. Identification and Expression Analysis of CBF Gene Family of Artemisia argyi [J]. Acta Horticulturae Sinica, 2025, 52(10): 2703-2712. |
| [13] | WANG Shanshan, GUO Rui, HE Ling, WU Chunhong, CHEN Chanyou, WAN Heping, ZHAO Huixia. Identification of Long Cowpea Lhc Gene Family and Its Expression Analysis Under Salt Stress [J]. Acta Horticulturae Sinica, 2025, 52(1): 111-122. |
| [14] | LIU Jianhao, JING Yanfu, LIU Yuexin, XU Yaoguang, YU Yang, GE Xiuxiu, XIE Hua. Identification of Peach NAC Gene Family and Role of PpNAC050 in Promoting Fruit Fructose Accumulation [J]. Acta Horticulturae Sinica, 2024, 51(9): 1983-1996. |
| [15] | ZHAO Jiaying, ZENG Zhouting, CEN Xinying, SHI Jiaoqi, LI Xiaoxian, SHEN Xiaoxia, YU Zhenming. Genome-Wide Identification and Expression Analysis of CCO Gene Family in Dendrobium officinale During Flower Development [J]. Acta Horticulturae Sinica, 2024, 51(9): 2075-2088. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd