
Acta Horticulturae Sinica ›› 2026, Vol. 53 ›› Issue (3): 683-696.doi: 10.16420/j.issn.0513-353x.2025-0310
• Genetic & Breeding·Germplasm Resources·Molecular Biology • Previous Articles Next Articles
CHEN Yushan1,2, GAO Huan1,2, WU Junkang1,2, WANG Limei1,2, HUANG Chunhui1,2, XU Xiaobiao1,2,**( ), LIAO Guanglian1,2,**(
), LIAO Guanglian1,2,**( )
)
Received:2025-07-02
Revised:2025-09-28
Online:2026-03-25
Published:2026-03-20
Contact:
XU Xiaobiao, LIAO Guanglian
CHEN Yushan, GAO Huan, WU Junkang, WANG Limei, HUANG Chunhui, XU Xiaobiao, LIAO Guanglian. Identification and Expression Analysis of AeBAM Gene Family in Actinidia eriantha[J]. Acta Horticulturae Sinica, 2026, 53(3): 683-696.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2025-0310
| 基因 Gene | 上游引物序列(5′-3′) Forward primer sequences | 下游引物序列(5′-3′) Revers primer sequences | 产物大小/bp Product |
|---|---|---|---|
| AeBAM1 | GAATGGCTACAAGAGGCT | TATGTCGGGATTGCTACG | 157 |
| AeBAM2 | GGAGGAAGGCTATGTGGG | TCCGTGTAGGCAAGGTCA | 283 |
| AeBAM3 | TTTGTGCCCACTTTATCC | CAACCTCTTGTAGCCATT | 384 |
| AeBAM4 | TACGCTATCCCTCTTATCC | ATTTCTGCCGCTTTCTTC | 109 |
| AeBAM5 | TTTCCTTGGGTTGTGATT | GTCTTTCTTTCCGATTGC | 281 |
| AeBAM6 | AGGGATGTTGCTTCTTCA | GGTTCATCTGCTGCTCATT | 266 |
| AeBAM7 | CCCGATGGCACTACCTAT | GCACCCTGGAACCAACAT | 298 |
| AeBAM8 | GATTCACTGGCACTACGGG | GGGCATCCTGAGGCTGTT | 173 |
| AeBAM9 | GCCGATTGATGGAGTAAG | GATGCGTGGAAGCAGAGC | 264 |
| AeBAM10 | GGCAAGGTGGTGTCAGTC | CATCCGATTTCCAGTTTT | 103 |
| AeBAM11 | AGAAGGGAAGCCTCAGTG | GTTAGTTCAGCAGGGTGT | 297 |
| AeBAM12 | TCAAGGATTGGCTGTGGC | GATGACGGCTGTTGATGC | 141 |
| AeBAM13 | AGTGCGGAGGAAATGTTGG | TCTAAGCACTGGTAACGAAT | 158 |
| AeBAM14 | GTGCGGAGGAAATGTTGG | AAGGATAGGTATTCGGCATT | 129 |
| AeBAM15 | AGTGCGGAGGAAATGTTGG | GCCAAGGATAGGTATTCGGG | 133 |
| AeBAM16 | TGGAACAATGATTACGGACAA | GTAGTGCCAATGAATCCCTG | 150 |
| AeBAM17 | ATCTGATTCCAGGGTGCC | CATTCATTGTTCGTGGCTTG | 89 |
Table 1 qRT-PCR primer sequences
| 基因 Gene | 上游引物序列(5′-3′) Forward primer sequences | 下游引物序列(5′-3′) Revers primer sequences | 产物大小/bp Product |
|---|---|---|---|
| AeBAM1 | GAATGGCTACAAGAGGCT | TATGTCGGGATTGCTACG | 157 |
| AeBAM2 | GGAGGAAGGCTATGTGGG | TCCGTGTAGGCAAGGTCA | 283 |
| AeBAM3 | TTTGTGCCCACTTTATCC | CAACCTCTTGTAGCCATT | 384 |
| AeBAM4 | TACGCTATCCCTCTTATCC | ATTTCTGCCGCTTTCTTC | 109 |
| AeBAM5 | TTTCCTTGGGTTGTGATT | GTCTTTCTTTCCGATTGC | 281 |
| AeBAM6 | AGGGATGTTGCTTCTTCA | GGTTCATCTGCTGCTCATT | 266 |
| AeBAM7 | CCCGATGGCACTACCTAT | GCACCCTGGAACCAACAT | 298 |
| AeBAM8 | GATTCACTGGCACTACGGG | GGGCATCCTGAGGCTGTT | 173 |
| AeBAM9 | GCCGATTGATGGAGTAAG | GATGCGTGGAAGCAGAGC | 264 |
| AeBAM10 | GGCAAGGTGGTGTCAGTC | CATCCGATTTCCAGTTTT | 103 |
| AeBAM11 | AGAAGGGAAGCCTCAGTG | GTTAGTTCAGCAGGGTGT | 297 |
| AeBAM12 | TCAAGGATTGGCTGTGGC | GATGACGGCTGTTGATGC | 141 |
| AeBAM13 | AGTGCGGAGGAAATGTTGG | TCTAAGCACTGGTAACGAAT | 158 |
| AeBAM14 | GTGCGGAGGAAATGTTGG | AAGGATAGGTATTCGGCATT | 129 |
| AeBAM15 | AGTGCGGAGGAAATGTTGG | GCCAAGGATAGGTATTCGGG | 133 |
| AeBAM16 | TGGAACAATGATTACGGACAA | GTAGTGCCAATGAATCCCTG | 150 |
| AeBAM17 | ATCTGATTCCAGGGTGCC | CATTCATTGTTCGTGGCTTG | 89 |
| 基因名 Gene name | 基因ID Gene ID | 氨基酸数 No. of amino acid | 分子量/Da Molecular weight | 等电点 pI | 稳定系数 Instability index | 脂溶指数 Aliphatic index | 亲水性 Hydrop- athicity | 已报道同源基因 Reported homologous genes |
|---|---|---|---|---|---|---|---|---|
| AeBAM1 | Chr02G799 | 751 | 84 142.08 | 5.82 | 34.45 | 77.75 | -0.35 | / |
| AeBAM2 | Chr02G1164 | 568 | 63 467.87 | 5.57 | 40.59 | 68.89 | -0.45 | / |
| AeBAM3 | Chr06G622 | 700 | 78 501.26 | 5.24 | 38.18 | 73.40 | -0.46 | / |
| AeBAM4 | Chr06G623 | 556 | 63 133.23 | 5.65 | 37.67 | 74.69 | -0.39 | / |
| AeBAM5 | Chr11G331 | 480 | 54 004.19 | 8.29 | 33.44 | 67.25 | -0.53 | AdBAM3L(Acc28966); AcBAM5(Actinidia10870.t1) |
| AeBAM6 | Chr11G561 | 541 | 60 915.04 | 6.65 | 37.90 | 67.80 | -0.31 | / |
| AeBAM7 | Chr11G638 | 666 | 74 694.92 | 5.89 | 39.40 | 70.33 | -0.45 | / |
| AeBAM8 | Chr13G233 | 564 | 62 786.18 | 5.90 | 45.97 | 70.27 | -0.42 | / |
| AeBAM9 | Chr16G812 | 532 | 58 600.29 | 6.18 | 28.35 | 81.73 | -0.26 | AcBAM13(Actinidia00051.t1) |
| AeBAM10 | Chr16G1012 | 189 | 21 149.85 | 10.01 | 44.29 | 111.43 | 0.17 | / |
| AeBAM11 | Chr19G603 | 523 | 59 697.34 | 8.93 | 51.50 | 80.38 | -0.30 | / |
| AeBAM12 | Chr20G1070 | 645 | 72 789.04 | 6.20 | 41.02 | 71.21 | -0.48 | / |
| AeBAM13 | Chr22G622 | 547 | 61 739.99 | 8.62 | 38.39 | 66.54 | -0.53 | AdBAM3.1(Acc15874) |
| AeBAM14 | Chr23G1204 | 501 | 56 841.28 | 8.24 | 37.43 | 67.19 | -0.58 | / |
| AeBAM15 | Chr23G1205 | 301 | 34 345.80 | 8.96 | 40.58 | 82.23 | -0.34 | / |
| AeBAM16 | Chr23G1206 | 501 | 56 428.79 | 8.00 | 37.67 | 66.99 | -0.56 | / |
| AeBAM17 | Chr25G384 | 546 | 61 462.78 | 8.58 | 36.62 | 70.57 | -0.51 | AdBAM3(Acc16695) |
Table 2 Protein physical and chemical properties analysis of AeBAM
| 基因名 Gene name | 基因ID Gene ID | 氨基酸数 No. of amino acid | 分子量/Da Molecular weight | 等电点 pI | 稳定系数 Instability index | 脂溶指数 Aliphatic index | 亲水性 Hydrop- athicity | 已报道同源基因 Reported homologous genes |
|---|---|---|---|---|---|---|---|---|
| AeBAM1 | Chr02G799 | 751 | 84 142.08 | 5.82 | 34.45 | 77.75 | -0.35 | / |
| AeBAM2 | Chr02G1164 | 568 | 63 467.87 | 5.57 | 40.59 | 68.89 | -0.45 | / |
| AeBAM3 | Chr06G622 | 700 | 78 501.26 | 5.24 | 38.18 | 73.40 | -0.46 | / |
| AeBAM4 | Chr06G623 | 556 | 63 133.23 | 5.65 | 37.67 | 74.69 | -0.39 | / |
| AeBAM5 | Chr11G331 | 480 | 54 004.19 | 8.29 | 33.44 | 67.25 | -0.53 | AdBAM3L(Acc28966); AcBAM5(Actinidia10870.t1) |
| AeBAM6 | Chr11G561 | 541 | 60 915.04 | 6.65 | 37.90 | 67.80 | -0.31 | / |
| AeBAM7 | Chr11G638 | 666 | 74 694.92 | 5.89 | 39.40 | 70.33 | -0.45 | / |
| AeBAM8 | Chr13G233 | 564 | 62 786.18 | 5.90 | 45.97 | 70.27 | -0.42 | / |
| AeBAM9 | Chr16G812 | 532 | 58 600.29 | 6.18 | 28.35 | 81.73 | -0.26 | AcBAM13(Actinidia00051.t1) |
| AeBAM10 | Chr16G1012 | 189 | 21 149.85 | 10.01 | 44.29 | 111.43 | 0.17 | / |
| AeBAM11 | Chr19G603 | 523 | 59 697.34 | 8.93 | 51.50 | 80.38 | -0.30 | / |
| AeBAM12 | Chr20G1070 | 645 | 72 789.04 | 6.20 | 41.02 | 71.21 | -0.48 | / |
| AeBAM13 | Chr22G622 | 547 | 61 739.99 | 8.62 | 38.39 | 66.54 | -0.53 | AdBAM3.1(Acc15874) |
| AeBAM14 | Chr23G1204 | 501 | 56 841.28 | 8.24 | 37.43 | 67.19 | -0.58 | / |
| AeBAM15 | Chr23G1205 | 301 | 34 345.80 | 8.96 | 40.58 | 82.23 | -0.34 | / |
| AeBAM16 | Chr23G1206 | 501 | 56 428.79 | 8.00 | 37.67 | 66.99 | -0.56 | / |
| AeBAM17 | Chr25G384 | 546 | 61 462.78 | 8.58 | 36.62 | 70.57 | -0.51 | AdBAM3(Acc16695) |

Fig. 2 Collinearity and Ka/Ks analysis of AeBAM gene family members A:Collinearity analysis among AeBAM family members;B:Ka/Ks analysis of AeBAM family members;C:Collinearity analysis of BAM family members across multiple species
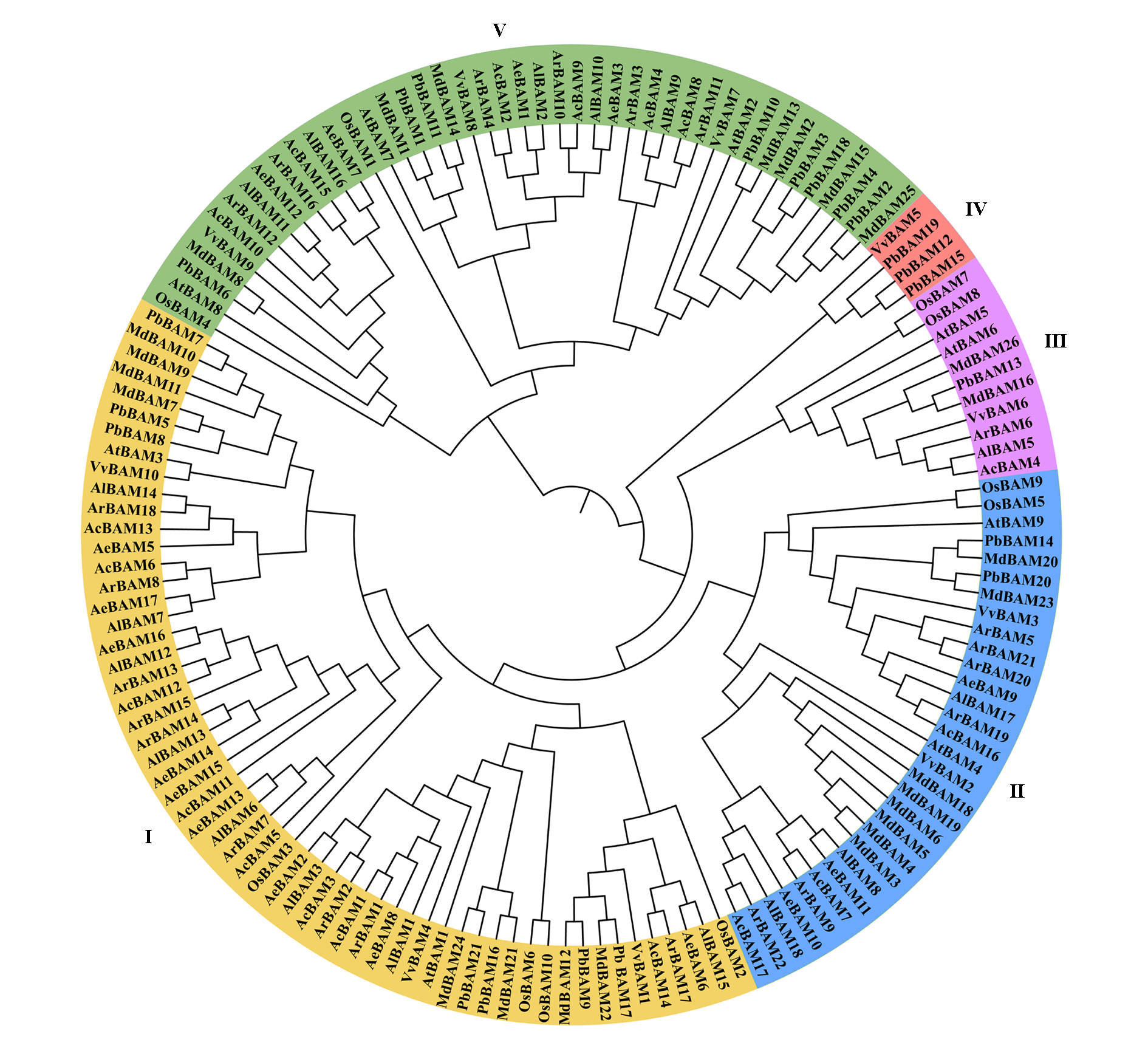
Fig. 3 Phylogenetic tree analysis of BAM between Actinidia eriantha and other species Ac:Actinidia chinensis;Ae:Actinidia eriantha;Al:Actinidia latifolia;Ar:Actinidia rufa;At:Arabidopsis thaliana;Md:Malus domestica;Os:Oryza sativa;Pb:Pyrus bretschneideri;Vv:Vitis vinifera
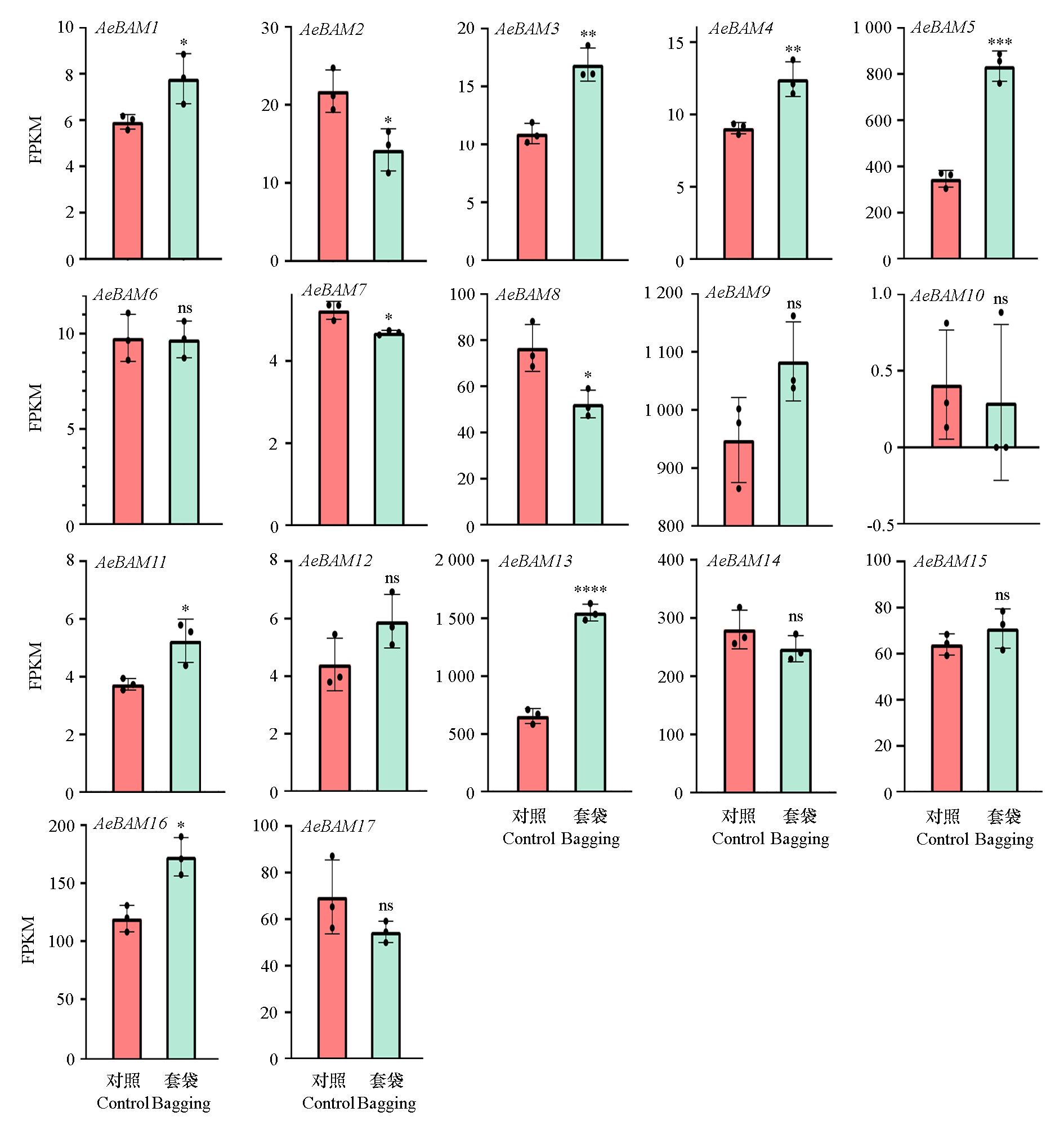
Fig. 6 Expression analysis of AeBAM family genes after fruit bagging * indicates significant differences at P < 0.05,** indicates significant differences at P < 0.01,*** indicates significant differences at P < 0.001,**** indicates significant differences at P < 0.0001,ns indicates no significant difference
| [1] |
|
| [2] |
doi: 10.1016/j.molp.2023.09.010 URL |
| [3] |
doi: 10.16420/j.issn.0513-353x.2024-0373 |
|
陈敏氡, 王彬, 白昌辉, 裘波音, 刘建汀, 温庆放, 朱海生, 李永平. 2025. 普通丝瓜Expansin家族基因的鉴定及其在果实膨大中的表达分析. 园艺学报,2025, 52 (6):1488-1504.
doi: 10.16420/j.issn.0513-353x.2024-0373 |
|
| [4] |
|
| [5] |
doi: 10.1371/journal.pcbi.1009703 URL |
| [6] |
doi: 10.1105/tpc.107.056507 URL |
| [7] |
|
| [8] |
doi: 10.3724/SP.J.1006.2017.01417 |
|
郝心愿, 岳川, 唐湖, 钱文俊, 王玉春, 王璐, 王新超, 杨亚军. 2017. 茶树β-淀粉酶基因CsBAM3的克隆及其响应低温的表达模式. 作物学报, 43 (10):1417-1425.
doi: 10.3724/SP.J.1006.2017.01417 |
|
| [9] |
doi: 10.1038/s41477-018-0287-6 pmid: 30374094 |
| [10] |
|
| [11] |
doi: 10.1046/j.1365-313X.1999.00625.x URL |
| [12] |
|
| [13] |
|
| [14] |
|
| [15] |
doi: 10.1186/s12870-021-02916-8 |
| [16] |
doi: 10.1016/j.jia.2023.07.018 URL |
| [17] |
|
| [18] |
doi: 10.1006/meth.2001.1262 pmid: 11846609 |
| [19] |
doi: 10.1186/s12864-022-08630-5 |
| [20] |
|
| [21] |
doi: 10.1104/pp.114.246421 URL |
| [22] |
|
| [23] |
doi: 10.1111/pce.2014.37.issue-12 URL |
| [24] |
doi: 10.1105/tpc.110.081950 pmid: 21487098 |
| [25] |
doi: 10.1111/jipb.v66.5 URL |
| [26] |
|
| [27] |
doi: 10.1016/j.pbi.2012.03.016 pmid: 22541711 |
| [28] |
|
|
田晓成, 祝令成, 邹晖, 李白云, 马锋旺, 李明军. 2023. 果实可溶性糖的积累模式及其调控研究进展. 园艺学报, 50 (4):885-895.
doi: 10.16420/j.issn.0513-353x.2021-1264 |
|
| [29] |
|
| [30] |
|
|
王成. 2023. 低温对薄皮甜瓜果实膨大期和采后淀粉代谢和糖积累的影响[硕士论文]. 沈阳:沈阳农业大学.
|
|
| [31] |
doi: 10.16420/j.issn.0513-353x.2024-0993 |
|
王心茹, 白帅帅, 孙雨乐, 武欣宇, 曾凤, 刘春英, 张玉喜, 盖树鹏, 袁延超. 2025. ‘凤丹’牡丹PoSnRK基因家族鉴定及其在花衰老中的表达分析. 园艺学报, 52 (9):2363-2376.
doi: 10.16420/j.issn.0513-353x.2024-0993 |
|
| [32] |
doi: 10.1186/s43897-022-00034-z |
| [33] |
|
|
赵粱怡. 2019. 梨β-淀粉酶基因家族生物信息学分析及PbBAM3基因抗寒性鉴定[硕士论文]. 南京: 南京农业大学.
|
| [1] | CHEN Yiquan, FAN Ronghui, LIN Bing, CHEN Yan, WU Jianshe, ZHONG Huaiqin. Screening and Validation of Reference Genes for qRT-PCR Analysis of Genes Related to Floral Scent Biosynthesis in Camellia japonica [J]. Acta Horticulturae Sinica, 2026, 53(2): 598-612. |
| [2] | WANG Yanqiu, BAN Jingjie, ZENG Yuhan, LAI Chunwang, HUANG Xiaoling, ZHOU Jianjin, WAN Kui, LAI Zhongxiong, LIN Yuling. Identification of the DREB Gene Family in Polygonatum cyrtonema and Analysis of Its Response to Salt Stress [J]. Acta Horticulturae Sinica, 2026, 53(1): 159-173. |
| [3] | WANG Baochen, LUO Chen, YAN Jinqiang, CHENG Xiaoxin, MO Renlian, LIU Wenrui, ZHANG Yuyang, JIANG Biao. Cloning,Expression,and Functional Research of BhYAB4b in Wax Gourd [J]. Acta Horticulturae Sinica, 2025, 52(9): 2353-2362. |
| [4] | ZHANG Ting, WANG Botao, ZHANG Lei, SONG Lihua, CAO Bing, MA Yaping. Identification of the LbWaxy Gene Family in Lycium barbarum and Analysis of Gene Expression Characteristics Under High Concentrations of CO2 [J]. Acta Horticulturae Sinica, 2025, 52(9): 2395-2409. |
| [5] | ZHANG Tuo, NIE Menghan, LIU Yuanyuan, DUAN Xinlian, HAN Xing, XIE Baogui, and YAN Junjie. Expression Analysis of the Catalase Genes FfCAT1 and FfCAT2 in Flammulina filiformis [J]. Acta Horticulturae Sinica, 2025, 52(7): 1769-1779. |
| [6] | YANG Chunmei, YU Yang, DING Yuge, XIA Jing, ZHOU Ling, and PENG Lei. Joint Transcription-Metabolism Analysis of Starch and Sucrose Metabolic Pathways in the Transformation of Axillary Buds into Flower Buds of ‘Guifei’Mango [J]. Acta Horticulturae Sinica, 2025, 52(6): 1412-1426. |
| [7] | SUN Tingzhen, SHU Qin, MA Wei, SHI Yuzi, ZHANG Meng, XIANG Chenggang, BO Kailiang, DUAN Ying, WANG Changlin. Identification and Phylogenetic Analysis of Gibberellin Oxidase Gene Family and Its Expression Pattern in Main Vine Growth in Cucurbita maxima [J]. Acta Horticulturae Sinica, 2025, 52(3): 603-622. |
| [8] | DENG Shuqin, GAO Yingrui, LI Yutong, WANG Ying, GONG Chunmei, BAI Juan. Response of Ubiquitin-ligase Gene CsPUB21 to Different Abiotic Stress in Camellia sinensis [J]. Acta Horticulturae Sinica, 2025, 52(3): 655-670. |
| [9] | TONG Zongjun, HAN Xing, DUAN Xinlian, LIU Yuanyuan, LIN Junbin, GAN Ying, CHEN Jie, XIE Baogui, GAN Bingcheng, YAN Junjie. Sequence Characterization of PRX Family and Expression Analysis in the Process of Fruiting Body Development in Flammulina filiformis [J]. Acta Horticulturae Sinica, 2025, 52(2): 337-348. |
| [10] | HAN Hongwei, CHANG Yanan, LIU Huifang, ZHUANG Hongmei, XING Jiayi, ZHANG Xiaoda, ZULIPIYE · Abudumilike, WANG Hao, WANG Qiang. Identification of Cytochrome P450 Family Genes in Melon and Expression Analysis in Response to Light Intensity [J]. Acta Horticulturae Sinica, 2025, 52(12): 3271-3287. |
| [11] | LIU Lu, WANG Meng, ZHENG Xiaomei, WANG Keqing, MIAO Ruyi, ZANG Qiaolu. Identification and Expression Analysis of CBF Gene Family of Artemisia argyi [J]. Acta Horticulturae Sinica, 2025, 52(10): 2703-2712. |
| [12] | HE Dandan, HE Hongtai, WANG Wenting, ZHOU Wenmei, LIU Yanmin, LIU Sushuang. Identification of Melon GolS Genes Family and the Expression Analysis in Response to Low Temperature Stress [J]. Acta Horticulturae Sinica, 2025, 52(1): 136-148. |
| [13] | GUO Changquan, LI Danqi, HUI Xinran, ZHENG Jingya, HOU Menglu, ZHU Yongxing. Identification and Expression Pattern Analysis of DUF966 Gene Family Members in Ginger [J]. Acta Horticulturae Sinica, 2024, 51(9): 2031-2047. |
| [14] | ZHAO Jiaying, ZENG Zhouting, CEN Xinying, SHI Jiaoqi, LI Xiaoxian, SHEN Xiaoxia, YU Zhenming. Genome-Wide Identification and Expression Analysis of CCO Gene Family in Dendrobium officinale During Flower Development [J]. Acta Horticulturae Sinica, 2024, 51(9): 2075-2088. |
| [15] | LIU Yanyan, DING Ying, LIU Xinghua, ZHENG Jiaqiu, LIU Zhiqin. Identification of Pepper CaSYT1 and Its Function in Phytophthora capsici Infection [J]. Acta Horticulturae Sinica, 2024, 51(3): 533-544. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd