
Acta Horticulturae Sinica ›› 2026, Vol. 53 ›› Issue (1): 131-148.doi: 10.16420/j.issn.0513-353x.2024-0952
• Genetic & Breeding·Germplasm Resources·Molecular Biology • Previous Articles Next Articles
ZHU Xuanyi1, ZHAO Yuqing1, TANG Feihong1, WU Haiyan1, ZHANG Chun3, ZHAO Kai2, PENG Donghui1, LAN Siren1, ZHOU Yuzhen1,*( )
)
Received:2025-03-13
Revised:2025-09-02
Online:2026-01-25
Published:2026-01-26
Contact:
ZHOU Yuzhen
ZHU Xuanyi, ZHAO Yuqing, TANG Feihong, WU Haiyan, ZHANG Chun, ZHAO Kai, PENG Donghui, LAN Siren, ZHOU Yuzhen. Whole-Genome Identification of the CNGC Gene Family in Cymbidium ensifolium and Transcriptional Reponses Under Abiotic Stresses[J]. Acta Horticulturae Sinica, 2026, 53(1): 131-148.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2024-0952
| 引物名称 Primer name | 正向引物(5′-3′) Forward primer | 反向引物(5′-3′) Reverse primer |
|---|---|---|
| CeCNGC1 | AGCCTATCAGACTCCTTCCACCTTG | CGAGCTTCCTTCCTGGCACTTG |
| CeCNGC2 | CTTCGTAATCCTTCCGCTTCCTCAG | GGTACACCTTCGGCAAGAACTGG |
| CeCNGC3 | AACGGTGAAGGCATTATCTGAGGTG | GCTTGCTATGGAGTCTGCGGAAC |
| CeCNGC4 | AGATGCGATATGCGAGCGATTAGTC | TCGTCCACTGGGTCACCTTCC |
| CeCNGC5 | ACAGTGAAGGCATTGTCGGAAGTC | TGAAGGTGTGCTGGAGTTGCTTAC |
| CeCNGC6 | AGTACAAGTGGTTGGCAACTCAAGG | GAGGCAGAGATGTCGCTTAATGTCC |
| CeCNGC7 | CAGGTTGTTCTATGGGTGGCAGTAC | CGGAGGAAGGAGATGGAGTGGTAG |
| CeCNGC8 | GTGGGAGCGGGCACTTTGATG | AGAGAAGCAAGCACTGGCATTAGG |
| CeCNGC9 | CACAGAGAGCATCTTCGTCGTCAG | TCGCCGCAGAAATCGCCTTG |
| CeCNGC10 | CACCGAGAGCACTTACATTGTCAGG | AAGCCACTCCTCCCACCATCAG |
| CeCNGC11 | GAGGTGCTTGCTGGTACTTACTTGG | GAAGTTGTGGTGCCGTAGTAGATGG |
| CeCNGC12 | GGAGCACCGCCCTGTCTTTATATG | CACGTAAGCCATCCTCATCTGTAGC |
| CeCNGC13 | CGTCGGCACAAGAGAAGGAAGATG | TCGCTTGGCTCTCCGCTCTC |
| TUB | GCAGTTTACGGCGATGTTCA | ACTCTTCCTCGTCAGCTGTG |
| 引物名称 Primer name | 正向引物(5′-3′) Forward primer | 反向引物(5′-3′) Reverse primer |
|---|---|---|
| CeCNGC1 | AGCCTATCAGACTCCTTCCACCTTG | CGAGCTTCCTTCCTGGCACTTG |
| CeCNGC2 | CTTCGTAATCCTTCCGCTTCCTCAG | GGTACACCTTCGGCAAGAACTGG |
| CeCNGC3 | AACGGTGAAGGCATTATCTGAGGTG | GCTTGCTATGGAGTCTGCGGAAC |
| CeCNGC4 | AGATGCGATATGCGAGCGATTAGTC | TCGTCCACTGGGTCACCTTCC |
| CeCNGC5 | ACAGTGAAGGCATTGTCGGAAGTC | TGAAGGTGTGCTGGAGTTGCTTAC |
| CeCNGC6 | AGTACAAGTGGTTGGCAACTCAAGG | GAGGCAGAGATGTCGCTTAATGTCC |
| CeCNGC7 | CAGGTTGTTCTATGGGTGGCAGTAC | CGGAGGAAGGAGATGGAGTGGTAG |
| CeCNGC8 | GTGGGAGCGGGCACTTTGATG | AGAGAAGCAAGCACTGGCATTAGG |
| CeCNGC9 | CACAGAGAGCATCTTCGTCGTCAG | TCGCCGCAGAAATCGCCTTG |
| CeCNGC10 | CACCGAGAGCACTTACATTGTCAGG | AAGCCACTCCTCCCACCATCAG |
| CeCNGC11 | GAGGTGCTTGCTGGTACTTACTTGG | GAAGTTGTGGTGCCGTAGTAGATGG |
| CeCNGC12 | GGAGCACCGCCCTGTCTTTATATG | CACGTAAGCCATCCTCATCTGTAGC |
| CeCNGC13 | CGTCGGCACAAGAGAAGGAAGATG | TCGCTTGGCTCTCCGCTCTC |
| TUB | GCAGTTTACGGCGATGTTCA | ACTCTTCCTCGTCAGCTGTG |
| 基因名 Gene name | 基因ID Gene ID | 编码序列长度/bp CDS length | 氨基酸数 Number of amino acids | 分子量/D Molecular weight | 等电点 pI |
|---|---|---|---|---|---|
| CeCNGC1 | JL014344 | 1 461 | 488 | 55 806.43 | 8.23 |
| CeCNGC2 | JL001908 | 2 200 | 735 | 82 623.44 | 9.56 |
| CeCNGC3 | JL010106 | 1 029 | 343 | 40 046.06 | 9.14 |
| CeCNGC4 | JL001369 | 1 591 | 531 | 60 926.40 | 8.34 |
| CeCNGC5 | JL000073 | 2 105 | 703 | 81 211.91 | 8.94 |
| CeCNGC6 | JL006714 | 1 497 | 501 | 58 400.86 | 9.60 |
| CeCNGC7 | JL008860 | 1 869 | 625 | 71 627.58 | 9.35 |
| CeCNGC8 | JL007202 | 2 268 | 759 | 86 931.94 | 9.25 |
| CeCNGC9 | JL005182 | 879 | 293 | 34 538.52 | 9.81 |
| CeCNGC10 | JL002960 | 2 174 | 726 | 83 914.57 | 9.11 |
| CeCNGC11 | JL010726 | 1 953 | 653 | 74 573.27 | 9.70 |
| CeCNGC12 | JL019980 | 2 146 | 717 | 81 825.83 | 9.43 |
| CeCNGC13 | JL019514 | 1 313 | 438 | 50 481.31 | 8.94 |
| 基因名 Gene name | 基因ID Gene ID | 不稳定系数 Instability index | 脂肪系数 Aliphatic index | 总平均亲水系数 Grand average of hydropathicity | 亚细胞定位 Subcellular localization |
| CeCNGC1 | JL014344 | 41.29 | 80.39 | -0.251 | 叶绿体Chloroplast |
| CeCNGC2 | JL001908 | 58.92 | 92.49 | 0.019 | 质膜Plasma membrane |
| CeCNGC3 | JL010106 | 54.80 | 79.91 | -0.457 | 叶绿体Chloroplast |
| CeCNGC4 | JL001369 | 43.95 | 90.19 | -0.158 | 质膜Plasma membrane |
| CeCNGC5 | JL000073 | 50.71 | 95.68 | -0.103 | 质膜Plasma membrane |
| CeCNGC6 | JL006714 | 50.76 | 92.26 | -0.103 | 叶绿体Chloroplast |
| CeCNGC7 | JL008860 | 50.00 | 96.74 | 0.012 | 质膜Plasma membrane |
| CeCNGC8 | JL007202 | 42.34 | 89.80 | -0.129 | 质膜Plasma membrane |
| CeCNGC9 | JL005182 | 67.42 | 80.48 | -0.616 | 细胞核nucleus |
| CeCNGC10 | JL002960 | 54.21 | 87.34 | -0.240 | 质膜Plasma membrane |
| CeCNGC11 | JL010726 | 55.62 | 87.35 | -0.270 | 质膜Plasma membrane |
| CeCNGC12 | JL019980 | 56.53 | 93.05 | 0.041 | 质膜Plasma membrane |
| CeCNGC13 | JL019514 | 53.37 | 76.64 | -0.432 | 叶绿体Chloroplast |
Table 2 Physicochemical properties and subcellular localization of CeCNGC family members
| 基因名 Gene name | 基因ID Gene ID | 编码序列长度/bp CDS length | 氨基酸数 Number of amino acids | 分子量/D Molecular weight | 等电点 pI |
|---|---|---|---|---|---|
| CeCNGC1 | JL014344 | 1 461 | 488 | 55 806.43 | 8.23 |
| CeCNGC2 | JL001908 | 2 200 | 735 | 82 623.44 | 9.56 |
| CeCNGC3 | JL010106 | 1 029 | 343 | 40 046.06 | 9.14 |
| CeCNGC4 | JL001369 | 1 591 | 531 | 60 926.40 | 8.34 |
| CeCNGC5 | JL000073 | 2 105 | 703 | 81 211.91 | 8.94 |
| CeCNGC6 | JL006714 | 1 497 | 501 | 58 400.86 | 9.60 |
| CeCNGC7 | JL008860 | 1 869 | 625 | 71 627.58 | 9.35 |
| CeCNGC8 | JL007202 | 2 268 | 759 | 86 931.94 | 9.25 |
| CeCNGC9 | JL005182 | 879 | 293 | 34 538.52 | 9.81 |
| CeCNGC10 | JL002960 | 2 174 | 726 | 83 914.57 | 9.11 |
| CeCNGC11 | JL010726 | 1 953 | 653 | 74 573.27 | 9.70 |
| CeCNGC12 | JL019980 | 2 146 | 717 | 81 825.83 | 9.43 |
| CeCNGC13 | JL019514 | 1 313 | 438 | 50 481.31 | 8.94 |
| 基因名 Gene name | 基因ID Gene ID | 不稳定系数 Instability index | 脂肪系数 Aliphatic index | 总平均亲水系数 Grand average of hydropathicity | 亚细胞定位 Subcellular localization |
| CeCNGC1 | JL014344 | 41.29 | 80.39 | -0.251 | 叶绿体Chloroplast |
| CeCNGC2 | JL001908 | 58.92 | 92.49 | 0.019 | 质膜Plasma membrane |
| CeCNGC3 | JL010106 | 54.80 | 79.91 | -0.457 | 叶绿体Chloroplast |
| CeCNGC4 | JL001369 | 43.95 | 90.19 | -0.158 | 质膜Plasma membrane |
| CeCNGC5 | JL000073 | 50.71 | 95.68 | -0.103 | 质膜Plasma membrane |
| CeCNGC6 | JL006714 | 50.76 | 92.26 | -0.103 | 叶绿体Chloroplast |
| CeCNGC7 | JL008860 | 50.00 | 96.74 | 0.012 | 质膜Plasma membrane |
| CeCNGC8 | JL007202 | 42.34 | 89.80 | -0.129 | 质膜Plasma membrane |
| CeCNGC9 | JL005182 | 67.42 | 80.48 | -0.616 | 细胞核nucleus |
| CeCNGC10 | JL002960 | 54.21 | 87.34 | -0.240 | 质膜Plasma membrane |
| CeCNGC11 | JL010726 | 55.62 | 87.35 | -0.270 | 质膜Plasma membrane |
| CeCNGC12 | JL019980 | 56.53 | 93.05 | 0.041 | 质膜Plasma membrane |
| CeCNGC13 | JL019514 | 53.37 | 76.64 | -0.432 | 叶绿体Chloroplast |
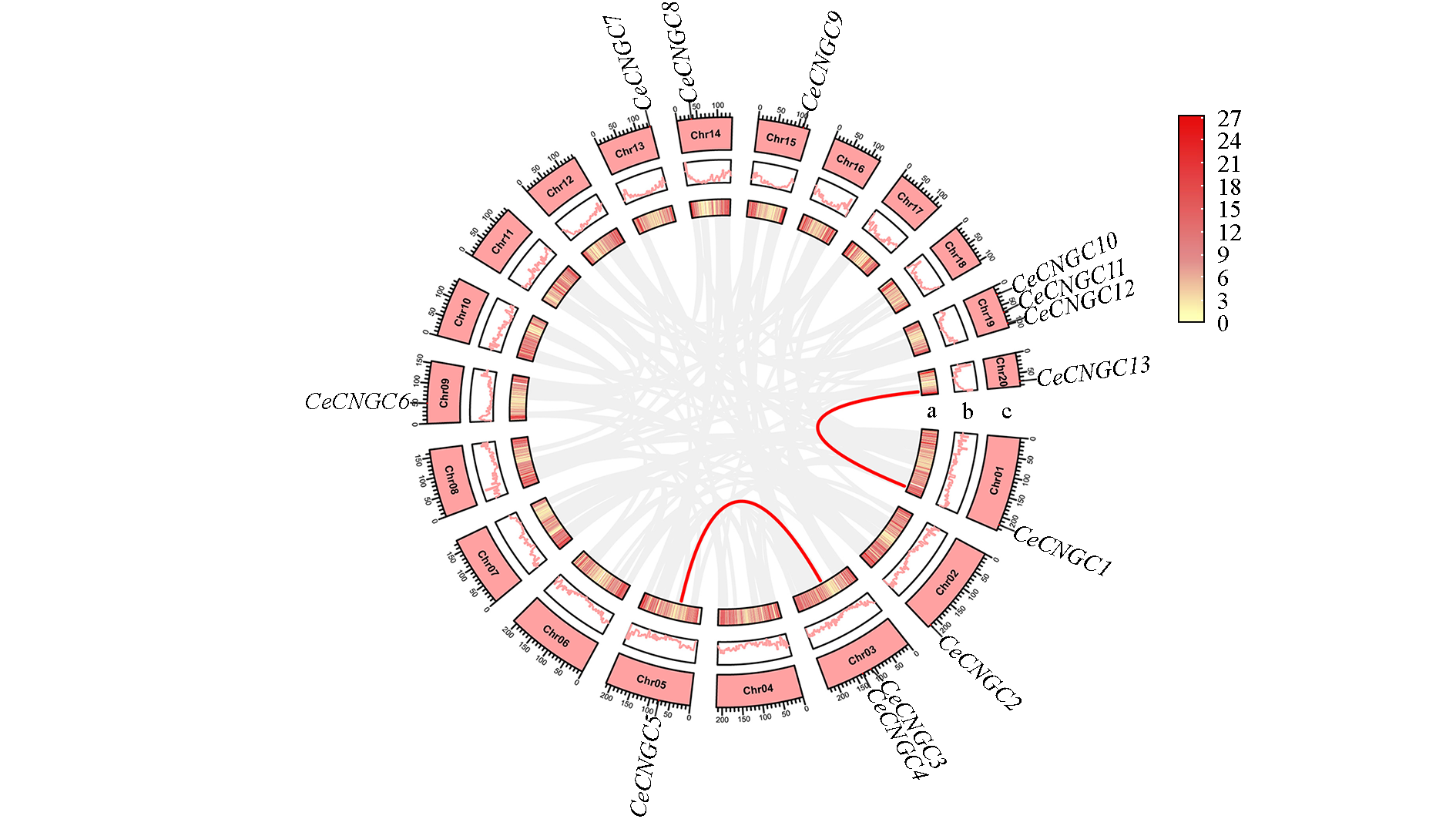
Fig. 2 Chromosomal locations and segmental duplication of the CeCNGCs in the Cymbidium ensifolium genome The red lines inside indicate pairs of CeCNGCs involved in uplicated events
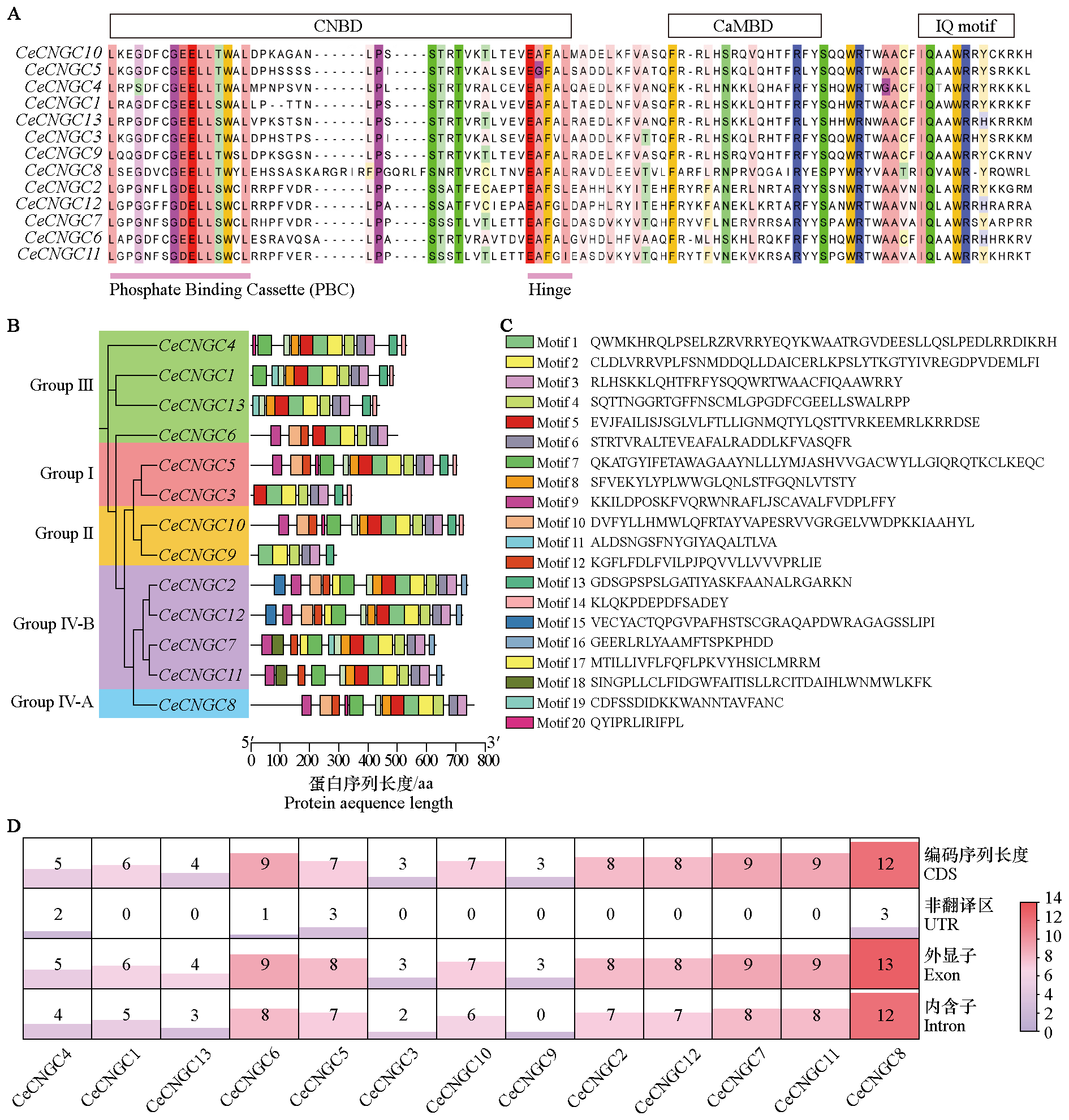
Fig. 3 Multiple sequence alignment of the CeCNGC family in Cymbidium ensifolium(A);motif composition and amino acid sequences(B,C)and CDS,UTR,exons,and introns(D)
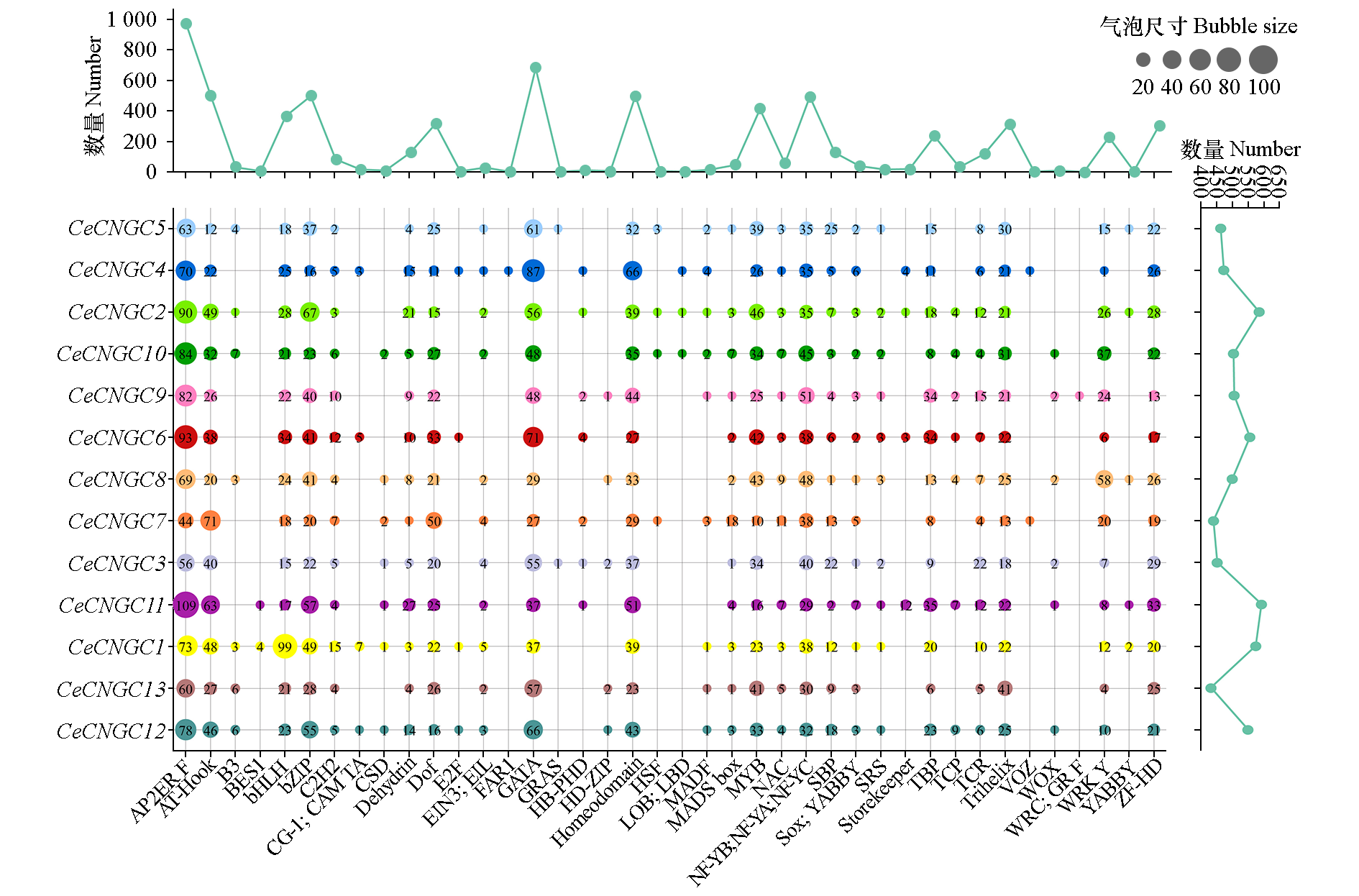
Fig. 4 Analysis of cis-acting elements in CeCNGCs promoters Bubble plot illustrates the distribution of cis-acting elements within the CeCNGC gene family. The horizontal line graph depicts the cumulative count of each cis-acting element across all CeCNGC,while the vertical line graph shows the total number of cis-acting elements present in each individual CeCNGC

Fig. 7 Predicted protein-protein interaction networks of CeCNGC proteins The outer ring represents proteins interacting with CeCNGCs,while the inner ring represents the CeCNGC proteins themselves. The darker colors indicating higher connectivity with other members. The orange-yellow circles specifically illustrate the protein interaction network involving only CeCNGC proteins
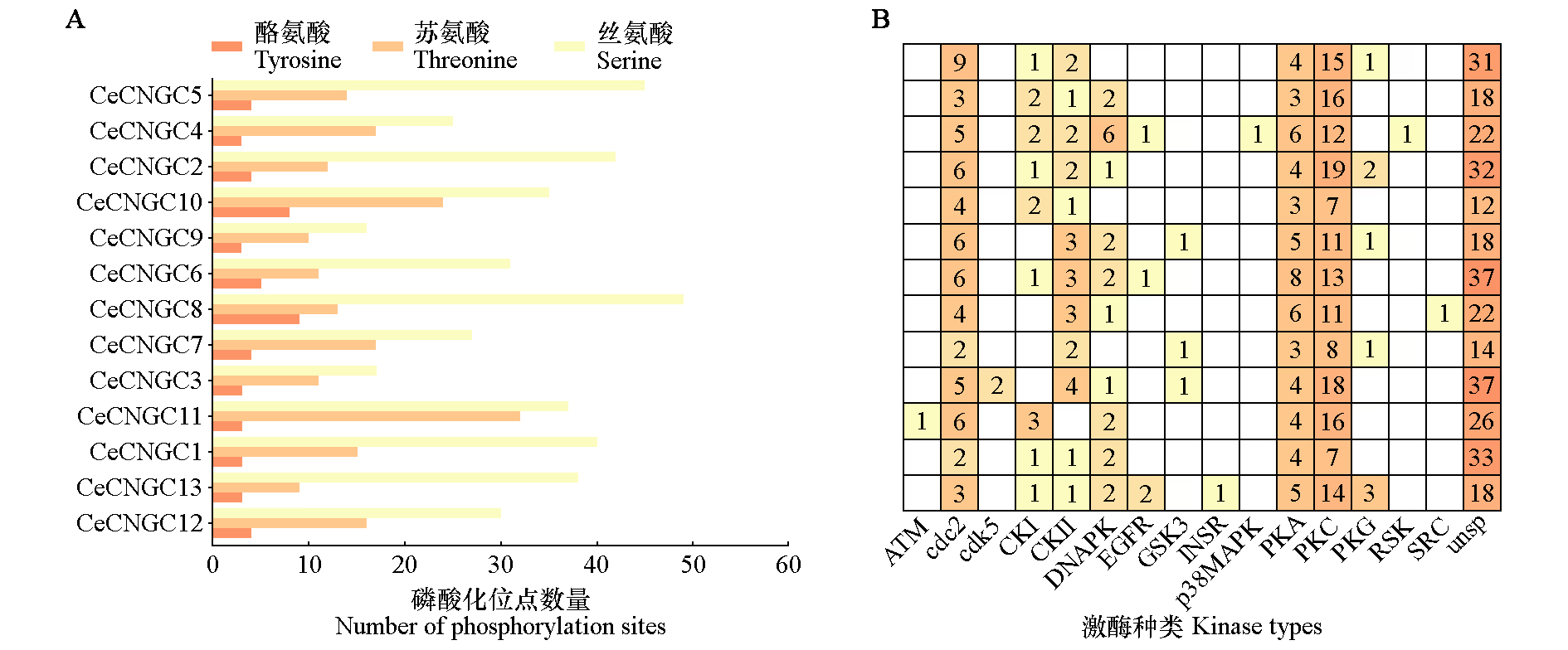
Fig. 8 The number of phosphorylation sites(A)and kinase binding sites(B)in CeCNGC proteins unsp indicates the prediction results of non-specific kinases
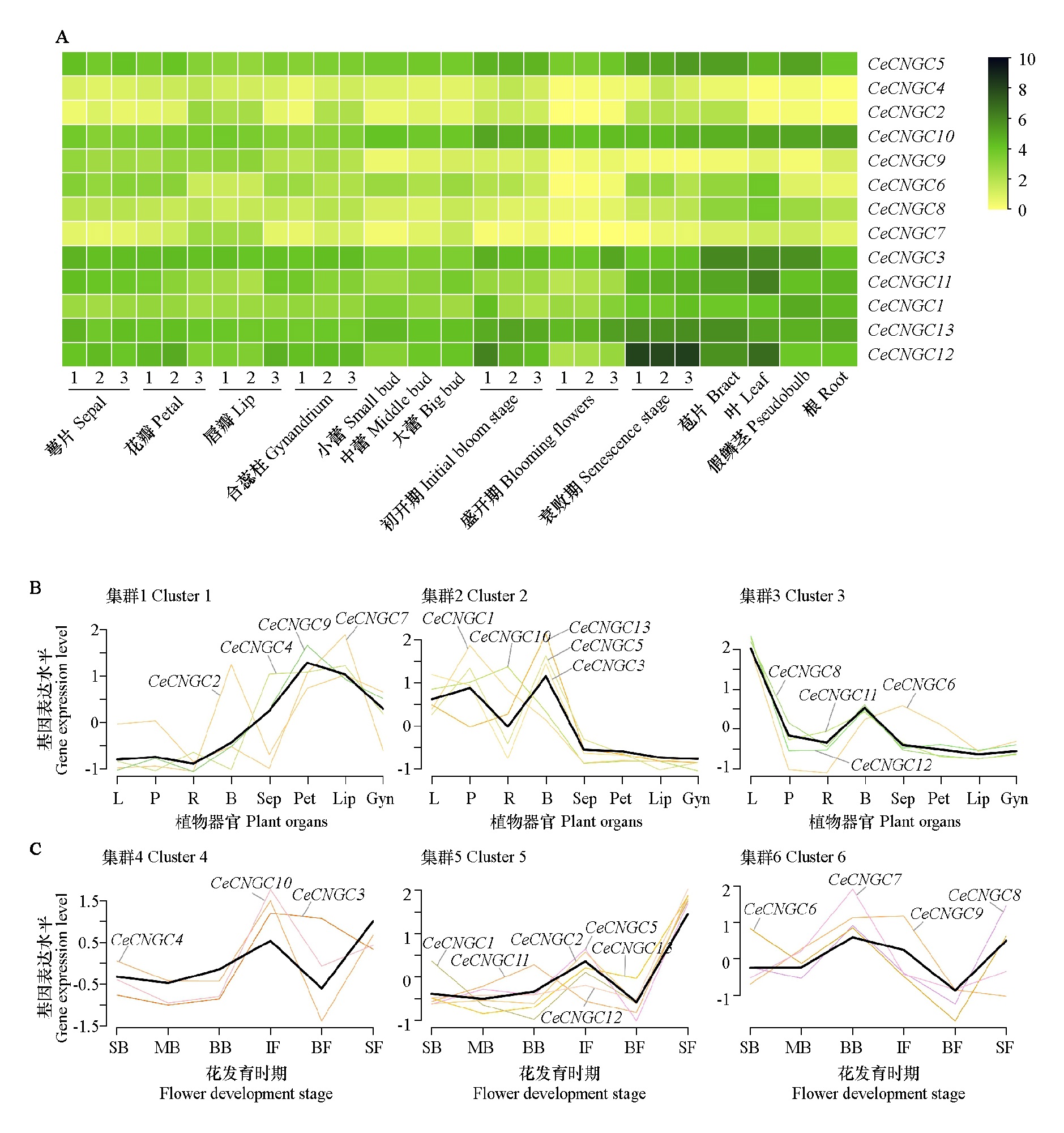
Fig. 9 Heatmap(A),clustering analysis of gene expression trends in different plant tissues(B)clustering analysis of gene expression trends at different flower development stages(C)and of CeCNGC family members The thick brown line represents the overall expression pattern of the genes within the cluster. L:Leaf;P:Pseudobulb;R:Root;B:Bract;Sep:Sepal;Pet:Petal;Lip:Lip;Gyn:Gynandrium;SB:Samll bud;MB:Middle bud;BB:Big bud;IF:Initial bloom stage;BF:Blooming flowers;SF:Senescence stage
| [1] |
doi: 10.1038/s41598-021-03712-y pmid: 34921218 |
| [2] |
doi: 10.1038/nbt.3519 pmid: 27043002 |
| [3] |
doi: 10.1016/j.molp.2023.09.010 URL |
| [4] |
doi: 10.1016/j.ygeno.2014.11.006 URL |
| [5] |
doi: 10.1111/jipb.v63.1 URL |
| [6] |
|
| [7] |
doi: 10.1104/pp.20.00591 pmid: 32527735 |
| [8] |
|
| [9] |
doi: 10.1104/pp.20.00425 pmid: 32576644 |
| [10] |
|
| [11] |
doi: 10.1016/j.cj.2020.07.007 URL |
| [12] |
|
| [13] |
doi: 10.16420/j.issn.0513-353x.2024-0946 |
|
贺子怡, 曾瑞珍, 谢利, 魏倩, 张志胜. 2025. 国兰育种:历史、现状和展望. 园艺学报, 52 (8):2233-2248.
doi: 10.16420/j.issn.0513-353x.2024-0946 |
|
| [14] |
doi: 10.3390/biom11081159 URL |
| [15] |
doi: 10.1186/s12864-017-4244-y URL |
| [16] |
doi: 10.1016/j.bbagrm.2011.05.001 pmid: 21605713 |
| [17] |
|
| [18] |
doi: 10.1105/tpc.109.072686 URL |
| [19] |
doi: 10.1111/nph.14966 pmid: 29332310 |
| [20] |
doi: 10.1186/s12870-023-04050-z |
| [21] |
doi: 10.1104/pp.126.4.1646 pmid: 11500563 |
| [22] |
doi: 10.1104/pp.106.091298 URL |
| [23] |
doi: 10.1186/1471-2164-15-853 |
| [24] |
doi: 10.1093/plcell/koae177 URL |
| [25] |
doi: 10.1007/s12298-021-01061-8 |
| [26] |
|
| [27] |
doi: 10.1093/dnares/dsv029 URL |
| [28] |
|
| [29] |
pmid: 12818663 |
| [30] |
|
| [31] |
|
| [32] |
doi: 10.1007/s00606-011-0429-z URL |
| [33] |
doi: 10.1016/j.molp.2020.11.022 URL |
| [34] |
doi: 10.1186/s12864-020-6601-5 |
| [35] |
doi: 10.1093/nar/gkr1293 URL |
| [36] |
doi: 10.1093/bioinformatics/btp033 pmid: 19151095 |
| [37] |
doi: 10.1016/j.tplants.2004.12.012 pmid: 15708346 |
| [38] |
|
| [39] |
|
| [40] |
|
| [41] |
doi: 10.1186/s12864-023-09307-3 |
| [42] |
doi: 10.1186/s12870-023-04604-1 |
| [43] |
|
| [1] | LI Meiqing, LUO Sifei, JIA Yaohao, WANG Weigui, and SUN Jin. Expression Analysis of the Cucumber Mg-Dechelatase Gene CsMDC and Functional Verification of Its Role in Regulating Chlorlorophyll Degradation [J]. Acta Horticulturae Sinica, 2025, 52(6): 1463-1476. |
| [2] | MA Yuwan, LIU Ao, LIU Xiangdong, ZHANG Yajing, DONG Xuanke, LI Yufan, CHEN Jiren. Cloning of the RcRAP2.7 Gene and its Expression Analysis Under Abiotic Stresses in Rosa chinensis‘Yueyuehong’ [J]. Acta Horticulturae Sinica, 2025, 52(4): 921-932. |
| [3] | WANG Yunyun, ZHOU Hui, QIU Keli, PAN Haifa, SHENG Yu, SHI Pei, XIE Qingmei, CHEN Hongli, ZHANG Jinyun, LI Dahui. Identification and Expression Analysis of Peach JMJ Histone Demethylase Gene Family [J]. Acta Horticulturae Sinica, 2025, 52(3): 575-590. |
| [4] | DENG Shuqin, GAO Yingrui, LI Yutong, WANG Ying, GONG Chunmei, BAI Juan. Response of Ubiquitin-ligase Gene CsPUB21 to Different Abiotic Stress in Camellia sinensis [J]. Acta Horticulturae Sinica, 2025, 52(3): 655-670. |
| [5] | LI Yadi, WANG Hanxiang, HU Baigeng, YANG Hui, HU Xinxi, XIONG Xingyao, WANG Wanxing. Research Progress on Plant Growth Promoting Rhizobacteria to Alleviate Abiotic Stress Tolerance of Horticultural Crops [J]. Acta Horticulturae Sinica, 2024, 51(8): 1964-1976. |
| [6] | LI Yachen, ZHENG Yanmei, SONG Wenpei, LI Dawei, LIANG Hong, ZHANG Xianzhi. Research Progress on the Role of Hydrogen-Rich Water in Plant Growth and Development and Stress Response [J]. Acta Horticulturae Sinica, 2024, 51(7): 1489-1500. |
| [7] | LUO Xinrui, ZHANG Xiaoxu, WANG Yuping, WANG Zhi, MA Yuanyuan, ZHOU Bingyue. Identification and Expression Analysis of Trihelix Gene Family in Common Bean [J]. Acta Horticulturae Sinica, 2024, 51(12): 2775-2790. |
| [8] | ZHANG Pan, YU Yongxu, CAO Linggai, ZHU Guangbing, WU Wei, GUO Yushuang, YIN Guoying, JIA Meng’ao. Research Progress of m6A Methylation Modification Response to Plant Biotic and Abiotic Stresses [J]. Acta Horticulturae Sinica, 2023, 50(9): 1841-1853. |
| [9] | WEI Jianming, LI Yunzhou, LIANG Yan. Advances in Research on Improving Tomato Disease Resistance and Stress Resistance by Grafting Technology [J]. Acta Horticulturae Sinica, 2023, 50(9): 1997-2014. |
| [10] | SHEN Yuxiao, ZOU Jinyu, LUO Ping, SHANG Wenqian, LI Yonghua, HE Songlin, WANG Zheng, SHI Liyun. Genome-wide Identification and Abiotic Stress Response Analysis of PP2C Family Genes in Rosa chinensis‘Old Blush’ [J]. Acta Horticulturae Sinica, 2023, 50(10): 2139-2156. |
| [11] | XU Xiaoping, CAO Qingying, CAI Roudi, GUAN Qingxu, ZHANG Zihao, CHEN Yukun, XU HAN, LIN Yuling, LAI Zhongxiong. Gene Cloning and Expression Analysis of miR408 and Its Target DlLAC12 in Globular Embryo Development and Abiotic Stress in Dimocarpus longan [J]. Acta Horticulturae Sinica, 2022, 49(9): 1866-1882. |
| [12] | JIA Xin, ZENG Zhen, CHEN Yue, FENG Hui, LÜ Yingmin, ZHAO Shiwei. Cloning and Expression Analysis of RcDREB2A Gene in Rosa chinensis‘Old Blush’ [J]. Acta Horticulturae Sinica, 2022, 49(9): 1945-1956. |
| [13] | MA Weifeng, LI Yanmei, MA Zonghuan, CHEN Baihong, MAO Juan. Identification of Apple POD Gene Family and Functional Analysis of MdPOD15 Gene [J]. Acta Horticulturae Sinica, 2022, 49(6): 1181-1199. |
| [14] | ZHOU Zhiming, YANG Jiabao, ZHANG Cheng, ZENG Linglu, MENG Wanqiu, SUN Li. Genome-wide Identification and Expression Analyses of Long-chain Acyl-CoA Synthetases Under Abiotic Stresses in Helianthus annuus [J]. Acta Horticulturae Sinica, 2022, 49(2): 352-364. |
| [15] | XIE Siyi, ZHOU Chengzhe, ZHU Chen, ZHAN Dongmei, CHEN Lan, WU Zuchun, LAI Zhongxiong, GUO Yuqiong. Genome-wide Identification and Expression Analysis of CsTIFY Transcription Factor Family Under Abiotic Stress and Hormone Treatments in Camellia sinensis [J]. Acta Horticulturae Sinica, 2022, 49(1): 100-116. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd