
园艺学报 ›› 2021, Vol. 48 ›› Issue (2): 233-242.doi: 10.16420/j.issn.0513-353x.2020-0350
梅闯1, 张小燕2, 闫鹏1,*( ), 艾沙江·买买提1, 冯贝贝1, 马凯1, 韩立群1, 董连新2, 王继勋1,*(
), 艾沙江·买买提1, 冯贝贝1, 马凯1, 韩立群1, 董连新2, 王继勋1,*( )
)
收稿日期:2020-11-29
修回日期:2021-01-18
出版日期:2021-02-25
发布日期:2021-03-09
通讯作者:
闫鹏,王继勋
E-mail:xaasyysyp@163.com;ee_wjx@163.com
基金资助:
MEI Chuang1, ZHANG Xiaoyan2, YAN Peng1,*( ), Aisajan Mamat1, FENG Beibei1, MA Kai1, HAN Liqun1, DONG Lianxin2, WANG Jixun1,*(
), Aisajan Mamat1, FENG Beibei1, MA Kai1, HAN Liqun1, DONG Lianxin2, WANG Jixun1,*( )
)
Received:2020-11-29
Revised:2021-01-18
Online:2021-02-25
Published:2021-03-09
Contact:
YAN Peng,WANG Jixun
E-mail:xaasyysyp@163.com;ee_wjx@163.com
摘要:
利用生物信息学方法在苹果全基因组中鉴定了TIFY转录因子家族基因,对基因结构、蛋白保守基序和进化关系进行了分析,同时基于新疆野苹果(Malus sieversii)转录组数据分析,明确了苹果TIFY家族基因在虫害胁迫条件下的差异表达情况。共鉴定到了16个苹果TIFY基因,其蛋白质大小介于209 ~ 449 aa之间。16个TIFY基因分布于苹果8条染色体中,所有的MdTIFY蛋白都具有保守的TIFY结构域。其中8个苹果MdTIFY与新疆野苹果转录组测序拼接转录本对应,这些基因均含有保守的TIFY功能域,多物种进化分析表明该类蛋白可聚类成7个组。7个TIFY基因的表达量在不同处理间表现出显著差异,虫害侵染后,抗虫株系中MsTIFY10B-a表达量升高34倍,MsTIFY9-c上升5.2倍。同时抗虫株系在自然条件下茉莉酸—异亮氨酸含量显著高于敏感株系。新疆野苹果中重要抗虫响应基因MsTIFY10B-a和MsTIFY9-c位于预测关联网络节点中心位置,使它们在调节植物对生物胁迫的响应中发挥重要作用。
中图分类号:
梅闯, 张小燕, 闫鹏, 艾沙江·买买提, 冯贝贝, 马凯, 韩立群, 董连新, 王继勋. 苹果TIFY家族基因鉴定及其在虫害胁迫下的表达分析[J]. 园艺学报, 2021, 48(2): 233-242.
MEI Chuang, ZHANG Xiaoyan, YAN Peng, Aisajan Mamat, FENG Beibei, MA Kai, HAN Liqun, DONG Lianxin, WANG Jixun. Identification of TIFY Family in Apple and Their Expression Analysis Under Insect Stress[J]. Acta Horticulturae Sinica, 2021, 48(2): 233-242.
| 物种 Species | 基因名 Gene name | 基因号 Gene ID | 物种 Species | 基因名 Gene name | 基因号 Gene ID |
|---|---|---|---|---|---|
| 拟南芥 Arabidopsis thaliana | AthTIFY11A | AT1G17380.1 | 番茄 Solanum lycoperiscon | SLTIFY5A | Solyc01g097060.2 |
| AthTIFY10A | AT1G19180.1 | SLTIFY10A | Solyc01g103600.2 | ||
| AthTIFY11B | AT1G72450.1 | SLTIFY10b | Solyc03g122190.2 | ||
| AthTIFY10B | AT1G74950.1 | SLTIFY8 | Solyc06g065650.2 | ||
| AthTIFY3A | AT3G43440.1 | SLTIFY10b-A | Solyc07g042170.2 | ||
| AthTIFY8 | AT4G32570.1 | SLTIFY5A-a | Solyc08g036620.2 | ||
| AthTIFY3B | AT5G20900.1 | SLTIFY5A-b | Solyc08g036640.2 | ||
| 水稻 Oryza sativa | OsTIFY11a | Os03t0180800-01 | SLTIFY5A-c | Solyc08g036660.2 | |
| OsTIFY11c | Os03t0180900-01 | SLTIFY10b-B | Solyc12g009220.1 | ||
| OsTIFY11b | Os03t0181100-01 | SLTIFY10b-C | Solyc12g049400.1 | ||
| OsTIFY11g | Os03t0396500-00 | 葡萄 Vitis vinifera | VvTIFY5A | VIT_04s0008g00110.t01 | |
| OsTIFY10a | Os03t0402800-01 | VvTIFY8 | VIT_04s0008g04950.t01 | ||
| OsTIFY3 | Os04t0653000-01 | VvTIFY10A | VIT_09s0002g00890.t01 | ||
| OsTIFY5 | Os07t0153000-00 | VvTIFY5A-a | VIT_10s0003g03790.t01 | ||
| OsTIFY10b | Os07t0615200-01 | VvTIFY5A-b | VIT_10s0003g03800.t01 | ||
| OsTIFY10c | Os09t0439200-01 | VvTIFY5A-c | VIT_10s0003g03810.t01 | ||
| OsTIFY11e | Os10t0391400-01 | VvTIFY5A-d | VIT_01s0011g05560.t01 | ||
| OsTIFY11d | Os10t0392400-01 | VvTIFY10A-a | VIT_11s0016g00710.t01 | ||
| VvTIFY10A-b | VIT_12s0035g00900.t01 |
表1 系统发育分析中的基因ID信息
Table 1 Gene ID information in phylogenetic analysis
| 物种 Species | 基因名 Gene name | 基因号 Gene ID | 物种 Species | 基因名 Gene name | 基因号 Gene ID |
|---|---|---|---|---|---|
| 拟南芥 Arabidopsis thaliana | AthTIFY11A | AT1G17380.1 | 番茄 Solanum lycoperiscon | SLTIFY5A | Solyc01g097060.2 |
| AthTIFY10A | AT1G19180.1 | SLTIFY10A | Solyc01g103600.2 | ||
| AthTIFY11B | AT1G72450.1 | SLTIFY10b | Solyc03g122190.2 | ||
| AthTIFY10B | AT1G74950.1 | SLTIFY8 | Solyc06g065650.2 | ||
| AthTIFY3A | AT3G43440.1 | SLTIFY10b-A | Solyc07g042170.2 | ||
| AthTIFY8 | AT4G32570.1 | SLTIFY5A-a | Solyc08g036620.2 | ||
| AthTIFY3B | AT5G20900.1 | SLTIFY5A-b | Solyc08g036640.2 | ||
| 水稻 Oryza sativa | OsTIFY11a | Os03t0180800-01 | SLTIFY5A-c | Solyc08g036660.2 | |
| OsTIFY11c | Os03t0180900-01 | SLTIFY10b-B | Solyc12g009220.1 | ||
| OsTIFY11b | Os03t0181100-01 | SLTIFY10b-C | Solyc12g049400.1 | ||
| OsTIFY11g | Os03t0396500-00 | 葡萄 Vitis vinifera | VvTIFY5A | VIT_04s0008g00110.t01 | |
| OsTIFY10a | Os03t0402800-01 | VvTIFY8 | VIT_04s0008g04950.t01 | ||
| OsTIFY3 | Os04t0653000-01 | VvTIFY10A | VIT_09s0002g00890.t01 | ||
| OsTIFY5 | Os07t0153000-00 | VvTIFY5A-a | VIT_10s0003g03790.t01 | ||
| OsTIFY10b | Os07t0615200-01 | VvTIFY5A-b | VIT_10s0003g03800.t01 | ||
| OsTIFY10c | Os09t0439200-01 | VvTIFY5A-c | VIT_10s0003g03810.t01 | ||
| OsTIFY11e | Os10t0391400-01 | VvTIFY5A-d | VIT_01s0011g05560.t01 | ||
| OsTIFY11d | Os10t0392400-01 | VvTIFY10A-a | VIT_11s0016g00710.t01 | ||
| VvTIFY10A-b | VIT_12s0035g00900.t01 |
| 基因 Gene | 基因号 Gene ID | 基因组位置 Genomic position | 开放阅读框/bp ORF | 蛋白大小/aa Protein size | 外显子 Number of exons |
|---|---|---|---|---|---|
| MdTIFY1 | MD02G1096100 | Chr02:7600659..7602728+ | 843 | 280 | 5 |
| MdTIFY2 | MD15G1220400 | Chr15:17843310..17845336+ | 837 | 278 | 5 |
| MdTIFY3 | MD14G1238100 | Chr14:31732049..31736399- | 1 134 | 377 | 7 |
| MdTIFY4 | MD16G1020800 | Chr16:1492513..1504489+ | 1 128 | 375 | 7 |
| MdTIFY5 | MD06G1228900 | Chr06:35860218..35864674- | 1 188 | 395 | 7 |
| MdTIFY6 | MD15G1225800 | Chr15:18345401..18348150+ | 621 | 206 | 5 |
| MdTIFY7 | MD02G1105900 | Chr02:8591750 8594168 | 633 | 210 | 5 |
| MdTIFY8 | MD13G1030000 | Chr13:2164003..2166858+ | 1 149 | 382 | 7 |
| MdTIFY9 | MD09G1178600 | Chr09:15274688..15275998- | 615 | 204 | 2 |
| MdTIFY10 | MD15G1434400 | Chr15:53470255..53474071+ | 810 | 269 | 5 |
| MdTIFY11 | MD16G1199200 | Chr16:18010085..18014820+ | 1 026 | 341 | 9 |
| MdTIFY12 | MD13G1199500 | Chr13:17721679..17726448+ | 1 050 | 349 | 9 |
| MdTIFY13 | MD08G1137500 | Chr08:13179781..13184749+ | 1 338 | 445 | 8 |
| MdTIFY14 | MD15G1116100 | Chr15:8188285..8192990+ | 1 350 | 449 | 7 |
| MdTIFY15 | MD16G1127400 | Chr16:9274777..9277069+ | 636 | 211 | 4 |
| MdTIFY16 | MD13G1127100 | Chr15:9515820..9518542+ | 630 | 209 | 4 |
表2 苹果TIFY家族成员基本信息
Table 2 Basic information of theTIFY family in apple plant
| 基因 Gene | 基因号 Gene ID | 基因组位置 Genomic position | 开放阅读框/bp ORF | 蛋白大小/aa Protein size | 外显子 Number of exons |
|---|---|---|---|---|---|
| MdTIFY1 | MD02G1096100 | Chr02:7600659..7602728+ | 843 | 280 | 5 |
| MdTIFY2 | MD15G1220400 | Chr15:17843310..17845336+ | 837 | 278 | 5 |
| MdTIFY3 | MD14G1238100 | Chr14:31732049..31736399- | 1 134 | 377 | 7 |
| MdTIFY4 | MD16G1020800 | Chr16:1492513..1504489+ | 1 128 | 375 | 7 |
| MdTIFY5 | MD06G1228900 | Chr06:35860218..35864674- | 1 188 | 395 | 7 |
| MdTIFY6 | MD15G1225800 | Chr15:18345401..18348150+ | 621 | 206 | 5 |
| MdTIFY7 | MD02G1105900 | Chr02:8591750 8594168 | 633 | 210 | 5 |
| MdTIFY8 | MD13G1030000 | Chr13:2164003..2166858+ | 1 149 | 382 | 7 |
| MdTIFY9 | MD09G1178600 | Chr09:15274688..15275998- | 615 | 204 | 2 |
| MdTIFY10 | MD15G1434400 | Chr15:53470255..53474071+ | 810 | 269 | 5 |
| MdTIFY11 | MD16G1199200 | Chr16:18010085..18014820+ | 1 026 | 341 | 9 |
| MdTIFY12 | MD13G1199500 | Chr13:17721679..17726448+ | 1 050 | 349 | 9 |
| MdTIFY13 | MD08G1137500 | Chr08:13179781..13184749+ | 1 338 | 445 | 8 |
| MdTIFY14 | MD15G1116100 | Chr15:8188285..8192990+ | 1 350 | 449 | 7 |
| MdTIFY15 | MD16G1127400 | Chr16:9274777..9277069+ | 636 | 211 | 4 |
| MdTIFY16 | MD13G1127100 | Chr15:9515820..9518542+ | 630 | 209 | 4 |
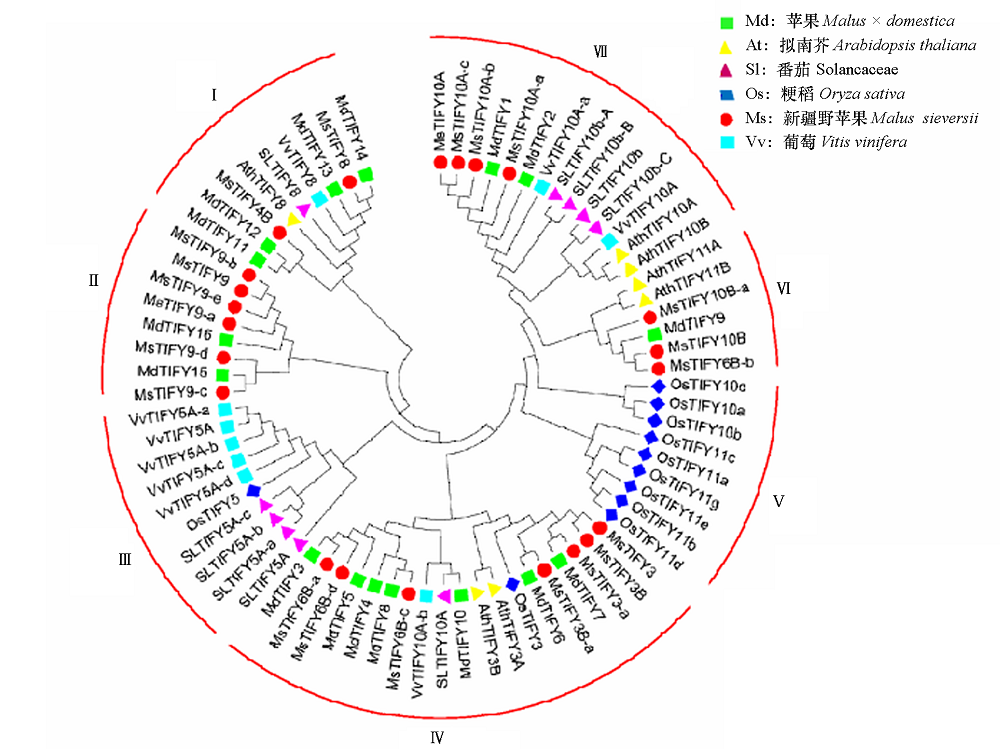
图1 苹果、拟南芥、番茄、粳稻、新疆野苹果、葡萄TIFY基因的进化树分析
Fig. 1 Phylogenetic tree analysis of TIFY genes in apple,Arabidopsis,tomato,japonica rice,Xinjiang wild apple,and grape
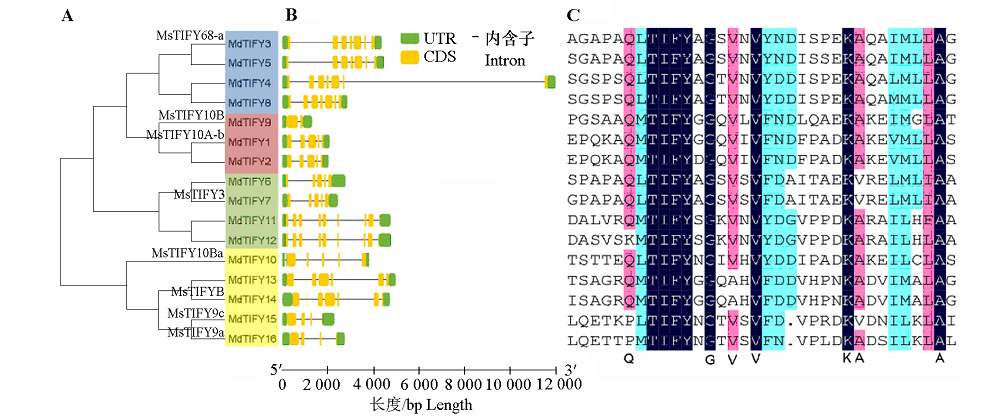
图2 苹果TIFY家族成员结构分析 A:系统进化分析,不同颜色代表聚类为一组;B:基因的外显子—内含子结构;C:氨基酸序列比对分析。
Fig. 2 Analysis of TIFY family in apple A:Phylogenetic analysis,different colors represent a clustered group;B:The exon-intron structure of the genes;C:Alignment analysis of amino acid sequence.
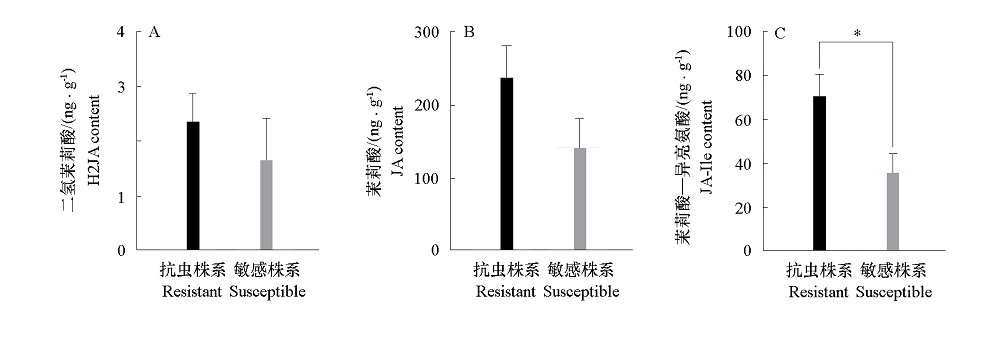
图5 新疆野苹果自然条件下茉莉素组分差异 * 表示P < 0.05显著水平。
Fig. 5 Differences of jasmin components under natural conditions in Xinjiang wild apple * Significance level of P < 0.05.
| [1] |
Bai Y, Meng Y, Huang D, Qi Y, Chen M. 2011. Origin and evolutionary analysis of the plant-specific TIFY transcription factor family. Genomics, 98(2):128-136.
doi: 10.1016/j.ygeno.2011.05.002 URL |
| [2] |
Barah P, Bones A M. 2015. Multidimensional approaches for studying plant defence against insects:from ecology to omics and synthetic biology. Journal of Experimental Botany, 66(2):479-493.
doi: 10.1093/jxb/eru489 URL |
| [3] |
Daccord N, Celton J M, Linsmith G, Becker C, Choisne N, Schijlen E, van de Geest H, Bianco L, Micheletti D, Velasco R, Di Pierro E A, Gouzy J, Rees D J G, Guerif P, Muranty H, Durel C E, Laurens F, Lespinasse Y, Gaillard S, Aubourg S, Quesneville H, Weigel D, van de Weg E, Troggio M, Bucher E. 2017. High-quality de novo assembly of the apple genome and methylome dynamics of early fruit development. Nature Genetics, 49(7):1099-1106.
doi: 10.1038/ng.3886 URL |
| [4] |
Dahl C C V, Baldwin I T. 2004. Methyl jasmonate and cis-jasmone do not dispose of the herbivore-induced jasmonate burst in Nicotiana attenuata. Physiologia Plantarum, 120(3):474-481.
doi: 10.1111/ppl.2004.120.issue-3 URL |
| [5] |
Dong Q, Zhao S, Duan D, Tian Y, Wang Y, Mao K, Zhou Z, Ma F. 2018. Structural and functional analyses of genes encoding VQ proteins in apple. Plant Science, 272:208-219.
doi: 10.1016/j.plantsci.2018.04.029 URL |
| [6] |
Ebel C, BenFeki A, Hanin M, Solano R, Chini A. 2018. Characterization of wheat(Triticum aestivum)TIFY family and role of Triticum durum TdTIFY11a in salt stress tolerance. PLoS ONE, 13(7):e0200566.
doi: 10.1371/journal.pone.0200566 URL |
| [7] | Engelberth J, Seidl-Adams I, Schultz J C, Tumlinson J H. 2007. Insect elicitors and exposure to green leafy volatiles differentially upregulate major octadecanoids and transcripts of 12-Oxo phytodienoic Acid Reductases in Zea mays. Molecular Plant, 20(6):707-716. |
| [8] |
Howe G A, Jander G. 2008. Plant immunity to insect herbivores. Annual Review of Plant Biology, 59:41-66.
doi: 10.1146/annurev.arplant.59.032607.092825 URL |
| [9] |
Hu C, Wei C, Ma Q, Dong H, Shi K, Zhou Y, Foyer C H, Yu J. 2021. Eethylene response factor 15 and 16 trigger jasmonate biosynthesis in tomato during herbivore resistance. Plant Physiology,Doi: 10.1093/plphys/kiaa089.
doi: 10.1093/plphys/kiaa089 |
| [10] |
Koo A J, Gao X, Jones A D, Howe G A. 2009. A rapid wound signal activates the systemic synthesis of bioactive jasmonates in Arabidopsis. Plant Journal, 59(6):974-986.
doi: 10.1111/tpj.2009.59.issue-6 URL |
| [11] | Li R, Jin J, Xu J, Wang L, Li J, Lou Y, Baldwin I T. 2020. Long non‐coding RNAs associate with jasmonate‐mediated plant defence against herbivores. Plant,Cell & Environment, pce: 13952. |
| [12] | Li X, Yin X, Wang H, Li J, Guo C, Gao H, Zheng Y, Fan C, Wang X. 2014. Genome-wide identification and analysis of the apple(Malus × domestica Borkh.)TIFY gene family. Tree Genetics & Genomes, 11(1):808. |
| [13] |
Liu Q, Wang X, Tzin V, Romeis J, Peng Y, Li Y. 2016. Combined transcriptome and metabolome analyses to understand the dynamic responses of rice plants to attack by the rice stem borer Chilo suppressalis(Lepidoptera:Crambidae). BMC Plant Biology, 16(1):259.
doi: 10.1186/s12870-016-0946-6 URL |
| [14] |
Mao K, Dong Q, Li C, Liu C, Ma F. 2017a. Genome wide identification and characterization of apple bHLH transcription factors and expression analysis in response to drought and salt stress. Frontiers in Plant Science, 8:480.
doi: 10.3389/fpls.2017.00480 pmid: 28443104 |
| [15] |
Mao Y B, Liu Y Q, Chen D Y, Chen F Y, Fang X, Hong G J, Wang L J, Wang J W, Chen X Y. 2017b. Jasmonate response decay and defense metabolite accumulation contributes to age-regulated dynamics of plant insect resistance. Nature Communications, 8:13925.
doi: 10.1038/ncomms13925 URL |
| [16] | Mei Chuang, Yan Peng, Ai Shajiang, Ma Kai, Han Liqun, Xu Zheng, Zhong Haixia, Diao Yongqiang, Wang Jixun. 2016. The relationship between bark thickness and branch roughnesson Agrilus mali damage in Xinjiang wild apple. Journal of Agricultural Science and Technology, 18(4):24-30. (in Chinese) |
| 梅闯, 闫鹏, 艾沙江·买买提, 马凯, 韩立群, 许正, 钟海霞, 刁永强, 王继勋. 2016. 新疆野苹果(Malus sieversii)受苹小吉丁虫危害程度与树皮厚度、径阶的关系. 中国农业科技导报, 18(4):24-30. | |
| [17] | Mei Chuang, Yan Peng, Ma Kai, Han Liqun, Xu Zheng, Lu Chunsheng, Fan Dingyu, Ai Shajiang, Wang Jixun. 2015. Agrilus mali Matsumura resistance of different type of Xinjiang wild apple. Xinjiang Agricultural Sciences, 52(10):1859-1865. (in Chinese). |
| 梅闯, 闫鹏, 马凯, 韩立群, 许正, 卢春生, 樊丁宇, 艾沙江, 王继勋. 2015. 新疆野苹果不同类型单株对苹果小吉丁虫的抗性差异. 新疆农业科学, 52(10):1859-1865. | |
| [18] |
Mei C, Yang J, Yan P, Li N, Ma K, Mamat A, Han L, Dong Q, Mao K, Ma F, Wang J. 2020. Full-length transcriptome and targeted metabolome analyses provide insights into defense mechanisms of Malus sieversii against Agrilus mali. PeerJ, 8:e8992.
doi: 10.7717/peerj.8992 URL |
| [19] |
Staswick P E. 2008. JAZing up jasmonate signaling. Trends in Plant Science, 13(2):66-71.
doi: 10.1016/j.tplants.2007.11.011 URL |
| [20] |
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. 2011. MEGA5:molecular evolutionary genetics analysis using maximum likelihood,evolutionary distance,and maximum parsimony methods. Molecular Biology and Evolution, 28(10):2731-2739.
doi: 10.1093/molbev/msr121 URL |
| [21] |
Thines B, Katsir L, Melotto M, Niu Y, Mandaokar A, Liu G, Nomura K, He S Y, Howe G A, Browse J. 2007. JAZ repressor proteins are targets of the SCF(COI1)complex during jasmonate signalling. Nature, 448(7154):661-665.
pmid: 17637677 |
| [22] |
Thireault C, Shyu C, Yoshida Y, St Aubin B, Campos M L, Howe G A. 2015. Repression of jasmonate signaling by a non-TIFY JAZ protein in Arabidopsis. The Plant journal, 82(4):669-679.
doi: 10.1111/tpj.12841 pmid: 25846245 |
| [23] |
Vanholme B, Grunewald W, Bateman A, Kohchi T, Gheysen G. 2007. The tify family previously known as ZIM. Trends in Plant Science, 12(6):239-244.
doi: 10.1016/j.tplants.2007.04.004 URL |
| [24] |
Velasco R, Zharkikh A, Affourtit J, Dhingra A, Cestaro A, Kalyanaraman A, Fontana P, Bhatnagar S K, Troggio M, Pruss D, Salvi S, Pindo M, Baldi P, Castelletti S, Cavaiuolo M, Coppola G, Costa F, Cova V, Dal Ri A, Goremykin V, Komjanc M, Longhi S, Magnago P, Malacarne G, Malnoy M, Micheletti D, Moretto M, Perazzolli M, Si-Ammour A, Vezzulli S, Zini E, Eldredge G, Fitzgerald L M, Gutin N, Lanchbury J, Macalma T, Mitchell J T, Reid J, Wardell B, Kodira C, Chen Z, Desany B, Niazi F, Palmer M, Koepke T, Jiwan D, Schaeffer S, Krishnan V, Wu C, Chu V T, King S T, Vick J, Tao Q, Mraz A, Stormo A, Stormo K, Bogden R, Ederle D, Stella A, Vecchietti A, Kater M M, Masiero S, Lasserre P, Lespinasse Y, Allan A C, Bus V, Chagne D, Crowhurst R N, Gleave A P, Lavezzo E, Fawcett J A, Proost S, Rouze P, Sterck L, Toppo S, Lazzari B, Hellens R P, Durel C E, Gutin A, Bumgarner R E, Gardiner S E, Skolnick M, Egholm M, Van de Peer Y, Salamini F, Viola R. 2010. The genome of the domesticated apple(Malus × domestica Borkh.). Nature Genetics, 42(10):833-839.
doi: 10.1038/ng.654 URL |
| [25] | Wang Ping, Lu Shixiong, Liang Guoping, Ma Zonghuan, Li Wenfang, Mao Juan, Chen Baihong. 2019. Bioinformatics identification and expression analysis of Trihelix transcription factor family in apple. Acta Horticulturae Sinica, 46(11):2082-2098. (in Chinese). |
| 王萍, 卢世雄, 梁国平, 马宗桓, 李文芳, 毛娟, 陈佰鸿. 2019. 苹果Trihelix转录因子家族生物信息学鉴定与基因表达分析. 园艺学报, 46(11):2082-2098. | |
| [26] |
Wang Y, Pan F, Chen D M, Chu W Y, Liu H L, Xiang Y. 2017. Genome-wide identification and analysis of the Populus trichocarpa TIFY gene family. Plant Physiology and Biochemistry, 115:360-371.
doi: 10.1016/j.plaphy.2017.04.015 URL |
| [27] |
War A R, Paulraj M G, Ahmad T, Buhroo A A, Hussain B, Ignacimuthu S, Sharma H C. 2012. Mechanisms of plant defense against insect herbivores. Plant Signal Behav, 7(10):1306-1320.
doi: 10.4161/psb.21663 URL |
| [28] | Xia W X, Yu H Y, Cao P, Luo J, Wang N. 2017. Identification of TIFY family genes and analysis of their expression profiles in response to phytohormone treatments and Melampsora larici-populina infection in poplar. Frontiers in Plant Science, 8:493. |
| [29] |
Xie Y, Chen P, Yan Y, Bao C, Li X, Wang L, Shen X, Li H, Liu X, Niu C, Zhu C, Fang N, Shao Y, Zhao T, Yu J, Zhu J, Xu L, van Nocker S, Ma F, Guan Q. 2018. An atypical R2R3 MYB transcription factor increases cold hardiness by CBF-dependent and CBF-independent pathways in apple. The New Phytologist, 218(1):201-218.
doi: 10.1111/nph.14952 URL |
| [30] | Xu G X, Guo C C, Shan H Y, Kong H Z. 2012. Divergence of duplicate genes in exon-intron structure. Proceedings of the National Academy of Sciences of the United States of America, 109(4):1187-1192. |
| [31] |
Ye H Y, Du H, Tang N, Li X H, Xiong L Z. 2009. Identification and expression profiling analysis of TIFY family genes involved in stress and phytohormone responses in rice. Plant Molecular Biology, 71(3):291-305.
doi: 10.1007/s11103-009-9524-8 URL |
| [32] |
Zhang Y, Gao M, Singer S D, Fei Z, Wang H, Wang X. 2012. Genome-wide identification and analysis of the TIFY gene family in grape. PLoS ONE, 7(9):e44465.
doi: 10.1371/journal.pone.0044465 URL |
| [1] | 于婷婷, 李 欢, 宁源生, 宋建飞, 彭璐琳, 贾竣淇, 张玮玮, 杨洪强. 苹果GRAS全基因组鉴定及其对生长素的响应分析[J]. 园艺学报, 2023, 50(2): 397-409. |
| [2] | 翟含含, 翟宇杰, 田义, 张叶, 杨丽, 温陟良, 陈海江. 桃SAUR家族基因分析及PpSAUR5功能鉴定[J]. 园艺学报, 2023, 50(1): 1-14. |
| [3] | 赵雪艳, 王琪, 王莉, 王方圆, 王庆, 李艳. 基于比较转录组的延胡索组织差异性表达分析[J]. 园艺学报, 2023, 50(1): 177-187. |
| [4] | 丁志杰, 包金波, 柔鲜古丽, 朱甜甜, 李雪丽, 苗浩宇, 田新民. 新疆野苹果与‘元帅’‘金冠’的叶绿体基因组比对研究[J]. 园艺学报, 2022, 49(9): 1977-1990. |
| [5] | 高彦龙, 吴玉霞, 张仲兴, 王双成, 张瑞, 张德, 王延秀. 苹果ELO家族基因鉴定及其在低温胁迫下的表达分析[J]. 园艺学报, 2022, 49(8): 1621-1636. |
| [6] | 刘金明, 郭彩华, 袁星, 亢超, 全绍文, 牛建新. 梨Dof家族基因鉴定及其在宿存与脱落萼片中的表达分析[J]. 园艺学报, 2022, 49(8): 1637-1649. |
| [7] | 邱子文, 刘林敏, 林永盛, 林晓洁, 李永裕, 吴少华, 杨超. 千层金MbEGS基因的克隆与功能分析[J]. 园艺学报, 2022, 49(8): 1747-1760. |
| [8] | 郑林, 王帅, 刘语诺, 杜美霞, 彭爱红, 何永睿, 陈善春, 邹修平. 柑橘响应黄龙病菌侵染的NAC基因的克隆及表达分析[J]. 园艺学报, 2022, 49(7): 1441-1457. |
| [9] | 马维峰, 李艳梅, 马宗桓, 陈佰鸿, 毛娟. 苹果POD家族基因的鉴定与MdPOD15的功能分析[J]. 园艺学报, 2022, 49(6): 1181-1199. |
| [10] | 张凯, 麻明英, 王萍, 李益, 金燕, 盛玲, 邓子牛, 马先锋. 柑橘HSP20家族基因鉴定及其响应溃疡病菌侵染表达分析[J]. 园艺学报, 2022, 49(6): 1213-1232. |
| [11] | 梁晨, 孙如意, 向锐, 孙艺萌, 师校欣, 杜国强, 王莉. 葡萄生长调控因子GRF家族基因的鉴定及表达分析[J]. 园艺学报, 2022, 49(5): 995-1007. |
| [12] | 肖学宸, 刘梦雨, 蒋梦琦, 陈燕, 薛晓东, 周承哲, 吴兴健, 吴君楠, 郭寅生, 叶开温, 赖钟雄, 林玉玲. 龙眼褪黑素合成途径SNAT、ASMT和COMT家族基因鉴定及表达分析[J]. 园艺学报, 2022, 49(5): 1031-1046. |
| [13] | 高玮林, 张力曼, 薛超玲, 张垚, 刘孟军, 赵锦. 枣E类MADS基因在花和果中的表达及其蛋白互作研究[J]. 园艺学报, 2022, 49(4): 739-748. |
| [14] | 刘梦雨, 蒋梦琦, 陈燕, 张舒婷, 薛晓东, 肖学宸, 赖钟雄, 林玉玲. 龙眼GDSL酯酶/脂肪酶基因的全基因组鉴定及表达分析[J]. 园艺学报, 2022, 49(3): 597-612. |
| [15] | 姜翠翠, 方智振, 周丹蓉, 林炎娟, 叶新福. ‘芙蓉李’糖转运蛋白家族基因鉴定及表达分析[J]. 园艺学报, 2022, 49(2): 252-264. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
版权所有 © 2012 《园艺学报》编辑部 京ICP备10030308号-2 国际联网备案号 11010802023439
编辑部地址: 北京市海淀区中关村南大街12号中国农业科学院蔬菜花卉研究所 邮编: 100081
电话: 010-82109523 E-Mail: yuanyixuebao@126.com
技术支持:北京玛格泰克科技发展有限公司