
Acta Horticulturae Sinica ›› 2021, Vol. 48 ›› Issue (11): 2251-2261.doi: 10.16420/j.issn.0513-353x.2021-0397
• Original article • Previous Articles Next Articles
YANG Ting1, XUE Zhenzhen1, LI Na2, LANG Xiaoan2, LI Lingfei2,*( ), ZHONG Chunmei1,*(
), ZHONG Chunmei1,*( )
)
Received:2021-05-17
Revised:2021-09-15
Published:2021-12-02
Contact:
LI Lingfei,ZHONG Chunmei
E-mail:lingfei_li@szbg.ac.cn;zhongchunmei@scau.edu.cn
CLC Number:
YANG Ting, XUE Zhenzhen, LI Na, LANG Xiaoan, LI Lingfei, ZHONG Chunmei. Reference Genes Selection and Validation in Begonia masoniana Leaves of Different Developmental Stages[J]. Acta Horticulturae Sinica, 2021, 48(11): 2251-2261.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2021-0397
| 时期 Stage | 叶片长/cm Leaf length | 叶片宽/cm Leaf width | 生长天数/d Growth days | 特征描述 Feature description |
|---|---|---|---|---|
| S1 | 1.2 ~ 1.6 | 1.0 ~ 1.2 | 2 ~ 3 | 叶片卷曲,表皮毛较长,全叶呈红色 Leaf curly,hirsute-villous,whole leaf is red |
| S2 | 2.2 ~ 2.8 | 1.8 ~ 2.2 | 6 ~ 8 | 叶片舒张,表皮毛长,叶片质地较厚,全叶呈红色 Leaf expansion,hirsute-villous,thick,whole leaf is red |
| S3 | 4.1 ~ 5.1 | 3.1 ~ 3.7 | 10 ~ 12 | 叶斑初显,沿主叶脉规则分布,全叶呈红色 Leaf variegation appearing,distributed along the main vein,whole leaf is red |
| S4 | 6.8 ~ 7.4 | 4.6 ~ 6.2 | 15 ~ 18 | 叶斑颜色继续加深,叶斑以外部分叶色偏黄 Leaf variegation deepen,other parts of leaf show yellowish |
| S5 | 10.2 ~ 12.8 | 7.7 ~ 8.7 | 23 ~ 25 | 叶斑明显,叶斑以外部分叶色逐渐转绿 Leaf variegation begin obvious,other parts of leaf turn green |
| S6 | 12.7 ~ 15.0 | 9.2 ~ 11.9 | 33 ~ 35 | 叶片继续生长,叶斑以外部分叶色全绿 Leaf continues to grow,and other parts except leaf variegation are all green |
| S7 | 20.1 ~ 21.5 | 15.2 ~ 16.1 | 48 ~ 52 | 叶片大小达到最大 Leaf size reaches maximum |
Table 1 Developmental stages of leaves in Begonia masoniana
| 时期 Stage | 叶片长/cm Leaf length | 叶片宽/cm Leaf width | 生长天数/d Growth days | 特征描述 Feature description |
|---|---|---|---|---|
| S1 | 1.2 ~ 1.6 | 1.0 ~ 1.2 | 2 ~ 3 | 叶片卷曲,表皮毛较长,全叶呈红色 Leaf curly,hirsute-villous,whole leaf is red |
| S2 | 2.2 ~ 2.8 | 1.8 ~ 2.2 | 6 ~ 8 | 叶片舒张,表皮毛长,叶片质地较厚,全叶呈红色 Leaf expansion,hirsute-villous,thick,whole leaf is red |
| S3 | 4.1 ~ 5.1 | 3.1 ~ 3.7 | 10 ~ 12 | 叶斑初显,沿主叶脉规则分布,全叶呈红色 Leaf variegation appearing,distributed along the main vein,whole leaf is red |
| S4 | 6.8 ~ 7.4 | 4.6 ~ 6.2 | 15 ~ 18 | 叶斑颜色继续加深,叶斑以外部分叶色偏黄 Leaf variegation deepen,other parts of leaf show yellowish |
| S5 | 10.2 ~ 12.8 | 7.7 ~ 8.7 | 23 ~ 25 | 叶斑明显,叶斑以外部分叶色逐渐转绿 Leaf variegation begin obvious,other parts of leaf turn green |
| S6 | 12.7 ~ 15.0 | 9.2 ~ 11.9 | 33 ~ 35 | 叶片继续生长,叶斑以外部分叶色全绿 Leaf continues to grow,and other parts except leaf variegation are all green |
| S7 | 20.1 ~ 21.5 | 15.2 ~ 16.1 | 48 ~ 52 | 叶片大小达到最大 Leaf size reaches maximum |
| 基因 Gene | 引物序列(5′-3′) Primer sequence |
|---|---|
| CHS | F:TGCACCACAAGCGGAGTCGA;R:AGCAAGATCCTTGGCGACCCT |
| F3H | F:TTCCAGAACCCAGCGCCAGA;R:TCAGCCTGGCAAGCTCAAGGT |
| F3’H | F:GCGACTTCATTCCGGCGCTT;R:ACCGCCGATCCGCTTGTGTT |
| FLS | F:TGAGCAGCCCGGAATCACCA;R:TGTCGCACACCACACGTTCCT |
| DFR | F:AGACATTGGCAGAGCAGGCG;R:AGTGATCAGGCTCGGTGGCA |
| UFGT | F:TGAATGCCGCCCCAGAAAGCT;R:CGGCGAACCACAAGAAAGCGT |
Table 2 Primer sequences for some key genes in anthocyanin synthesis pathway derived from qRT-PCR
| 基因 Gene | 引物序列(5′-3′) Primer sequence |
|---|---|
| CHS | F:TGCACCACAAGCGGAGTCGA;R:AGCAAGATCCTTGGCGACCCT |
| F3H | F:TTCCAGAACCCAGCGCCAGA;R:TCAGCCTGGCAAGCTCAAGGT |
| F3’H | F:GCGACTTCATTCCGGCGCTT;R:ACCGCCGATCCGCTTGTGTT |
| FLS | F:TGAGCAGCCCGGAATCACCA;R:TGTCGCACACCACACGTTCCT |
| DFR | F:AGACATTGGCAGAGCAGGCG;R:AGTGATCAGGCTCGGTGGCA |
| UFGT | F:TGAATGCCGCCCCAGAAAGCT;R:CGGCGAACCACAAGAAAGCGT |
| 基因 Gene | 基因名称 Gene name | 引物序列(5′-3′) Primer sequence | 斜率 k | 扩增效率/% E | 相关系数 R2 |
|---|---|---|---|---|---|
| PP2A | Serine/threonine-PP2A catalytic subunit | F:TCCTGATGGAGTGCAAGCCGT R:ACAGACTGTCACCGGGCACT | -3.31 | 100.50 | 0.9984 |
| EF1α | Elongation factor 1α | F:TGCAGCTTGGCTCGGCAGAT R:TGGCAGGGGCGTGATCTAACA | -3.41 | 96.45 | 0.9996 |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase | F:TGACGTTGTGGCAGCTCAGGT R:TCCGGCCAATGCGTCCGAAT | -3.32 | 100.08 | 0.9961 |
| CYP | Cyclophilin | F:ACGGTGCCAAGTTTGCCGAC R:AACTGTGAGCCGTTGGTTCCG | -3.34 | 99.25 | 0.9988 |
| elF4A | Eukaryotic translation initiation factor 4A-1 | F:ACCTCCATCGAATCGGCCGT R:TGCTGGCAACTCCTCCACCA | -3.20 | 105.35 | 0.9995 |
| ACT2 | Actin 2 | F:TTGCTCACAGAGGCACCGCT R:ACCGGTAGTACGACCACTGGCA | -3.33 | 99.66 | 0.9935 |
| 18S | 18S Ribosomal RNA | F:TGACGGATCGCACGGCCTTT R:TTCTCCGTCACCCGTCACCA | -3.45 | 94.92 | 0.9966 |
| UBQ10 | Polyubiquitin 10 | F:ACGCTTCGAGGTAGGTTTCTTGT R:ACGCATCGAAAACAACAACCGCA | -3.50 | 93.07 | 0.9991 |
| ACT7 | Actin 7 | F:TCGTGCTTGGAGGTTCGGCT R:CCAGCCTTCACCATTCCAGTTCCA | -3.27 | 102.21 | 0.9964 |
| PPR | Pentatricopeptide repeat superfamily protein | F:AGAAGTGCGAGTGGCTACCT R:CACAAACCCGCCGATCAGAGT | -3.19 | 105.82 | 0.9996 |
| HDH | Histidinol dehydrogenase | F:ATGTCCCTGGGGGTACTGCTGT R:TGCCATCCTGACTTGGGGGA | -3.36 | 98.44 | 0.9997 |
Table 3 Primer sequences and amplification parameters for 11 candidate reference genes of Begonia masoniana
| 基因 Gene | 基因名称 Gene name | 引物序列(5′-3′) Primer sequence | 斜率 k | 扩增效率/% E | 相关系数 R2 |
|---|---|---|---|---|---|
| PP2A | Serine/threonine-PP2A catalytic subunit | F:TCCTGATGGAGTGCAAGCCGT R:ACAGACTGTCACCGGGCACT | -3.31 | 100.50 | 0.9984 |
| EF1α | Elongation factor 1α | F:TGCAGCTTGGCTCGGCAGAT R:TGGCAGGGGCGTGATCTAACA | -3.41 | 96.45 | 0.9996 |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase | F:TGACGTTGTGGCAGCTCAGGT R:TCCGGCCAATGCGTCCGAAT | -3.32 | 100.08 | 0.9961 |
| CYP | Cyclophilin | F:ACGGTGCCAAGTTTGCCGAC R:AACTGTGAGCCGTTGGTTCCG | -3.34 | 99.25 | 0.9988 |
| elF4A | Eukaryotic translation initiation factor 4A-1 | F:ACCTCCATCGAATCGGCCGT R:TGCTGGCAACTCCTCCACCA | -3.20 | 105.35 | 0.9995 |
| ACT2 | Actin 2 | F:TTGCTCACAGAGGCACCGCT R:ACCGGTAGTACGACCACTGGCA | -3.33 | 99.66 | 0.9935 |
| 18S | 18S Ribosomal RNA | F:TGACGGATCGCACGGCCTTT R:TTCTCCGTCACCCGTCACCA | -3.45 | 94.92 | 0.9966 |
| UBQ10 | Polyubiquitin 10 | F:ACGCTTCGAGGTAGGTTTCTTGT R:ACGCATCGAAAACAACAACCGCA | -3.50 | 93.07 | 0.9991 |
| ACT7 | Actin 7 | F:TCGTGCTTGGAGGTTCGGCT R:CCAGCCTTCACCATTCCAGTTCCA | -3.27 | 102.21 | 0.9964 |
| PPR | Pentatricopeptide repeat superfamily protein | F:AGAAGTGCGAGTGGCTACCT R:CACAAACCCGCCGATCAGAGT | -3.19 | 105.82 | 0.9996 |
| HDH | Histidinol dehydrogenase | F:ATGTCCCTGGGGGTACTGCTGT R:TGCCATCCTGACTTGGGGGA | -3.36 | 98.44 | 0.9997 |
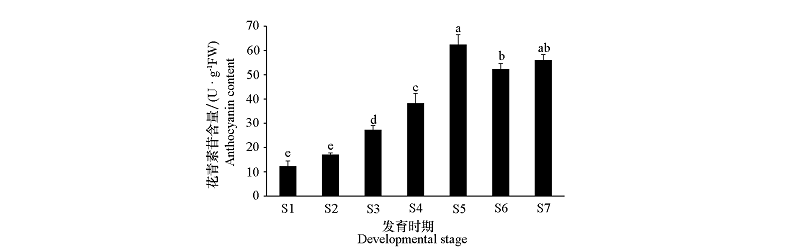
Fig. 2 Anthocyanin content from Begonia masoniana leaves of different developmental stages Different lowercase letters indicate significant differences among different periods at 0.05 level.
| 基因 Gene | 几何平均数 GM | 算数平均数 AM | 最小值 Min | 最大值 Max | 标准偏差 SD | 稳定性排序 Stability rank |
|---|---|---|---|---|---|---|
| PP2A | 24.02 | 24.02 | 23.94 | 24.26 | 0.07 | 1 |
| ACT7 | 21.82 | 21.82 | 21.64 | 21.94 | 0.10 | 2 |
| elF4A | 24.26 | 24.26 | 23.86 | 24.54 | 0.14 | 3 |
| PPR | 26.54 | 26.54 | 26.29 | 26.97 | 0.15 | 4 |
| HDH | 24.88 | 24.88 | 24.55 | 25.23 | 0.19 | 5 |
| ACT2 | 19.45 | 19.45 | 19.06 | 19.83 | 0.26 | 6 |
| EF1α | 24.24 | 24.25 | 23.66 | 24.97 | 0.37 | 7 |
| 18S | 23.08 | 23.09 | 22.53 | 23.77 | 0.42 | 8 |
| CYP | 21.51 | 21.53 | 20.28 | 22.87 | 0.77 | 9 |
| UBQ10 | 24.28 | 24.30 | 22.75 | 26.02 | 0.82 | 10 |
| GAPDH | 21.50 | 21.55 | 19.88 | 23.89 | 1.27 | 11 |
Table 4 Expression stabilities of eleven candidate reference genes analyzed by BestKeeper
| 基因 Gene | 几何平均数 GM | 算数平均数 AM | 最小值 Min | 最大值 Max | 标准偏差 SD | 稳定性排序 Stability rank |
|---|---|---|---|---|---|---|
| PP2A | 24.02 | 24.02 | 23.94 | 24.26 | 0.07 | 1 |
| ACT7 | 21.82 | 21.82 | 21.64 | 21.94 | 0.10 | 2 |
| elF4A | 24.26 | 24.26 | 23.86 | 24.54 | 0.14 | 3 |
| PPR | 26.54 | 26.54 | 26.29 | 26.97 | 0.15 | 4 |
| HDH | 24.88 | 24.88 | 24.55 | 25.23 | 0.19 | 5 |
| ACT2 | 19.45 | 19.45 | 19.06 | 19.83 | 0.26 | 6 |
| EF1α | 24.24 | 24.25 | 23.66 | 24.97 | 0.37 | 7 |
| 18S | 23.08 | 23.09 | 22.53 | 23.77 | 0.42 | 8 |
| CYP | 21.51 | 21.53 | 20.28 | 22.87 | 0.77 | 9 |
| UBQ10 | 24.28 | 24.30 | 22.75 | 26.02 | 0.82 | 10 |
| GAPDH | 21.50 | 21.55 | 19.88 | 23.89 | 1.27 | 11 |
| [1] |
Andersen C L, Jensen J L, Orntoft T F. 2004. Normalization of real-time quantitative reverse transcription-PCR data:a model-based variance estimation approach to identify genes suited for normalization,applied to bladder and colon cancer data sets. Cancer Res, 64 (15):5245-5250.
doi: 10.1158/0008-5472.CAN-04-0496 URL |
| [2] | Bai Xue. 2020. Study on genes and substances related to anthocyanin biosynthesis by low temperature and high light stress in the leaves of Begonia semperflorens[M. D. Dissertation]. Zhengzhou: Henan Agricultural University. (in Chinese) |
| 白雪. 2020. 四季秋海棠叶片在低温和高光胁迫下次生花色素苷生物合成相关基因以及物质的研究[硕士论文]. 郑州: 河南农业大学. | |
| [3] |
Bustin S A. 2002. Quantification of mRNA using real-time reverse transcription PCR(RT-PCR):trends and problems. J Mol Endocrinol, 29 (1):23-39.
pmid: 12200227 |
| [4] | Cui Weihua, Guan Kaiyun. 2013. Diversity of leaf variegation in Chinese Begonias. Plant Diversity and Resources, 35 (2):119-127. (in Chinese) |
| 崔卫华, 管开云. 2013. 中国秋海棠属植物叶片斑纹多样性研究. 植物分类与资源学报, 35 (2):119-127. | |
| [5] | de Spiegelaere W, Dern-Wieloch J, Weigel R, Schumacher V, Schorle H, Nettersheim D, Bergmann M, Brehm R, Kliesch S, Vandekerckhove L, Fink C. 2015. Reference gene validation for RT-qPCR,a note on different available software packages. PLoS ONE, 10 (3):e122515. |
| [6] | Dong Lina, Liu Yan. 2019. Supplement to Begonia L. in Flora of Guangxi. Guihaia, 39 (1):16-39. (in Chinese) |
| 董莉娜, 刘演. 2019. 《广西植物志》秋海棠属(Begonia L.)增订. 广西植物, 39 (1):16-39. | |
| [7] |
Dong Y, Qu Y, Qi R, Bai X, Tian G, Wang Y, Wang J, Zhang K. 2018. Transcriptome analysis of the biosynthesis of anthocyanins in Begonia semperflorens under low-temperature and high-light conditions. Forests, 9 (2):87.
doi: 10.3390/f9020087 URL |
| [8] |
Fu Z Z, Shang H Q, Jiang H, Gao J, Dong X Y, Wang H J, Li Y M, Wang L M, Zhang J, Shu Q Y, Chao Y C, Xu M L, Wang R, Wang L S, Zhang H C. 2020. Systematic identification of the light-quality responding anthocyanin synthesis-related transcripts in petunia petals. Horticultural Plant Journal, 6 (6):428-438.
doi: 10.1016/j.hpj.2020.11.006 URL |
| [9] |
Gutierrez L, Mauriat M, Guenin S, Pelloux J, Lefebvre J F, Louvet R, Rusterucci C, Moritz T, Guerineau F, Bellini C, van Wuytswinkel O. 2008. The lack of a systematic validation of reference genes:a serious pitfall undervalued in reverse transcription-polymerase chain reaction (RT-PCR)analysis in plants. Plant Biotechnol J, 6 (6):609-618.
doi: 10.1111/j.1467-7652.2008.00346.x pmid: 18433420 |
| [10] |
Harborne J B, Williams C A. 2000. Advances in flavonoid research since 1992. Phytochemistry, 55 (6):481-504.
pmid: 11130659 |
| [11] |
Holton T A, Cornish E C. 1995. Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell, 7 (7):1071-1083.
doi: 10.2307/3870058 URL |
| [12] |
Hughes N M, Vogelmann T C, Smith W K. 2008. Optical effects of abaxial anthocyanin on absorption of red wavelengths by understorey species:revisiting the back-scatter hypothesis. J Exp Bot, 59 (12):3435-3442.
doi: 10.1093/jxb/ern193 URL |
| [13] |
Khaldoun O A S, Abd E H. 2015. Biochemical and genetic evidences of anthocyanin biosynthesis and accumulation in a selected tomato mutant. Rendiconti Lincei, 26 (3):293-306.
doi: 10.1007/s12210-015-0446-x URL |
| [14] |
Li Y M, Zhang K M, Jin H H, Zhu L, Li Y H. 2015. Isolation and expression analysis of four putative structural genes involved in anthocyanin biosynthesis in Begonia semperflorens. J Hortic Sci Biotech, 90 (4):444-450.
doi: 10.1080/14620316.2015.11513208 URL |
| [15] |
Li Y, Qu Y, Wang Y, Bai X, Tian G, Liu Z, Li Y, Zhang K. 2019. Selection of suitable reference genes for qRT-PCR analysis of Begonia semperflorens under stress conditions. Mol Biol Rep, 46 (6):6027-6037.
doi: 10.1007/s11033-019-05038-5 URL |
| [16] |
Li W X, Wang L, He Z C, Lu Z G, Cui J W, Xu N T, Jin B, Wang L. 2020. Physiological and transcriptomic changes during autumn coloration and senescence in Ginkgo biloba leaves. Horticultural Plant Journal, 6 (6):396-408.
doi: 10.1016/j.hpj.2020.11.002 URL |
| [17] | Liang Lijun, Yang Yichen, Wang Erhuan, Xing Bingcong, Liang Zongsuo. 2018. Research progress on biosynthesis and regulation of plant anthocyanin. Journal of Anhui Agricultural Sciences, 46 (21):18-24. (in Chinese) |
| 梁立军, 杨祎辰, 王二欢, 邢丙聪, 梁宗锁. 2018. 植物花青素生物合成与调控研究进展. 安徽农业科学, 46 (21):18-24. | |
| [18] |
Livak K J, Schmittgen T D. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2-∆∆C(T) method. Methods, 25 (4):402-408.
pmid: 11846609 |
| [19] |
Maroufi A, van Bockstaele E, de Loose M. 2010. Validation of reference genes for gene expression analysis in chicory(Cichorium intybus)using quantitative real-time PCR. BMC Mol Biol, 11:15.
doi: 10.1186/1471-2199-11-15 pmid: 20156357 |
| [20] | Mei Beijian, Ai Hua. 1987. Rapid propagation of iron cross Begonia. Plant Physiology Physiol Communications,(2):27-30. (in Chinese) |
| 梅贝坚, 艾华. 1987. 铁十字秋海棠试管快速繁殖. 植物生理学通讯,(2):27-30. | |
| [21] |
Pfaffl M W, Tichopad A, Prgomet C, Neuvians T P. 2004. Determination of stable housekeeping genes,differentially regulated target genes and sample integrity:BestKeeper-Excel-based tool using pair-wise correlations. Biotechnol Lett, 26 (6):509-515.
doi: 10.1023/B:BILE.0000019559.84305.47 URL |
| [22] | Ren Yan, Zhang Weiguang. 2006. Research advances of anthocyanin. China Food Additives,(4):71-77. (in Chinese) |
| 任雁, 张惟广. 2006. 花色素苷的研究进展. 中国食品添加剂,(4):71-77. | |
| [23] |
Shi M Z, Xie D Y. 2014. Biosynthesis and metabolic engineering of anthocyanins in Arabidopsis thaliana. Recent Pat Biotechnol, 8 (1):47-60.
doi: 10.2174/1872208307666131218123538 URL |
| [24] |
Steyn W J, Wand S J E, Holcroft D M, Jacobs G. 2002. Anthocyanins in vegetative tissues:a proposed unified function in photoprotect. New Phytol, 155 (3):349-361.
doi: 10.1046/j.1469-8137.2002.00482.x pmid: 33873306 |
| [25] |
Sparvoli F, Martin C, Scienza A, Gavazzi G, Tonelli C. 1994. Cloning and molecular analysis of structural genes involved in flavonoid and stilbene biosynthesis in grape(Vitis vinifera L.). Plant Mol Biol, 24 (5):743-755.
pmid: 8193299 |
| [26] |
Tanaka Y, Sasaki N, Ohmiya A. 2008. Biosynthesis of plant pigments:anthocyanins,betalains and carotenoids. Plant J, 54 (4):733-749.
doi: 10.1111/j.1365-313X.2008.03447.x URL |
| [27] | Vandesompele J, de Preter K, Pattyn F, Poppe B, van Roy N, de Paepe A, Speleman F. 2002. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol, 3 (7):H34. |
| [28] |
Wang J, Guo M, Li Y, Wu R, Zhang K. 2018. High-throughput transcriptome sequencing reveals the role of anthocyanin metabolism in Begonia semperflorens under high light stress. Photochem Photobiol, 94 (1):105-114.
doi: 10.1111/php.2018.94.issue-1 URL |
| [29] |
Winkel-Shirley B. 2001. Flavonoid biosynthesis. A colorful model for genetics,biochemistry,cell biology,and biotechnology. Plant Physiol, 126 (2):485-493.
pmid: 11402179 |
| [30] |
Wu Z J, Tian C, Jiang Q, Li X H, Zhuang J. 2016. Selection of suitable reference genes for qRT-PCR normalization during leaf development and hormonal stimuli in tea plant(Camellia sinensis). Sci Rep, 6:19748.
doi: 10.1038/srep19748 URL |
| [31] | Zeng Defu, Zhou Jianchan, Zhong Chunmei, Xie Jun. 2018. Screening of reference genes in Dioscorea composita tubers of different development stages. Plant Physiology Journal, 54 (3):509-517. (in Chinese) |
| 曾德福, 周建婵, 钟春梅, 谢君. 2018. 菊叶薯蓣不同发育时期块茎内参基因的筛选. 植物生理学报, 54 (3):509-517. | |
| [32] |
Zhang K M, Wang J W, Guo M L, Du W L, Wu R H, Wang X. 2016. Short-day signals are crucial for the induction of anthocyanin biosynthesis in Begonia semperflorens under low temperature condition. J Plant Physiol, 204:1-7.
doi: 10.1016/j.jplph.2016.06.021 URL |
| [33] | Zhang Ning, Hu Zongli, Chen Xuqing, Hou Xiaoshu, Li Yong, Chen Guoping. 2008. Analysis of metabolic pathway and establishment of regulating model of anthocyanin synthesis. China Biotechnology, 28 (1):97-105.. (in Chinese) |
| 张宁, 胡宗利, 陈绪清, 侯晓姝, 李勇, 陈国平. 2008. 植物花青素代谢途径分析及调控模型建立. 中国生物工程杂志, 28 (1):97-105. |
| [1] | ZHAO Xueyan, WANG Qi, WANG Li, WANG Fangyuan, WANG Qing, LI Yan. Comparative Transcriptome Analysis of Differential Expression in Different Tissues of Corydalis yanhusuo [J]. Acta Horticulturae Sinica, 2023, 50(1): 177-187. |
| [2] | GAO Yanlong, WU Yuxia, ZHANG Zhongxing, WANG Shuangcheng, ZHANG Rui, ZHANG De, WANG Yanxiu. Bioinformatics Analysis of Apple ELO Gene Family and Its Expression Analysis Under Low Temperature Stress [J]. Acta Horticulturae Sinica, 2022, 49(8): 1621-1636. |
| [3] | QIU Ziwen, LIU Linmin, LIN Yongsheng, LIN Xiaojie, LI Yongyu, WU Shaohua, YANG Chao. Cloning and Functional Analysis of the MbEGS Gene from Melaleuca bracteata [J]. Acta Horticulturae Sinica, 2022, 49(8): 1747-1760. |
| [4] | ZHENG Lin, WANG Shuai, LIU Yunuo, DU Meixia, PENG Aihong, HE Yongrui, CHEN Shanchun, ZOU Xiuping. Gene Cloning and Expression Analysis of NAC Gene in Citrus in Response to Huanglongbing [J]. Acta Horticulturae Sinica, 2022, 49(7): 1441-1457. |
| [5] | ZHANG Qiuyue, LIU Changlai, YU Xiaojing, YANG Jiading, FENG Chaonian. Screening of Reference Genes for Differentially Expressed Genes in Pyrus betulaefolia Plant Under Salt Stress by qRT-PCR [J]. Acta Horticulturae Sinica, 2022, 49(7): 1557-1570. |
| [6] | MA Weifeng, LI Yanmei, MA Zonghuan, CHEN Baihong, MAO Juan. Identification of Apple POD Gene Family and Functional Analysis of MdPOD15 Gene [J]. Acta Horticulturae Sinica, 2022, 49(6): 1181-1199. |
| [7] | ZHANG Kai, MA Mingying, WANG Ping, LI Yi, JIN Yan, SHENG Ling, DENG Ziniu, MA Xianfeng. Identification of HSP20 Family Genes in Citrus and Their Expression in Pathogen Infection Responses Citrus Canker [J]. Acta Horticulturae Sinica, 2022, 49(6): 1213-1232. |
| [8] | LIANG Chen, SUN Ruyi, XIANG Rui, SUN Yimeng, SHI Xiaoxin, DU Guoqiang, WANG Li. Genome-wide Identification of Grape GRF Family and Expression Analysis [J]. Acta Horticulturae Sinica, 2022, 49(5): 995-1007. |
| [9] | XIAO Xuechen, LIU Mengyu, JIANG Mengqi, CHEN Yan, XUE Xiaodong, ZHOU Chengzhe, WU Xingjian, WU Junnan, GUO Yinsheng, YEH Kaiwen, LAI Zhongxiong, LIN Yuling. Whole-genome Identification and Expression Analysis of SNAT,ASMT and COMT Families of Melatonin Synthesis Pathway in Dimocarpus longan [J]. Acta Horticulturae Sinica, 2022, 49(5): 1031-1046. |
| [10] | GAO Weilin, ZHANG Liman, XUE Chaoling, ZHANG Yao, LIU Mengjun, ZHAO Jin. Expression of E-type MADS-box Genes in Flower and Fruits and Protein Interaction Analysis in Chinese Jujube [J]. Acta Horticulturae Sinica, 2022, 49(4): 739-748. |
| [11] | LIU Mengyu, JIANG Mengqi, CHEN Yan, ZHANG Shuting, XUE Xiaodong, XIAO Xuechen, LAI Zhongxiong, LIN Yuling. Genome-wide Identification and Expression Analysis of GDSL Esterase/Lipase Genes in Dimocarpus longan [J]. Acta Horticulturae Sinica, 2022, 49(3): 597-612. |
| [12] | JIANG Cuicui, FANG Zhizhen, ZHOU Danrong, LIN Yanjuan, YE Xinfu. Identification and Expression Analysis of Sugar Transporter Family Genes in‘Furongli’(Prunus salicina) [J]. Acta Horticulturae Sinica, 2022, 49(2): 252-264. |
| [13] | WANG Zhiyu, CHANG Beibei, LIU Qi, CHENG Xiaofan, DU Xiaoyun, YU Xiaoli, SONG Laiqing, ZHAO Lingling. Study on Expression and Anthocyanin Accumulation of Solute Carrier Gene MdSLC35F2-like in Apple [J]. Acta Horticulturae Sinica, 2022, 49(11): 2293-2303. |
| [14] | HUANG Renwei, REN Yinghong, QI Weiliang, ZENG Rui, LIU Xinyu, DENG Binyan. Cloning of Mulberry MaERF105-Like Gene and Its Expression Under Drought Stress [J]. Acta Horticulturae Sinica, 2022, 49(11): 2439-2448. |
| [15] | HOU Tianze, YI Shuangshuang, ZHANG Zhiqun, WANG Jian, LI Chonghui. Selection and Validation of Reference Genes for RT-qPCR in Phalaenopsis- type Dendrobium Hybrid [J]. Acta Horticulturae Sinica, 2022, 49(11): 2489-2501. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd