
园艺学报 ›› 2023, Vol. 50 ›› Issue (4): 815-830.doi: 10.16420/j.issn.0513-353x.2021-1238
王同欢1, 吴雨馨1, 武艺圆1, 李鑫鑫1, 刘梦阳1, 杨莲莲1,2, 李佳鹏1,2, 张忠山1,2, 曹访1, 仲雪婷1,2, 王占旗1,2,*( )
)
收稿日期:2022-08-13
修回日期:2022-12-11
出版日期:2023-04-25
发布日期:2023-04-27
通讯作者:
*(E-mail:zhqwang@zjhu.edu.cn)
基金资助:
WANG Tonghuan1, WU Yuxin1, WU Yiyuan1, LI Xinxin1, LIU Mengyang1, YANG Lianlian1,2, LI Jiapeng1,2, ZHANG Zhongshan1,2, CAO Fang1, ZHONG Xueting1,2, WANG Zhanqi1,2,*( )
)
Received:2022-08-13
Revised:2022-12-11
Online:2023-04-25
Published:2023-04-27
Contact:
*(E-mail:zhqwang@zjhu.edu.cn)
摘要:
利用生物信息的方法鉴定了菊花脑(Chrysanthemum nankingense)GRAS基因家族成员,并利用已发表的RNA-Seq数据分析了CnGRAS在低温胁迫应答中的表达情况。结果表明:CnGRAS家族包含80个成员,可分为10个亚家族,同一个亚家族中各成员的基因结构和蛋白功能域高度保守。复制进化分析发现,CnGRAS家族的复制模式仅存在串联重复。GRAS在菊花脑中的表达具有组织特异性,多数在根和叶中表达水平较高。GO富集分析表明CnGRAS主要定位于细胞核,具有多种分子功能,参与多个生物学过程。RNA-Seq数据分析表明,14个CnGRAS至少在1种低温胁迫(短期低温或长期低温)条件下呈现差异表达,其中CnGRAS32、CnGRAS33、CnGRAS40、CnGRAS44、CnGRAS56、CnGRAS74和CnGRAS79呈现出显著上调表达,说明这些基因可能在菊花脑响应低温胁迫中起着重要作用。
中图分类号:
王同欢, 吴雨馨, 武艺圆, 李鑫鑫, 刘梦阳, 杨莲莲, 李佳鹏, 张忠山, 曹访, 仲雪婷, 王占旗. 菊花脑GRAS家族鉴定及其低温胁迫响应表达分析[J]. 园艺学报, 2023, 50(4): 815-830.
WANG Tonghuan, WU Yuxin, WU Yiyuan, LI Xinxin, LIU Mengyang, YANG Lianlian, LI Jiapeng, ZHANG Zhongshan, CAO Fang, ZHONG Xueting, WANG Zhanqi. Genome-wide Identification and Expression Analysis of the GRAS Gene Family in Response to Cold Stress in Chrysanthemum nankingense[J]. Acta Horticulturae Sinica, 2023, 50(4): 815-830.
| 基因名 Gene name | 基因ID Gene ID | 方向 Strand | 开放阅读框/bp ORF | 外显子数 Number of exons | 蛋白质/aa Protein | 等电点 pI | 分子量/kD MW | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CnGRAS1 | CHR00004937-RA | 反向 Reverse | 1 401 | 6 | 466 | 4.6 | 52.8 | ||||||
| CnGRAS2 | CHR00004939-RA | 反向 Reverse | 948 | 6 | 315 | 4.5 | 35.8 | ||||||
| CnGRAS3 | CHR00004941-RA | 反向 Reverse | 1 305 | 4 | 434 | 4.6 | 48.9 | ||||||
| CnGRAS4 | CHR00004943-RA | 反向 Reverse | 1 005 | 5 | 334 | 5.0 | 37.9 | ||||||
| CnGRAS5 | CHR00010605-RA | 反向 Reverse | 1 464 | 3 | 487 | 5.7 | 54.8 | ||||||
| CnGRAS6 | CHR00010606-RA | 反向 Reverse | 1 551 | 6 | 516 | 5.5 | 57.9 | ||||||
| CnGRAS7 | CHR00030831-RA | 反向 Reverse | 1 317 | 6 | 438 | 5.3 | 49.3 | ||||||
| CnGRAS8 | CHR00009270-RA | 反向 Reverse | 882 | 3 | 293 | 5.2 | 33.7 | ||||||
| CnGRAS9 | CHR00003961-RA | 反向 Reverse | 3 216 | 3 | 1 071 | 5.5 | 121.1 | ||||||
| CnGRAS10 | CHR00055273-RA | 反向 Reverse | 762 | 4 | 253 | 5.7 | 28.5 | ||||||
| CnGRAS11 | CHR00032218-RA | 反向 Reverse | 1 407 | 2 | 468 | 6.1 | 52.6 | ||||||
| CnGRAS12 | CHR00031862-RA | 正向 Forward | 624 | 3 | 207 | 8.9 | 22.7 | ||||||
| CnGRAS13 | CHR00060013-RA | 正向 Forward | 2 187 | 7 | 728 | 5.1 | 81.5 | ||||||
| CnGRAS14 | CHR00049603-RA | 反向 Reverse | 1 965 | 3 | 654 | 5.5 | 71.8 | ||||||
| CnGRAS15 | CHR00044507-RA | 正向 Forward | 1 587 | 2 | 528 | 5.9 | 59.0 | ||||||
| CnGRAS16 | CHR00088841-RA | 正向 Forward | 684 | 2 | 227 | 6.4 | 25.3 | ||||||
| CnGRAS17 | CHR00057450-RA | 反向 Reverse | 1 809 | 4 | 602 | 6.4 | 66.5 | ||||||
| CnGRAS18 | CHR00032070-RA | 正向 Forward | 1 605 | 4 | 534 | 5.8 | 61.0 | ||||||
| CnGRAS19 | CHR00001839-RA | 正向 Forward | 1 656 | 1 | 551 | 6.2 | 62.0 | ||||||
| CnGRAS20 | CHR00003428-RA | 正向 Forward | 1 560 | 3 | 519 | 4.9 | 58.8 | ||||||
| CnGRAS21 | CHR00080196-RA | 反向 Reverse | 1 863 | 4 | 620 | 5.8 | 69.5 | ||||||
| CnGRAS22 | CHR00056344-RA | 反向 Reverse | 1 509 | 5 | 502 | 5.2 | 56.5 | ||||||
| CnGRAS23 | CHR00043261-RA | 反向 Reverse | 1 617 | 6 | 538 | 5.4 | 61.0 | ||||||
| CnGRAS24 | CHR00007360-RA | 正向 Forward | 1 647 | 6 | 548 | 5.7 | 61.3 | ||||||
| CnGRAS25 | CHR00084234-RA | 反向 Reverse | 1 971 | 5 | 656 | 6.3 | 75.5 | ||||||
| CnGRAS26 | CHR00026458-RA | 正向 Forward | 1 587 | 7 | 528 | 5.1 | 59.7 | ||||||
| CnGRAS27 | CHR00012285-RA | 正向 Forward | 1 392 | 5 | 463 | 5.8 | 53.1 | ||||||
| CnGRAS28 | CHR00012287-RA | 正向 Forward | 1 389 | 2 | 462 | 6.2 | 53.3 | ||||||
| CnGRAS29 | CHR00001229-RA | 反向 Reverse | 1 659 | 5 | 552 | 6.4 | 62.5 | ||||||
| CnGRAS30 | CHR00090421-RA | 正向 Forward | 1 326 | 5 | 441 | 5.8 | 50.0 | ||||||
| CnGRAS31 | CHR00090422-RA | 正向 Forward | 1 770 | 12 | 589 | 8.2 | 67.0 | ||||||
| CnGRAS32 | CHR00006827-RA | 反向 Reverse | 1 989 | 4 | 662 | 6.5 | 73.5 | ||||||
| CnGRAS33 | CHR00080708-RA | 反向 Reverse | 1 845 | 7 | 614 | 5.7 | 68.2 | ||||||
| CnGRAS34 | CHR00009185-RA | 正向 Forward | 1 761 | 2 | 586 | 7.2 | 66.5 | ||||||
| CnGRAS35 | CHR00014432-RA | 反向 Reverse | 1 284 | 4 | 427 | 7.3 | 48.9 | ||||||
| CnGRAS36 | CHR00027166-RA | 正向 Forward | 1 995 | 5 | 664 | 5.8 | 73.2 | ||||||
| CnGRAS37 | CHR00030490-RA | 反向 Reverse | 1 221 | 3 | 406 | 6.1 | 46.3 | ||||||
| CnGRAS38 | CHR00089224-RA | 反向 Reverse | 1 389 | 8 | 462 | 5.8 | 51.9 | ||||||
| CnGRAS39 | CHR00032024-RA | 反向 Reverse | 840 | 2 | 279 | 7.2 | 31.8 | ||||||
| CnGRAS40 | CHR00059102-RA | 反向 Reverse | 1 176 | 5 | 391 | 5.1 | 43.2 | ||||||
| CnGRAS41 | CHR00020653-RA | 反向 Reverse | 660 | 7 | 219 | 9.1 | 25.6 | ||||||
| CnGRAS42 | CHR00023444-RA | 正向 Forward | 1 461 | 4 | 486 | 5.6 | 54.6 | ||||||
| CnGRAS43 | CHR00047978-RA | 正向 Forward | 2 112 | 5 | 703 | 7.6 | 79.9 | ||||||
| CnGRAS44 | CHR00037746-RA | 反向(reverse) | 1 350 | 8 | 449 | 5.7 | 50.2 | ||||||
| CnGRAS45 | CHR00050945-RA | 正向 Forward | 1 464 | 3 | 487 | 5.3 | 54.8 | ||||||
| CnGRAS46 | CHR00043205-RA | 反向(reverse) | 1 656 | 6 | 551 | 5.3 | 62.2 | ||||||
| CnGRAS47 | CHR00020406-RA | 正向 Forward | 1 047 | 3 | 348 | 7.7 | 38.9 | ||||||
| CnGRAS48 | CHR00034025-RA | 反向 Reverse | 1 608 | 3 | 535 | 5.2 | 59.6 | ||||||
| CnGRAS49 | CHR00074287-RA | 正向 Forward | 1 500 | 2 | 499 | 5.4 | 55.5 | ||||||
| CnGRAS50 | CHR00022194-RA | 正向 Forward | 1 269 | 3 | 422 | 5.7 | 48.2 | ||||||
| CnGRAS51 | CHR00013495-RA | 正向 Forward | 771 | 4 | 256 | 5.9 | 28.4 | ||||||
| CnGRAS52 | CHR00043827-RA | 反向 Reverse | 1 221 | 4 | 406 | 5.9 | 46.8 | ||||||
| CnGRAS53 | CHR00043828-RA | 反向 Reverse | 2 172 | 5 | 723 | 5.4 | 82.3 | ||||||
| CnGRAS54 | CHR00082830-RA | 反向 Reverse | 1 779 | 4 | 592 | 5.7 | 101.7 | ||||||
| CnGRAS55 | CHR00031262-RA | 正向 Forward | 1 413 | 3 | 470 | 8.0 | 53.8 | ||||||
| CnGRAS56 | CHR00066643-RA | 正向 Forward | 2 175 | 7 | 724 | 5.3 | 82.6 | ||||||
| CnGRAS57 | CHR00036461-RA | 正向 Forward | 1 665 | 5 | 554 | 6.3 | 61.9 | ||||||
| CnGRAS58 | CHR00037682-RA | 反向 Reverse | 1 548 | 3 | 515 | 6.0 | 56.4 | ||||||
| CnGRAS59 | CHR00081984-RA | 正向 Forward | 963 | 2 | 320 | 8.6 | 37.1 | ||||||
| CnGRAS60 | CHR00076473-RA | 反向 Reverse | 1 209 | 2 | 402 | 5.4 | 44.2 | ||||||
| CnGRAS61 | CHR00077126-RA | 正向 Forward | 1 326 | 6 | 441 | 5.2 | 49.7 | ||||||
| CnGRAS62 | CHR00021855-RA | 正向 Forward | 1 356 | 4 | 451 | 5.9 | 50.9 | ||||||
| CnGRAS63 | CHR00039693-RA | 反向 Reverse | 1 584 | 4 | 527 | 5.6 | 58.6 | ||||||
| CnGRAS64 | CHR00022872-RA | 正向 Forward | 1 023 | 4 | 340 | 5.0 | 37.8 | ||||||
| CnGRAS65 | CHR00086408-RA | 反向 Reverse | 1 323 | 3 | 440 | 5.1 | 49.7 | ||||||
| CnGRAS66 | CHR00081508-RA | 正向 Forward | 1 632 | 4 | 543 | 6.3 | 61.2 | ||||||
| CnGRAS67 | CHR00087940-RA | 反向 Reverse | 1 878 | 5 | 625 | 5.4 | 69.6 | ||||||
| CnGRAS68 | CHR00058033-RA | 反向 Reverse | 1 488 | 3 | 495 | 7.5 | 54.8 | ||||||
| CnGRAS69 | CHR00087040-RA | 反向 Reverse | 1 032 | 4 | 343 | 6.0 | 39.3 | ||||||
| CnGRAS70 | CHR00087041-RA | 反向 Reverse | 1 914 | 10 | 637 | 5.4 | 71.9 | ||||||
| CnGRAS71 | CHR00087042-RA | 反向 Reverse | 1 446 | 3 | 481 | 5.9 | 54.8 | ||||||
| CnGRAS72 | CHR00087043-RA | 反向 Reverse | 2 208 | 4 | 735 | 4.9 | 83.8 | ||||||
| CnGRAS73 | CHR00087044-RA | 反向 Reverse | 969 | 8 | 322 | 8.2 | 36.3 | ||||||
| CnGRAS74 | CHR00087600-RA | 反向 Reverse | 1 371 | 4 | 456 | 6.2 | 51.3 | ||||||
| CnGRAS75 | CHR00091555-RA | 反向 Reverse | 1 257 | 4 | 418 | 5.3 | 46.5 | ||||||
| CnGRAS76 | CHR00065020-RA | 反向 Reverse | 1 581 | 5 | 526 | 4.8 | 57.4 | ||||||
| CnGRAS77 | CHR00065743-RA | 正向 Forward | 1 515 | 6 | 504 | 6.1 | 57.8 | ||||||
| CnGRAS78 | CHR00069367-RA | 正向 Forward | 1 488 | 6 | 495 | 5.3 | 54.2 | ||||||
| CnGRAS79 | CHR00093834-RA | 反向 Reverse | 705 | 3 | 234 | 4.6 | 26.1 | ||||||
| CnGRAS80 | CHR00074223-RA | 正向 Forward | 735 | 5 | 244 | 9.1 | 28.6 | ||||||
表1 菊花脑GRAS家族的结构和理化特征
Table 1 Structural,physical and chemical features of GRAS gene family in Chrysanthemum nankingense
| 基因名 Gene name | 基因ID Gene ID | 方向 Strand | 开放阅读框/bp ORF | 外显子数 Number of exons | 蛋白质/aa Protein | 等电点 pI | 分子量/kD MW | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CnGRAS1 | CHR00004937-RA | 反向 Reverse | 1 401 | 6 | 466 | 4.6 | 52.8 | ||||||
| CnGRAS2 | CHR00004939-RA | 反向 Reverse | 948 | 6 | 315 | 4.5 | 35.8 | ||||||
| CnGRAS3 | CHR00004941-RA | 反向 Reverse | 1 305 | 4 | 434 | 4.6 | 48.9 | ||||||
| CnGRAS4 | CHR00004943-RA | 反向 Reverse | 1 005 | 5 | 334 | 5.0 | 37.9 | ||||||
| CnGRAS5 | CHR00010605-RA | 反向 Reverse | 1 464 | 3 | 487 | 5.7 | 54.8 | ||||||
| CnGRAS6 | CHR00010606-RA | 反向 Reverse | 1 551 | 6 | 516 | 5.5 | 57.9 | ||||||
| CnGRAS7 | CHR00030831-RA | 反向 Reverse | 1 317 | 6 | 438 | 5.3 | 49.3 | ||||||
| CnGRAS8 | CHR00009270-RA | 反向 Reverse | 882 | 3 | 293 | 5.2 | 33.7 | ||||||
| CnGRAS9 | CHR00003961-RA | 反向 Reverse | 3 216 | 3 | 1 071 | 5.5 | 121.1 | ||||||
| CnGRAS10 | CHR00055273-RA | 反向 Reverse | 762 | 4 | 253 | 5.7 | 28.5 | ||||||
| CnGRAS11 | CHR00032218-RA | 反向 Reverse | 1 407 | 2 | 468 | 6.1 | 52.6 | ||||||
| CnGRAS12 | CHR00031862-RA | 正向 Forward | 624 | 3 | 207 | 8.9 | 22.7 | ||||||
| CnGRAS13 | CHR00060013-RA | 正向 Forward | 2 187 | 7 | 728 | 5.1 | 81.5 | ||||||
| CnGRAS14 | CHR00049603-RA | 反向 Reverse | 1 965 | 3 | 654 | 5.5 | 71.8 | ||||||
| CnGRAS15 | CHR00044507-RA | 正向 Forward | 1 587 | 2 | 528 | 5.9 | 59.0 | ||||||
| CnGRAS16 | CHR00088841-RA | 正向 Forward | 684 | 2 | 227 | 6.4 | 25.3 | ||||||
| CnGRAS17 | CHR00057450-RA | 反向 Reverse | 1 809 | 4 | 602 | 6.4 | 66.5 | ||||||
| CnGRAS18 | CHR00032070-RA | 正向 Forward | 1 605 | 4 | 534 | 5.8 | 61.0 | ||||||
| CnGRAS19 | CHR00001839-RA | 正向 Forward | 1 656 | 1 | 551 | 6.2 | 62.0 | ||||||
| CnGRAS20 | CHR00003428-RA | 正向 Forward | 1 560 | 3 | 519 | 4.9 | 58.8 | ||||||
| CnGRAS21 | CHR00080196-RA | 反向 Reverse | 1 863 | 4 | 620 | 5.8 | 69.5 | ||||||
| CnGRAS22 | CHR00056344-RA | 反向 Reverse | 1 509 | 5 | 502 | 5.2 | 56.5 | ||||||
| CnGRAS23 | CHR00043261-RA | 反向 Reverse | 1 617 | 6 | 538 | 5.4 | 61.0 | ||||||
| CnGRAS24 | CHR00007360-RA | 正向 Forward | 1 647 | 6 | 548 | 5.7 | 61.3 | ||||||
| CnGRAS25 | CHR00084234-RA | 反向 Reverse | 1 971 | 5 | 656 | 6.3 | 75.5 | ||||||
| CnGRAS26 | CHR00026458-RA | 正向 Forward | 1 587 | 7 | 528 | 5.1 | 59.7 | ||||||
| CnGRAS27 | CHR00012285-RA | 正向 Forward | 1 392 | 5 | 463 | 5.8 | 53.1 | ||||||
| CnGRAS28 | CHR00012287-RA | 正向 Forward | 1 389 | 2 | 462 | 6.2 | 53.3 | ||||||
| CnGRAS29 | CHR00001229-RA | 反向 Reverse | 1 659 | 5 | 552 | 6.4 | 62.5 | ||||||
| CnGRAS30 | CHR00090421-RA | 正向 Forward | 1 326 | 5 | 441 | 5.8 | 50.0 | ||||||
| CnGRAS31 | CHR00090422-RA | 正向 Forward | 1 770 | 12 | 589 | 8.2 | 67.0 | ||||||
| CnGRAS32 | CHR00006827-RA | 反向 Reverse | 1 989 | 4 | 662 | 6.5 | 73.5 | ||||||
| CnGRAS33 | CHR00080708-RA | 反向 Reverse | 1 845 | 7 | 614 | 5.7 | 68.2 | ||||||
| CnGRAS34 | CHR00009185-RA | 正向 Forward | 1 761 | 2 | 586 | 7.2 | 66.5 | ||||||
| CnGRAS35 | CHR00014432-RA | 反向 Reverse | 1 284 | 4 | 427 | 7.3 | 48.9 | ||||||
| CnGRAS36 | CHR00027166-RA | 正向 Forward | 1 995 | 5 | 664 | 5.8 | 73.2 | ||||||
| CnGRAS37 | CHR00030490-RA | 反向 Reverse | 1 221 | 3 | 406 | 6.1 | 46.3 | ||||||
| CnGRAS38 | CHR00089224-RA | 反向 Reverse | 1 389 | 8 | 462 | 5.8 | 51.9 | ||||||
| CnGRAS39 | CHR00032024-RA | 反向 Reverse | 840 | 2 | 279 | 7.2 | 31.8 | ||||||
| CnGRAS40 | CHR00059102-RA | 反向 Reverse | 1 176 | 5 | 391 | 5.1 | 43.2 | ||||||
| CnGRAS41 | CHR00020653-RA | 反向 Reverse | 660 | 7 | 219 | 9.1 | 25.6 | ||||||
| CnGRAS42 | CHR00023444-RA | 正向 Forward | 1 461 | 4 | 486 | 5.6 | 54.6 | ||||||
| CnGRAS43 | CHR00047978-RA | 正向 Forward | 2 112 | 5 | 703 | 7.6 | 79.9 | ||||||
| CnGRAS44 | CHR00037746-RA | 反向(reverse) | 1 350 | 8 | 449 | 5.7 | 50.2 | ||||||
| CnGRAS45 | CHR00050945-RA | 正向 Forward | 1 464 | 3 | 487 | 5.3 | 54.8 | ||||||
| CnGRAS46 | CHR00043205-RA | 反向(reverse) | 1 656 | 6 | 551 | 5.3 | 62.2 | ||||||
| CnGRAS47 | CHR00020406-RA | 正向 Forward | 1 047 | 3 | 348 | 7.7 | 38.9 | ||||||
| CnGRAS48 | CHR00034025-RA | 反向 Reverse | 1 608 | 3 | 535 | 5.2 | 59.6 | ||||||
| CnGRAS49 | CHR00074287-RA | 正向 Forward | 1 500 | 2 | 499 | 5.4 | 55.5 | ||||||
| CnGRAS50 | CHR00022194-RA | 正向 Forward | 1 269 | 3 | 422 | 5.7 | 48.2 | ||||||
| CnGRAS51 | CHR00013495-RA | 正向 Forward | 771 | 4 | 256 | 5.9 | 28.4 | ||||||
| CnGRAS52 | CHR00043827-RA | 反向 Reverse | 1 221 | 4 | 406 | 5.9 | 46.8 | ||||||
| CnGRAS53 | CHR00043828-RA | 反向 Reverse | 2 172 | 5 | 723 | 5.4 | 82.3 | ||||||
| CnGRAS54 | CHR00082830-RA | 反向 Reverse | 1 779 | 4 | 592 | 5.7 | 101.7 | ||||||
| CnGRAS55 | CHR00031262-RA | 正向 Forward | 1 413 | 3 | 470 | 8.0 | 53.8 | ||||||
| CnGRAS56 | CHR00066643-RA | 正向 Forward | 2 175 | 7 | 724 | 5.3 | 82.6 | ||||||
| CnGRAS57 | CHR00036461-RA | 正向 Forward | 1 665 | 5 | 554 | 6.3 | 61.9 | ||||||
| CnGRAS58 | CHR00037682-RA | 反向 Reverse | 1 548 | 3 | 515 | 6.0 | 56.4 | ||||||
| CnGRAS59 | CHR00081984-RA | 正向 Forward | 963 | 2 | 320 | 8.6 | 37.1 | ||||||
| CnGRAS60 | CHR00076473-RA | 反向 Reverse | 1 209 | 2 | 402 | 5.4 | 44.2 | ||||||
| CnGRAS61 | CHR00077126-RA | 正向 Forward | 1 326 | 6 | 441 | 5.2 | 49.7 | ||||||
| CnGRAS62 | CHR00021855-RA | 正向 Forward | 1 356 | 4 | 451 | 5.9 | 50.9 | ||||||
| CnGRAS63 | CHR00039693-RA | 反向 Reverse | 1 584 | 4 | 527 | 5.6 | 58.6 | ||||||
| CnGRAS64 | CHR00022872-RA | 正向 Forward | 1 023 | 4 | 340 | 5.0 | 37.8 | ||||||
| CnGRAS65 | CHR00086408-RA | 反向 Reverse | 1 323 | 3 | 440 | 5.1 | 49.7 | ||||||
| CnGRAS66 | CHR00081508-RA | 正向 Forward | 1 632 | 4 | 543 | 6.3 | 61.2 | ||||||
| CnGRAS67 | CHR00087940-RA | 反向 Reverse | 1 878 | 5 | 625 | 5.4 | 69.6 | ||||||
| CnGRAS68 | CHR00058033-RA | 反向 Reverse | 1 488 | 3 | 495 | 7.5 | 54.8 | ||||||
| CnGRAS69 | CHR00087040-RA | 反向 Reverse | 1 032 | 4 | 343 | 6.0 | 39.3 | ||||||
| CnGRAS70 | CHR00087041-RA | 反向 Reverse | 1 914 | 10 | 637 | 5.4 | 71.9 | ||||||
| CnGRAS71 | CHR00087042-RA | 反向 Reverse | 1 446 | 3 | 481 | 5.9 | 54.8 | ||||||
| CnGRAS72 | CHR00087043-RA | 反向 Reverse | 2 208 | 4 | 735 | 4.9 | 83.8 | ||||||
| CnGRAS73 | CHR00087044-RA | 反向 Reverse | 969 | 8 | 322 | 8.2 | 36.3 | ||||||
| CnGRAS74 | CHR00087600-RA | 反向 Reverse | 1 371 | 4 | 456 | 6.2 | 51.3 | ||||||
| CnGRAS75 | CHR00091555-RA | 反向 Reverse | 1 257 | 4 | 418 | 5.3 | 46.5 | ||||||
| CnGRAS76 | CHR00065020-RA | 反向 Reverse | 1 581 | 5 | 526 | 4.8 | 57.4 | ||||||
| CnGRAS77 | CHR00065743-RA | 正向 Forward | 1 515 | 6 | 504 | 6.1 | 57.8 | ||||||
| CnGRAS78 | CHR00069367-RA | 正向 Forward | 1 488 | 6 | 495 | 5.3 | 54.2 | ||||||
| CnGRAS79 | CHR00093834-RA | 反向 Reverse | 705 | 3 | 234 | 4.6 | 26.1 | ||||||
| CnGRAS80 | CHR00074223-RA | 正向 Forward | 735 | 5 | 244 | 9.1 | 28.6 | ||||||
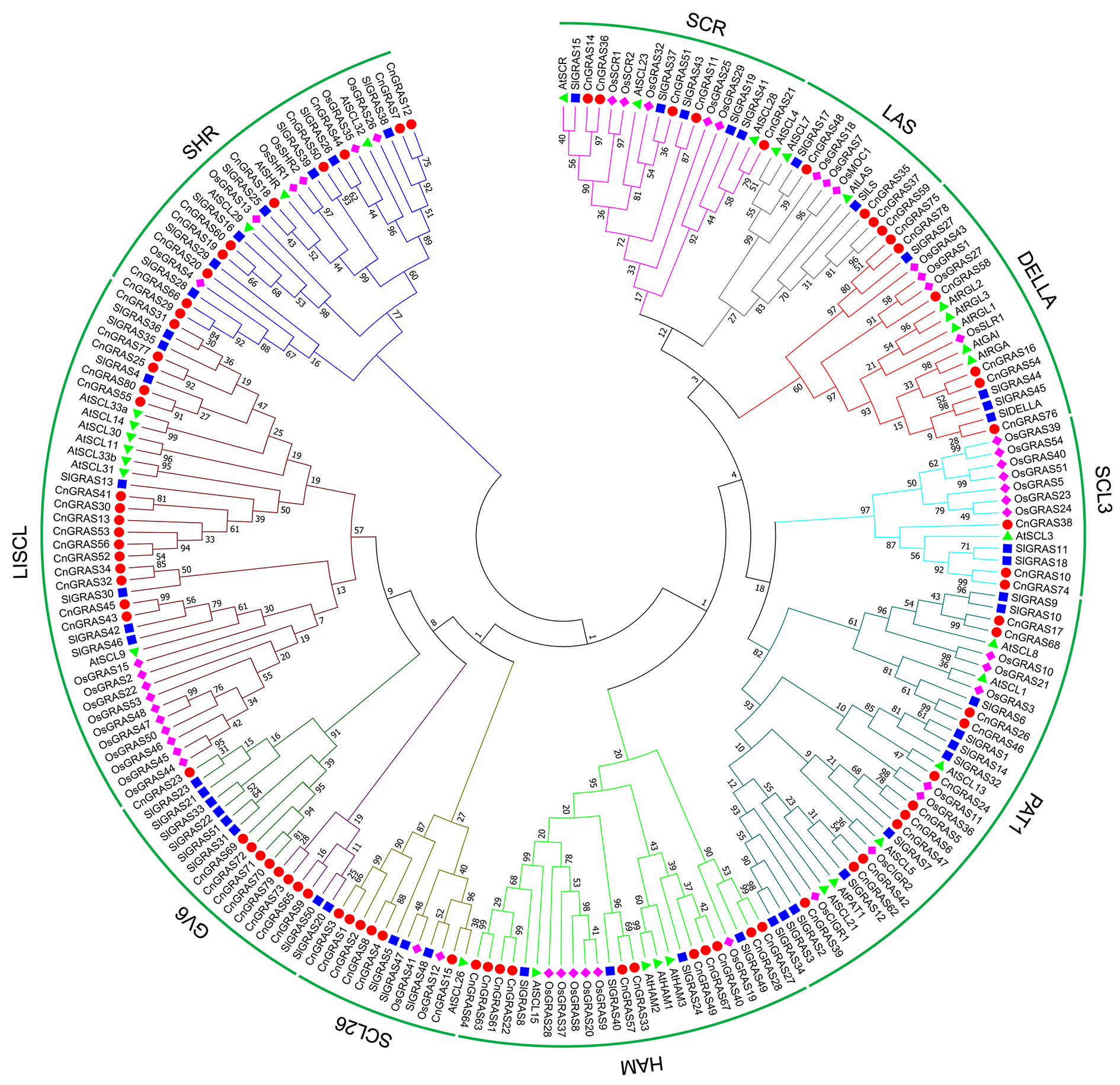
图1 菊花脑(Cn)、拟南芥(At)、番茄(Sl)和水稻(Os)GRAS基因家族系统进化分析
Fig. 1 Phylogenetic analysis of GRAS gene family in Chrysanthemum nankingense(Cn),Arabidopsis thaliana(At),Solanum lycopersicum(Sl)and Oryza sativa(Os)
| 基因对 Gene pair | 非同义替换率(Ka) Non-synonymous substitution rate | 同义替换率(Ks) Synonymous substitution rate | Ka/Ks | 复制时间/Myr Duplication date | 纯化选择 Purifying selection | ||
|---|---|---|---|---|---|---|---|
| CnGRAS2 | CnGRAS3 | 0.038 | 0.087 | 0.438 | 2.9 | Yes | |
| CnGRAS5 | CnGRAS6 | 0.151 | 0.555 | 0.272 | 18.5 | Yes | |
| CnGRAS27 | CnGRAS28 | 0.024 | 0.080 | 0.305 | 2.7 | Yes | |
| CnGRAS30 | CnGRAS31 | 0.470 | 2.249 | 0.209 | 75.0 | Yes | |
| CnGRAS52 | CnGRAS53 | 0.142 | 0.415 | 0.344 | 13.8 | Yes | |
| CnGRAS69 | CnGRAS70 | 0.275 | 0.821 | 0.335 | 27.4 | Yes | |
| CnGRAS70 | CnGRAS71 | 0.116 | 0.150 | 0.769 | 5.0 | Yes | |
| CnGRAS71 | CnGRAS72 | 0.145 | 0.271 | 0.534 | 9.0 | Yes | |
| CnGRAS72 | CnGRAS73 | 0.630 | 2.365 | 0.266 | 78.8 | Yes | |
表2 菊花脑GRAS重复基因对的进化分析
Table 2 Divergence between duplicated GRAS gene pairs in Chrysanthemum nankingense
| 基因对 Gene pair | 非同义替换率(Ka) Non-synonymous substitution rate | 同义替换率(Ks) Synonymous substitution rate | Ka/Ks | 复制时间/Myr Duplication date | 纯化选择 Purifying selection | ||
|---|---|---|---|---|---|---|---|
| CnGRAS2 | CnGRAS3 | 0.038 | 0.087 | 0.438 | 2.9 | Yes | |
| CnGRAS5 | CnGRAS6 | 0.151 | 0.555 | 0.272 | 18.5 | Yes | |
| CnGRAS27 | CnGRAS28 | 0.024 | 0.080 | 0.305 | 2.7 | Yes | |
| CnGRAS30 | CnGRAS31 | 0.470 | 2.249 | 0.209 | 75.0 | Yes | |
| CnGRAS52 | CnGRAS53 | 0.142 | 0.415 | 0.344 | 13.8 | Yes | |
| CnGRAS69 | CnGRAS70 | 0.275 | 0.821 | 0.335 | 27.4 | Yes | |
| CnGRAS70 | CnGRAS71 | 0.116 | 0.150 | 0.769 | 5.0 | Yes | |
| CnGRAS71 | CnGRAS72 | 0.145 | 0.271 | 0.534 | 9.0 | Yes | |
| CnGRAS72 | CnGRAS73 | 0.630 | 2.365 | 0.266 | 78.8 | Yes | |
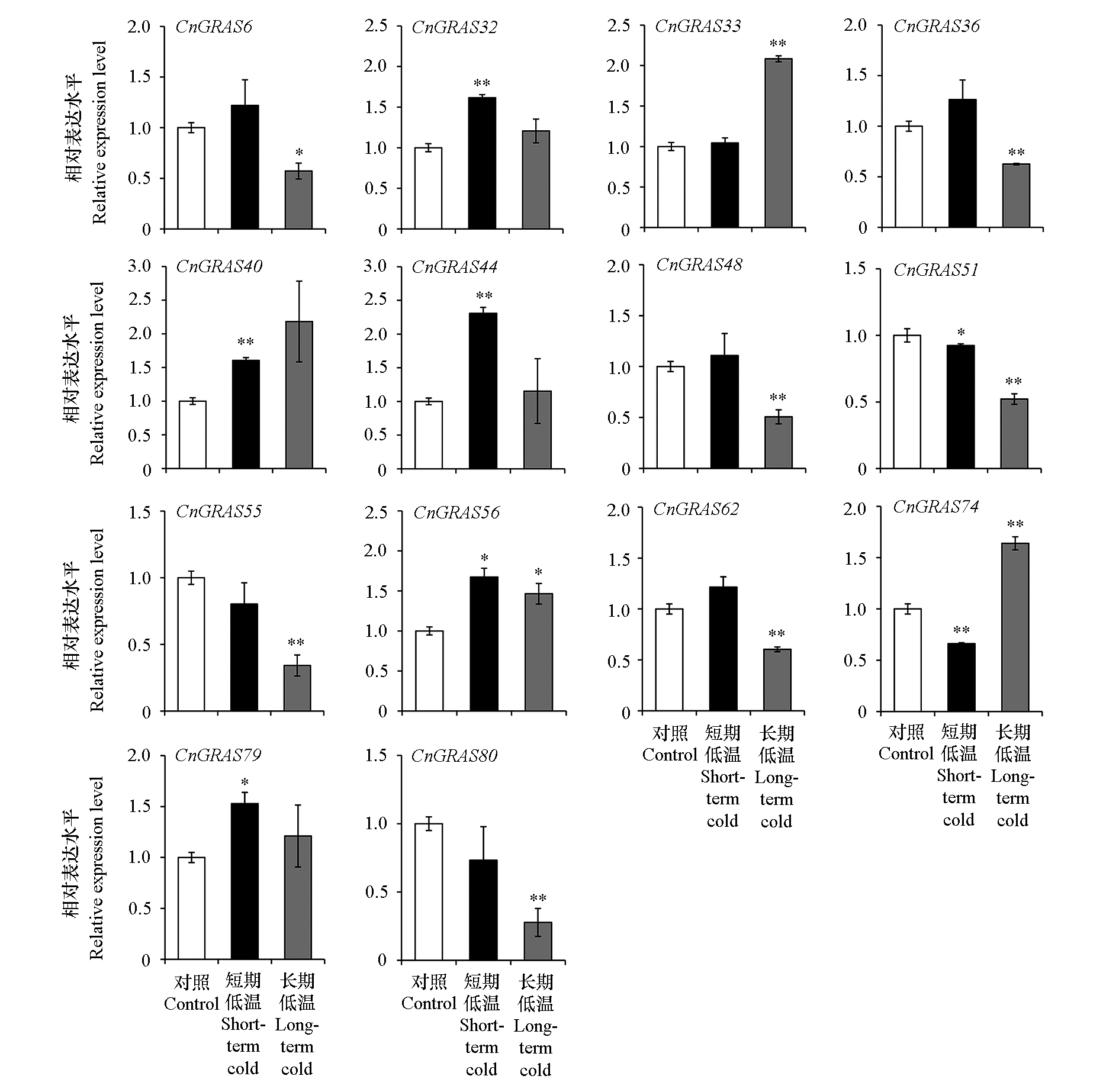
图7 菊花脑在短期和长期低温(STC和LTC)胁迫下GRAS的表达 Student’s t-test,*P < 0.05,**P < 0.01,ns无显著性差异。
Fig. 7 Expression of GRAS gene family in Chrysanthemum nankingense under short-term cold or long-term cold(STC or LTC)stress Student’s t-test,*P < 0.05,**P < 0.01,ns,not significant.
| [1] |
Bailey T L, Johnson J, Grant C E, Noble W S. 2015. The MEME suite. Nucleic Acids Research, 43 (W1):W39-W49.
doi: 10.1093/nar/gkv416 URL |
| [2] | Blum M, Chang H Y, Chuguransky S, Grego T, Kandasaamy S, Mitchell A, Nuka G, Paysan-Lafosse T, Qureshi M, Raj S, Richardson L, Salazar G A, Williams L, Bork P, Bridge A, Gough J, Haft D H, Letunic I, Marchler-Bauer A, Mi H, Natale D A, Necci M, Orengo C A, Pandurangan A P, Rivoire C, Sigrist C J A, Sillitoe I, Thanki N, Thomas P D, Tosatto S C E, Wu C H, Bateman A, Finn R D. 2021. The InterPro protein families and domains database:20 years on. Nucleic Acids Research, 49 (D1):D344-D354. |
| [3] |
Chen C, Chen H, Zhang Y, Thomas H R, Frank M H, He Y, Xia R. 2020. TBtools:an integrative toolkit developed for interactive analyses of big biological data. Molecular Plant, 13 (8):1194-1202.
doi: 10.1016/j.molp.2020.06.009 URL |
| [4] |
Chen H, Li H, Lu X, Chen L, Liu J, Wu H. 2019. Identification and expression analysis of GRAS transcription factors to elucidate candidate genes related to stolons,fruit ripening and abiotic stresses in woodland strawberry(Fragaria vesca). International Journal of Molecular Sciences, 20 (18):4593.
doi: 10.3390/ijms20184593 URL |
| [5] |
Grimplet J, Agudelo-Romero P, Teixeira R T, Martinez-Zapater J M, Fortes A M. 2016. Structural and functional analysis of the GRAS gene family in grapevine indicates a role of GRAS proteins in the control of development and stress responses. Frontiers in Plant Science, 7:353.
doi: 10.3389/fpls.2016.00353 pmid: 27065316 |
| [6] |
Hu B, Jin J, Guo A Y, Zhang H, Luo J, Gao G. 2015. GSDS 2.0:an upgraded gene feature visualization server. Bioinformatics, 31 (8):1296-1297.
doi: 10.1093/bioinformatics/btu817 URL |
| [7] |
Huang W, Hu B, Liu J, Zhou Y, Liu S. 2020. Identification and characterization of tonoplast sugar transporter(TST)gene family in cucumber. Horticultural Plant Journal, 6 (3),145-157.
doi: 10.1016/j.hpj.2020.03.005 URL |
| [8] |
Huerta-Cepas J, Forslund K, Coelho L P, Szklarczyk D, Jensen L J, von Mering C, Bork P. 2017. Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Molecular Biology and Evolution, 34 (8):2115-2122.
doi: 10.1093/molbev/msx148 pmid: 28460117 |
| [9] |
Jin J F, Wang Z Q, He Q Y, Wang J Y, Li P F, Xu J M, Zheng S J, Fan W, Yang J L. 2020. Genome-wide identification and expression analysis of the NAC transcription factor family in tomato(Solanum lycopersicum)during aluminum stress. BMC Genomics, 21 (1):288.
doi: 10.1186/s12864-020-6689-7 |
| [10] |
Koch M A, Haubold B, Mitchell-Olds T. 2000. Comparative evolutionary analysis of chalcone synthase and alcohol dehydrogenase loci in Arabidopsis,Arabis,and related genera(Brassicaceae). Molecular Biology and Evolution, 17 (10):1483-1498.
doi: 10.1093/oxfordjournals.molbev.a026248 pmid: 11018155 |
| [11] |
Kovaka S, Zimin A V, Pertea G M, Razaghi R, Salzberg S L, Pertea M. 2019. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biology, 20 (1):278.
doi: 10.1186/s13059-019-1910-1 pmid: 31842956 |
| [12] |
Kumar S, Stecher G, Tamura K. 2016. MEGA7:Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution, 33 (7):1870-1874.
doi: 10.1093/molbev/msw054 URL |
| [13] |
Landi S, Capasso G, Esposito S. 2021. Different G6PDH isoforms show specific roles in acclimation to cold stress at various growth stages of barley(Hordeum vulgare)and Arabidopsis thaliana. Plant Physiology and Biochemistry, 169:190-202.
doi: 10.1016/j.plaphy.2021.11.017 URL |
| [14] |
Lee M H, Kim B, Song S K, Heo J O, Yu N I, Lee S A, Kim M, Kim D G, Sohn S O, Lim C E, Chang K S, Lee M M, Lim J. 2008. Large-scale analysis of the GRAS gene family in Arabidopsis thaliana. Plant Molecular Biology, 67 (6):659-670.
doi: 10.1007/s11103-008-9345-1 URL |
| [15] | Lescot M. 2002. PlantCARE,a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Research, 30 (1):325-327. |
| [16] |
Li M, Sun B, Xie F, Gong R, Luo Y, Zhang F, Yan Z, Tang H. 2019. Identification of the GRAS gene family in the Brassica juncea genome provides insight into its role in stem swelling in stem mustard. PeerJ, 7:e6682.
doi: 10.7717/peerj.6682 URL |
| [17] |
Li Y, Liang J, Zeng X, Guo H, Luo Y, Kear P, Zhang S, Zhu G. 2021. Genome-wide analysis of MYB gene family in potato provides insights into tissue-specific regulation of anthocyanin biosynthesis. Horticultural Plant Journal, 7 (2):129-141.
doi: 10.1016/j.hpj.2020.12.001 URL |
| [18] |
Liu B, Sun Y, Xue J, Jia X, Li R. 2018. Genome-wide characterization and expression analysis of GRAS gene family in pepper(Capsicum annuum L.). PeerJ, 6:e4796.
doi: 10.7717/peerj.4796 URL |
| [19] |
Liu M, Huang L, Ma Z, Sun W, Wu Q, Tang Z, Bu T, Li C, Chen H. 2019. Genome-wide identification,expression analysis and functional study of the GRAS gene family in Tartary buckwheat(Fagopyrum tataricum). BMC Plant Biology, 19 (1):342.
doi: 10.1186/s12870-019-1951-3 |
| [20] |
Liu X, Widmer A. 2014. Genome-wide comparative analysis of the GRAS gene family in Populus,Arabidopsis and rice. Plant Molecular Biology Reporter, 32 (6):1129-1145.
doi: 10.1007/s11105-014-0721-5 URL |
| [21] |
Liu Y, Shi Y, Zhu N, Zhong S, Bouzayen M, Li Z. 2020. SlGRAS 4 mediates a novel regulatory pathway promoting chilling tolerance in tomato. Plant Biotechnology Journal, 18 (7):1620-1633.
doi: 10.1111/pbi.v18.7 URL |
| [22] |
Lu X, Liu W, Xiang C, Li X, Wang Q, Wang T, Liu Z, Zhang J, Gao L, Zhang W. 2020. Genome-wide characterization of GRAS family and their potential roles in cold tolerance of cucumber(Cucumis sativus L.). International Journal of Molecular Sciences, 21 (11):3857.
doi: 10.3390/ijms21113857 URL |
| [23] | Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar G A, Sonnhammer E L L, Tosatto S C E, Paladin L, Raj S, Richardson L J, Finn R D, Bateman A. 2021. Pfam:The protein families database in 2021. Nucleic Acids Research, 49 (D1):D412-D419. |
| [24] |
Mistry J, Finn R D, Eddy S R, Bateman A, Punta M. 2013. Challenges in homology search:HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Research, 41 (12):e121.
doi: 10.1093/nar/gkt263 URL |
| [25] |
Niu Y, Zhao T, Xu X, Li J. 2017. Genome-wide identification and characterization of GRAS transcription factors in tomato(Solanum lycopersicum). PeerJ, 5(11):e3955.
doi: 10.7717/peerj.3955 URL |
| [26] |
Pei Q, Yu T, Wu T, Yang Q, Gong K, Zhou R, Cui C, Yu Y, Zhao W, Kang X, Cao R, Song X. 2021. Comprehensive identification and analyses of the Hsf gene family in the whole-genome of three Apiaceae species. Horticultural Plant Journal, 7 (5),457-468.
doi: 10.1016/j.hpj.2020.08.005 URL |
| [27] |
Ren L, Sun J, Chen S, Gao J, Dong B, Liu Y, Xia X, Wang Y, Liao Y, Teng N, Fang W, Guan Z, Chen F, Jiang J. 2014. A transcriptomic analysis of Chrysanthemum nankingense provides insights into the basis of low temperature tolerance. BMC Genomics, 15 (1):844.
doi: 10.1186/1471-2164-15-844 |
| [28] |
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio J C, Guirao-Rico S, Librado P, Ramos-Onsins S E, Sánchez-Gracia A. 2017. DnaSP 6:DNA sequence polymorphism analysis of large data sets. Molecular Biology and Evolution, 34 (12):3299-3302.
doi: 10.1093/molbev/msx248 URL |
| [29] |
Shan Z, Luo X, Wu M, Wei L, Fan Z, Zhu Y. 2020. Genome-wide identification and expression of GRAS gene family members in cassava. BMC Plant Biology, 20 (1):46.
doi: 10.1186/s12870-020-2242-8 |
| [30] |
Shang Q M, Li L, Dong C J. 2012. Multiple tandem duplication of the phenylalanine ammonia-lyase genes in Cucumis sativus L.. Planta, 236 (4):1093-1105.
doi: 10.1007/s00425-012-1659-1 URL |
| [31] |
Shannon P, Markiel A, Ozier O, Baliga N S, Wang J T, Ramage D, Amin N, Schwikowski B, Ideker T. 2003. Cytoscape:a software environment for integrated models of biomolecular interaction networks. Genome Research, 13 (11):2498-2504.
doi: 10.1101/gr.1239303 pmid: 14597658 |
| [32] |
Sidhu N S, Pruthi G, Singh S, Bishnoi R, Singla D. 2020. Genome-wide identification and analysis of GRAS transcription factors in the bottle gourd genome. Scientific Reports, 10 (1):14338.
doi: 10.1038/s41598-020-71240-2 pmid: 32868844 |
| [33] |
Song C, Liu Y, Song A, Dong G, Zhao H, Sun W, Ramakrishnan S, Wang Y, Wang S, Li T, Niu Y, Jiang J, Dong B, Xia Y, Chen S, Hu Z, Chen F, Chen S. 2018. The Chrysanthemum nankingense genome provides insights into the evolution and diversification of chrysanthemum flowers and medicinal traits. Molecular Plant, 11 (12):1482-1491.
doi: S1674-2052(18)30308-3 pmid: 30342096 |
| [34] |
Szklarczyk D, Gable A L, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva N T, Morris J H, Bork P, Jensen L J, Mering C V. 2019. STRING v11:protein-protein association networks with increased coverage,supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Research, 47 (D1):D607-D613.
doi: 10.1093/nar/gky1131 |
| [35] |
Tang Mingjia, Xu Jin, Lin Rui, Song Jianing, Yu Jingquan, Zhou Yanhong. 2022. Advances in physiological and molecular mechanism of tomato responses to light and temperature stress. Acta Horticulturae Sinica, 49 (10):2174-2188. (in Chinese)
doi: 10.16420/j.issn.0513-353x.2022-0600 URL |
|
唐明佳, 徐进, 林锐, 宋珈凝, 喻景权, 周艳虹. 2022. 番茄响应光温逆境的生理分子机制研究进展. 园艺学报, 49 (10):2174-2188.
doi: 10.16420/j.issn.0513-353x.2022-0600 URL |
|
| [36] |
To V T, Shi Q, Zhang Y, Shi J, Shen C, Zhang D, Cai W. 2020. Genome-wide analysis of the GRAS gene family in barley(Hordeum vulgare L.). Genes, 11 (5):553.
doi: 10.3390/genes11050553 URL |
| [37] |
Upadhyay R K, Mattoo A K. 2018. Genome-wide identification of tomato(Solanum lycopersicum L.)lipoxygenases coupled with expression profiles during plant development and in response to methyl-jasmonate and wounding. Journal of Plant Physiology, 231:318-328.
doi: S0176-1617(18)30562-5 pmid: 30368230 |
| [38] |
Wang Y, Lü J, Chen D, Zhang J, Qi K, Cheng R, Zhang H, Zhang S. 2018a. Genome-wide identification,evolution,and expression analysis of the KT/HAK/KUP family in pear. Genome, 61 (10):755-765.
doi: 10.1139/gen-2017-0254 URL |
| [39] |
Wang Y, Tang H, Debarry J D, Tan X, Li J, Wang X, Lee T H, Jin H, Marler B, Guo H, Kissinger J C, Paterson A H. 2012. MCScanX:a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Research, 40 (7):e49.
doi: 10.1093/nar/gkr1293 URL |
| [40] |
Wang Y X, Liu Z W, Wu Z J, Li H, Wang W L, Cui X, Zhuang J. 2018b. Genome-wide identification and expression analysis of GRAS family transcription factors in tea plant(Camellia sinensis). Scientific Reports, 8 (1):3949.
doi: 10.1038/s41598-018-22275-z |
| [41] |
Wang Z, Li S, Jian S, Ye F, Wang T, Gong L, Li X. 2021a. Low temperature tolerance is impaired by polystyrene nanoplastics accumulated in cells of barley(Hordeum vulgare L.)plants. Journal of Hazardous Materials, 426:127826.
doi: 10.1016/j.jhazmat.2021.127826 URL |
| [42] |
Wang Z, Wong D C J, Wang Y, Xu G, Ren C, Liu Y, Kuang Y, Fan P, Li S, Xin H, Liang Z. 2021b. GRAS-domain transcription factor PAT 1 regulates jasmonic acid biosynthesis in grape cold stress response. Plant Physiology, 186 (3):1660-1678.
doi: 10.1093/plphys/kiab142 URL |
| [43] |
Won S Y, Jung J A, Kim J S. 2021. Genome-wide analysis of the MADS-Box gene family in chrysanthemum. Computational Biology and Chemistry, 90:107424.
doi: 10.1016/j.compbiolchem.2020.107424 URL |
| [44] | Yang M, Derbyshire M K, Yamashita R A, Marchler-Bauer A. 2020. NCBI’s conserved domain database and tools for protein domain analysis. Current Protocols in Bioinformatics, 69 (1):e90. |
| [45] |
Yuan Y, Fang L, Karungo S K, Zhang L, Gao Y, Li S, Xin H. 2016. Overexpression of VaPAT1,a GRAS transcription factor from Vitis amurensis,confers abiotic stress tolerance in Arabidopsis. Plant Cell Reports, 35 (3):655-666.
doi: 10.1007/s00299-015-1910-x URL |
| [46] |
Zhang H, Liu X, Wang X, Sun M, Song R, Mao P, Jia S. 2021. Genome-wide identification of GRAS gene family and their responses to abiotic stress in Medicago sativa. International Journal of Molecular Sciences, 22 (14):7729.
doi: 10.3390/ijms22147729 URL |
| [47] | Zhang Z L, Ogawa M, Fleet C M, Zentella R, Hu J, Heo J O, Lim J, Kamiya Y, Yamaguchi S, Sun T P. 2011. Scarecrow-like 3 promotes gibberellin signaling by antagonizing master growth repressor DELLA in Arabidopsis. Proceedings of the National Academy of Sciences of the United States of American, 108 (5):2160-2165. |
| [1] | 郑嘉瑞, 杨晓燕, 叶家保, 廖咏玲, 许锋. MYC2转录因子在植物中的功能研究进展[J]. 园艺学报, 2023, 50(4): 896-908. |
| [2] | 刘语诺, 曹亚, 王帅, 杜美霞, 郑林, 陈善春, 邹修平. 柑橘CsMYB41和CsMYB63响应溃疡病菌侵染的表达[J]. 园艺学报, 2023, 50(3): 495-507. |
| [3] | 俞沁含, 李俊铎, 崔莹, 王佳慧, 郑巧玲, 徐伟荣. 山葡萄转录因子VaMYB4a互作蛋白的筛选与鉴定[J]. 园艺学报, 2023, 50(3): 508-522. |
| [4] | 毛可欣, 安淼, 王海荣, 王世金, 吕巍, 郭盈添, 李健, 李国田. 猕猴桃MYB家族成员鉴定及其低温表达分析[J]. 园艺学报, 2023, 50(3): 534-548. |
| [5] | 叶子茂, 申晚霞, 刘梦雨, 王彤, 张晓楠, 余歆, 刘小丰, 赵晓春. R2R3-MYB转录因子CitMYB21对柑橘类黄酮生物合成的影响[J]. 园艺学报, 2023, 50(2): 250-264. |
| [6] | 宋艳红, 陈亚铎, 张晓玉, 宋盼, 刘丽锋, 李刚, 赵霞, 周厚成. 森林草莓FvbHLH130转录因子调控植株提前开花[J]. 园艺学报, 2023, 50(2): 295-306. |
| [7] | 韩睿, 钟雄辉, 陈登辉, 崔建, 乐祥庆, 颉建明, 康俊根. 黑腐病菌效应因子XopR的甘蓝靶标基因BobHLH34的克隆及功能分析[J]. 园艺学报, 2023, 50(2): 319-330. |
| [8] | 任菲, 卢苗苗, 刘吉祥, 陈信立, 刘道凤, 眭顺照, 马婧. 蜡梅胚胎晚期丰富蛋白基因CpLEA的表达及抗性分析[J]. 园艺学报, 2023, 50(2): 359-370. |
| [9] | 田明康, 徐智祥, 刘秀群, 眭顺照, 李名扬, 李志能. 蜡梅AP2亚家族转录因子鉴定及CpAP2-L11功能研究[J]. 园艺学报, 2023, 50(2): 382-396. |
| [10] | 于婷婷, 李欢, 宁源生, 宋建飞, 彭璐琳, 贾竣淇, 张玮玮, 杨洪强. 苹果GRAS全基因组鉴定及其对生长素的响应分析[J]. 园艺学报, 2023, 50(2): 397-409. |
| [11] | 蔺海娇, 梁雨晨, 李玲, 马军, 张璐, 兰振颖, 苑泽宁. 薰衣草CBF途径相关耐寒基因挖掘与调控网络分析[J]. 园艺学报, 2023, 50(1): 131-144. |
| [12] | 赵雪艳, 王琪, 王莉, 王方圆, 王庆, 李艳. 基于比较转录组的延胡索组织差异性表达分析[J]. 园艺学报, 2023, 50(1): 177-187. |
| [13] | 贾鑫, 曾臻, 陈月, 冯慧, 吕英民, 赵世伟. 月季‘月月粉’RcDREB2A的克隆与表达分析[J]. 园艺学报, 2022, 49(9): 1945-1956. |
| [14] | 高彦龙, 吴玉霞, 张仲兴, 王双成, 张瑞, 张德, 王延秀. 苹果ELO家族基因鉴定及其在低温胁迫下的表达分析[J]. 园艺学报, 2022, 49(8): 1621-1636. |
| [15] | 邱子文, 刘林敏, 林永盛, 林晓洁, 李永裕, 吴少华, 杨超. 千层金MbEGS基因的克隆与功能分析[J]. 园艺学报, 2022, 49(8): 1747-1760. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
版权所有 © 2012 《园艺学报》编辑部 京ICP备10030308号-2 国际联网备案号 11010802023439
编辑部地址: 北京市海淀区中关村南大街12号中国农业科学院蔬菜花卉研究所 邮编: 100081
电话: 010-82109523 E-Mail: yuanyixuebao@126.com
技术支持:北京玛格泰克科技发展有限公司