
园艺学报 ›› 2023, Vol. 50 ›› Issue (6): 1255-1268.doi: 10.16420/j.issn.0513-353x.2022-0226
贺威智1,2, 雷伟奇1, 郭祥鑫1, 李蕊莲1,3, 陈冠群1,*( )
)
收稿日期:2022-12-29
修回日期:2023-05-05
出版日期:2023-06-25
发布日期:2023-06-27
通讯作者:
* (E-mail:chengq@sjtu.edu.cn)基金资助:
HE Weizhi1,2, LEI Weiqi1, GUO Xiangxin1, LI Ruilian1,3, CHEN Guanqun1,*( )
)
Received:2022-12-29
Revised:2023-05-05
Published:2023-06-25
Online:2023-06-27
摘要:
利用生物信息鉴定百子莲MYB家族成员,挖掘并验证了调控百子莲蓝色花色形成的关键基因。以全长转录组测序数据作为参考序列,鉴定得到百子莲123个MYB家族成员,包括60个R2R3-MYB、4个3R-MYB、2个4R-MYB和57个1R-MYB基因。对调控类黄酮生物合成的主要亚家族R2R3-MYB进行分析发现,R2-和R3-repeats的序列较为保守,根据系统进化关系分为22个亚组(A1 ~ A22)。特殊的是,相较拟南芥的25个亚组,百子莲缺乏正向调控花色素苷合成的典型亚组S6。与花色形成相关的其他亚组有A18、A17和A16,其成员ApMYB4、ApMYB6、ApMYB7、ApMYB12和ApMYB111定位于细胞核,ApMYB123集中分布于核仁中。在蓝色百子莲花瓣着色过程中,ApMYB12显著上调表达,而ApMYB111显著下调表达。ApMYB4、ApMYB6和ApMYB7在蓝花各发育阶段的表达量均低于白花,预示这些基因可能是百子莲蓝色形成的负调控因子。ApMYB123在白花的开放前期大量表达,说明该基因可能在此阶段促进花瓣合成原花青素。在烟草中过表达ApMYB12,烟草花瓣颜色较野生型变浅,花色素苷含量减少54.55% ~ 62.50%,而黄酮醇含量提高125% ~ 170%,证实ApMYB12调控辅助色素黄酮醇的生物合成,从而影响蓝色的形成。
中图分类号:
贺威智, 雷伟奇, 郭祥鑫, 李蕊莲, 陈冠群. 百子莲MYB家族鉴定及蓝色形成关键基因功能分析[J]. 园艺学报, 2023, 50(6): 1255-1268.
HE Weizhi, LEI Weiqi, GUO Xiangxin, LI Ruilian, CHEN Guanqun. Identification of the MYB Gene Family and Functional Analysis of Key Genes Related to Blue Flower Coloration in Agapanthus praecox[J]. Acta Horticulturae Sinica, 2023, 50(6): 1255-1268.
| 基因名称 Gene name | 正向引物序列(5′-3′) Forward primer sequence | 反向引物序列(5′-3′) Reverse primer sequence |
|---|---|---|
| ApActin | AACACCACCAGACTTATGT | GCCGTTGAACTTGATGAG |
| ApMYB4 | CTACTGGAACACGCACATCAA | CTCTCGTCGGTAGAATCCTCAT |
| ApMYB6 | AGTTGTCGGCTTCGTTGGA | TGTCTGTTCTTCCTGGTAGTCTT |
| ApMYB7 | GACCTATCCATTAGCCTTCCTTATC | CTGTTGTTCTTGTCGCTGTCA |
| ApMYB12 | GACCTCGACCAAGACTTGTTAC | ATCTAGCACCTTCTCCTCAACTT |
| ApMYB111 | GGCACGGTATCAGCGGTAT | TTCTCATGCTCGTCTCTTGTTG |
| ApMYB123 | CCTTATCATCCGACTCCACTCT | TTGCTGAGGTGACTGTTCCA |
| pHB-ApMYB4 | CTCTCTCTCAAGCTTGGATCCATGGGTAGGTCTCCATGTTG | CATACTAGTGAGCTCCTGCAGAGCATTATACCCATCCTGTA |
| pHB-ApMYB6 | CTCTCTCTCAAGCTTGGATCCATGGGTAGATCTCCTTGTTG | CATACTAGTGAGCTCCTGCAGCTTCATCTGAAAACTTCTAT |
| pHB-ApMYB7 | CTCTCTCTCAAGCTTGGATCCATGGGAAGGTCCCCATGTTG | CATACTAGTGAGCTCCTGCAGTATCCATCCTTCCTCCAACG |
| pHB-ApMYB12 | CTCTCTCTCAAGCTTGGATCCATGGGAAGGGCTCCATGTTG | CATACTAGTGAGCTCCTGCAGCAACACATCTGAAAGAAGCC |
| pHB-ApMYB111 | CTCTCTCTCAAGCTTGGATCCATGGGGAGAGTTCCATGTTG | CATACTAGTGAGCTCCTGCAGTAATTTGCACCCACTCTCCC |
| pHB-ApMYB123 | CTCTCTCTCAAGCTTGGATCCATGGGGAGAGCTCCATGCTG | CATACTAGTGAGCTCCTGCAGAATCAACAGAGGCTCTGCAG |
| NtPAL | ATTGAGGTCATCCGTTCTGC | ACCGTGTAACGCCTTGTTTC |
| Nt4CL | TCATTGACGAGGATGACGAG | TGGGATGGTTGAGAAGAAGG |
| NtCHS | TTGTTCGAGCTTGTCTCTGC | AGCCCAGGAACATCTTTGAG |
| NtCHI | GTCAGGCCATTGAAAAGCTC | CTAATCGTCAATGCCCCAAC |
| NtFLS | TTTGGCACTTGGTGTTGTGG | ACTTGACATCATACCAATGG |
| NtDFR | AACCAACAGTCAGGGGAATG | TTGGACATCGACAGTTCCAG |
| NtUFGT | ATGTTGAAGGGCTAAAAGAAAGAGC | CAAGTCCCAGCTGATACATATTCCC |
| NtGAPDH | GGTGTCCACAGACTTCGTGG | GACTCCTCACAGCAGCACCA |
表1 PCR引物列表
Table 1 Primer list for PCR
| 基因名称 Gene name | 正向引物序列(5′-3′) Forward primer sequence | 反向引物序列(5′-3′) Reverse primer sequence |
|---|---|---|
| ApActin | AACACCACCAGACTTATGT | GCCGTTGAACTTGATGAG |
| ApMYB4 | CTACTGGAACACGCACATCAA | CTCTCGTCGGTAGAATCCTCAT |
| ApMYB6 | AGTTGTCGGCTTCGTTGGA | TGTCTGTTCTTCCTGGTAGTCTT |
| ApMYB7 | GACCTATCCATTAGCCTTCCTTATC | CTGTTGTTCTTGTCGCTGTCA |
| ApMYB12 | GACCTCGACCAAGACTTGTTAC | ATCTAGCACCTTCTCCTCAACTT |
| ApMYB111 | GGCACGGTATCAGCGGTAT | TTCTCATGCTCGTCTCTTGTTG |
| ApMYB123 | CCTTATCATCCGACTCCACTCT | TTGCTGAGGTGACTGTTCCA |
| pHB-ApMYB4 | CTCTCTCTCAAGCTTGGATCCATGGGTAGGTCTCCATGTTG | CATACTAGTGAGCTCCTGCAGAGCATTATACCCATCCTGTA |
| pHB-ApMYB6 | CTCTCTCTCAAGCTTGGATCCATGGGTAGATCTCCTTGTTG | CATACTAGTGAGCTCCTGCAGCTTCATCTGAAAACTTCTAT |
| pHB-ApMYB7 | CTCTCTCTCAAGCTTGGATCCATGGGAAGGTCCCCATGTTG | CATACTAGTGAGCTCCTGCAGTATCCATCCTTCCTCCAACG |
| pHB-ApMYB12 | CTCTCTCTCAAGCTTGGATCCATGGGAAGGGCTCCATGTTG | CATACTAGTGAGCTCCTGCAGCAACACATCTGAAAGAAGCC |
| pHB-ApMYB111 | CTCTCTCTCAAGCTTGGATCCATGGGGAGAGTTCCATGTTG | CATACTAGTGAGCTCCTGCAGTAATTTGCACCCACTCTCCC |
| pHB-ApMYB123 | CTCTCTCTCAAGCTTGGATCCATGGGGAGAGCTCCATGCTG | CATACTAGTGAGCTCCTGCAGAATCAACAGAGGCTCTGCAG |
| NtPAL | ATTGAGGTCATCCGTTCTGC | ACCGTGTAACGCCTTGTTTC |
| Nt4CL | TCATTGACGAGGATGACGAG | TGGGATGGTTGAGAAGAAGG |
| NtCHS | TTGTTCGAGCTTGTCTCTGC | AGCCCAGGAACATCTTTGAG |
| NtCHI | GTCAGGCCATTGAAAAGCTC | CTAATCGTCAATGCCCCAAC |
| NtFLS | TTTGGCACTTGGTGTTGTGG | ACTTGACATCATACCAATGG |
| NtDFR | AACCAACAGTCAGGGGAATG | TTGGACATCGACAGTTCCAG |
| NtUFGT | ATGTTGAAGGGCTAAAAGAAAGAGC | CAAGTCCCAGCTGATACATATTCCC |
| NtGAPDH | GGTGTCCACAGACTTCGTGG | GACTCCTCACAGCAGCACCA |
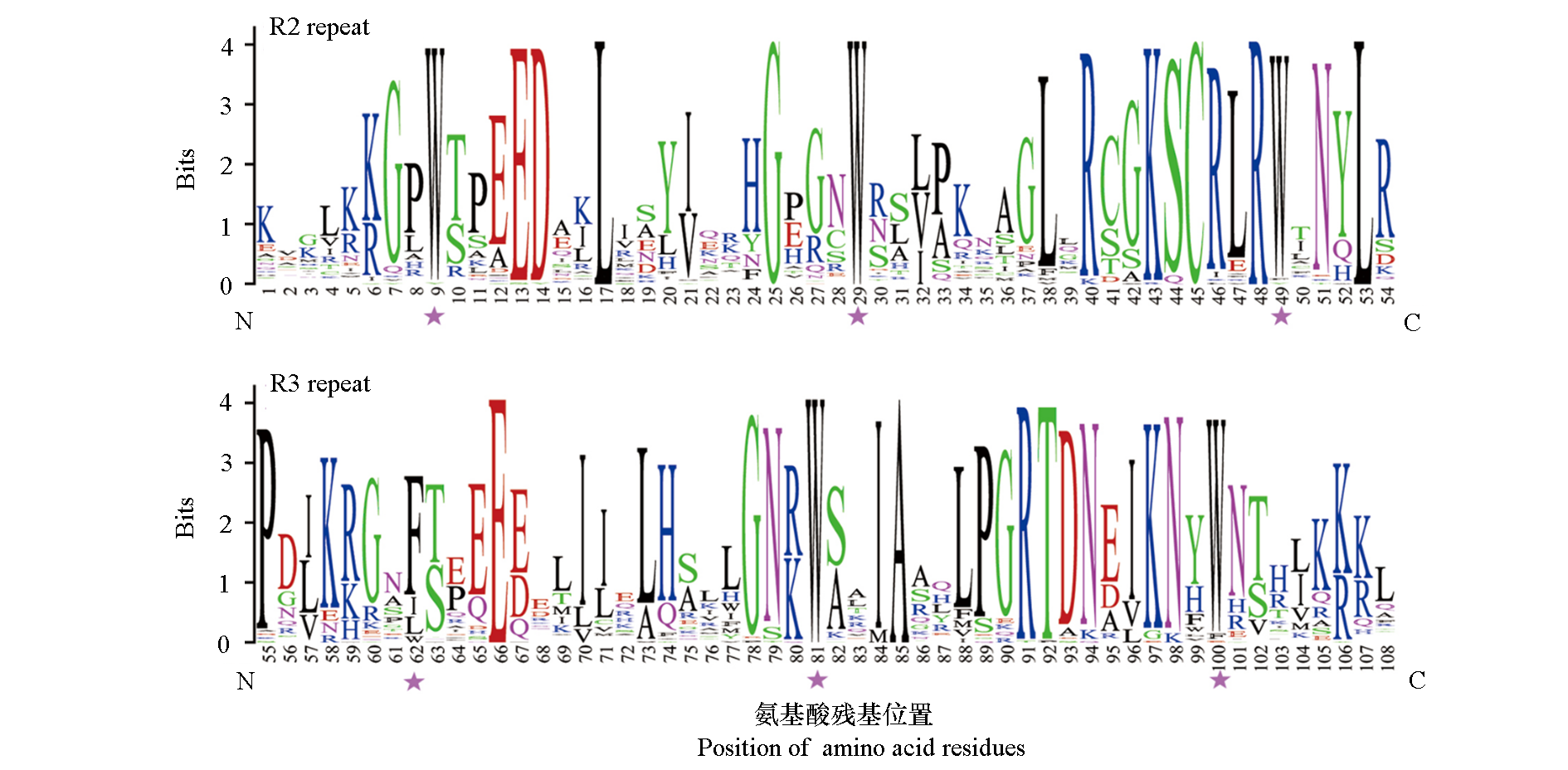
图2 百子莲R2R3-MYB结构域序列分析 Bits值表示序列中每个氨基酸位点的保守程度,星号代表MYB结构域中的保守氨基酸残基色氨酸(W)。
Fig. 2 Consensus sequence of R2R3-MYB domains in Agapanthus praecox The bit score of Y-axis indicated the conservation rate for each X-axis amino acid position in the sequence. Asterisks indicate the conserved residues Trp(W)in the MYB domain.
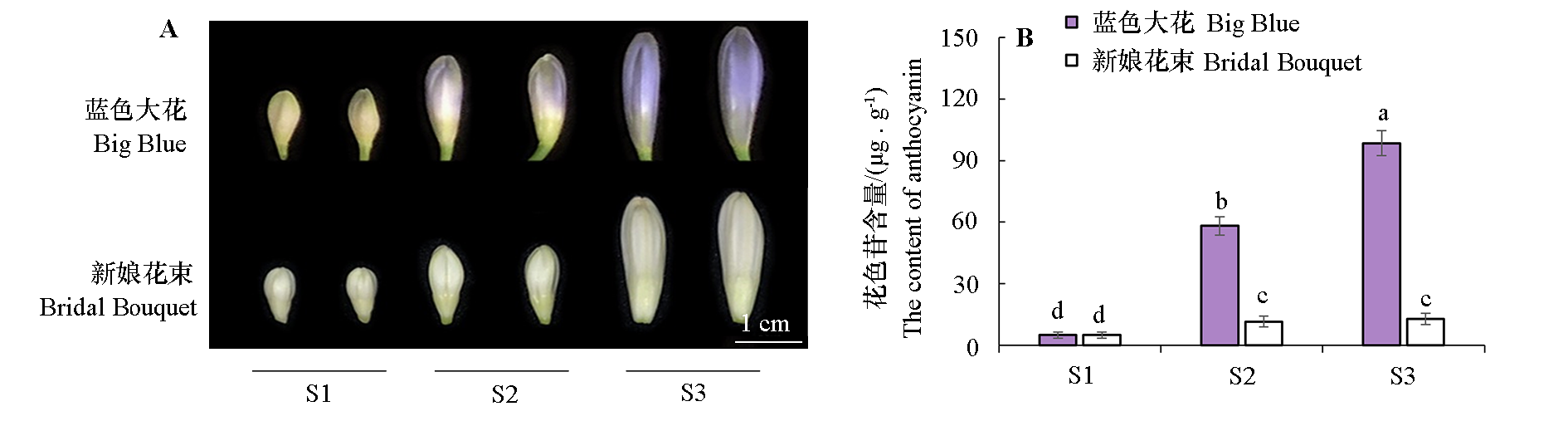
图3 蓝花‘蓝色大花’与白花‘新娘花束’百子莲花瓣发育过程中的花色素苷含量 S1:现蕾期;S2:花蕾伸长期;S3:开放前期。不同小写字母表示花色素苷含量差异显著(P < 0.05)。
Fig. 3 The anthocyanin contents during flower development in blue-flowered‘Big Blue’and white-flowered‘Bridal Bouquet’ S1:Bud emergence period;S2:Bud elongation period;S3:Pre-opening period. The different lowercase letters respectively indicate the significant difference in the anthocyanin content(P < 0.05).

图4 蓝花‘蓝色大花’与白花‘新娘花束’百子莲花瓣着色过程中类黄酮合成相关MYB基因表达热图(A)和蛋白保守基序(B) S1:现蕾期;S2:花蕾伸长期;S3:开放前期。
Fig. 4 Heat map of expression(A)and protein conserved motifs(B)of MYB genes related to flavonoid biosynthesis during flower coloration in blue-flowered‘Big Blue’and white-flowered‘Bridal Bouquet’ S1:Bud emergence period;S2:Bud elongation period;S3:Pre-opening period.

图6 烟草转ApMYB12株系(7#、8#)及野生型(WT)的花色表型、花瓣和叶片总黄酮醇和总花色素苷含量 **代表野生型与转基因株系间差异极显著(P < 0.01)。
Fig. 6 Flower color phenotypes, flavonol and anthocyanin content in petal and leaf of ApMYB12-transgenic tobacco lines(7#,8#)and wild type ** represents the significant difference between wild-type and transgenic lines(P < 0.01).
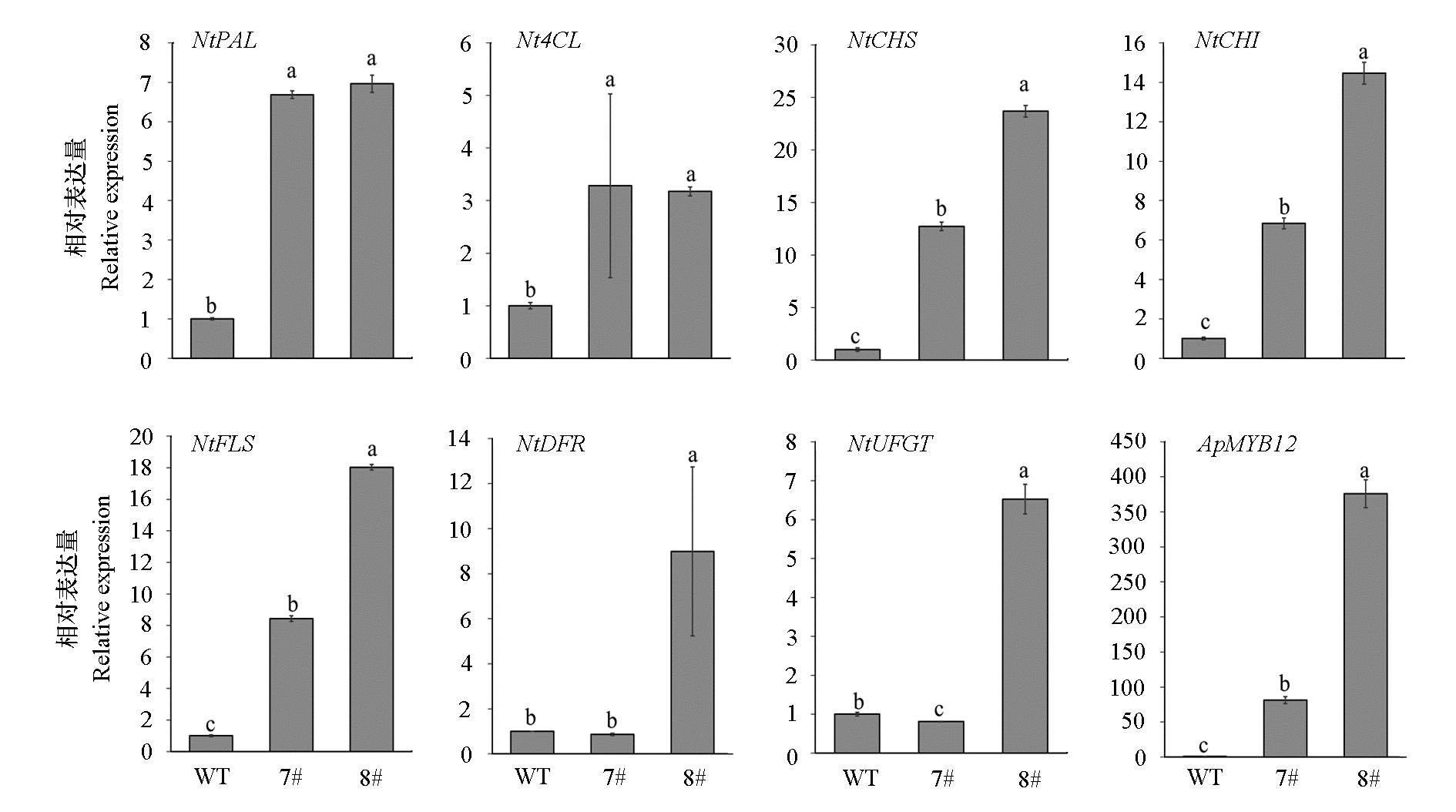
图8 烟草转ApMYB12株系(7#、8#)及野生型(WT)中类黄酮合成途径关键基因的表达量 不同小写字母表示野生型和转基因株系间具有显著差异(P < 0.05)。
Fig. 8 The expression of key genes in the flavonoid biosynthesis pathway of of ApMYB12-transgenic tobacco lines(7#,8#)and wild type Different lowercase letters indicate significant differences between wild-type and transgenic lines(P < 0.05).
| [1] |
Abe H, Urao T, Ito T, Seki M, Shinozaki K. 2003. Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell, 15 (1):63-78.
doi: 10.1105/tpc.006130 URL |
| [2] |
Bendahmane M, Dubois A, Raymond O, Bris M L. 2013. Genetics and genomics of flower initiation and development in roses. Journal of Experimental Botany, 64 (4):847-857.
doi: 10.1093/jxb/ers387 pmid: 23364936 |
| [3] |
Bloor S J, Falshaw R. 2000. Covalently linked anthocyanin-flavonol pigments from blue Agapanthus flowers. Phytochemistry, 53 (5):575-579.
pmid: 10724183 |
| [4] | Cao Y, Han Y, Li D, Lin Y, Cai Y. 2016. MYB transcription factors in Chinese pear(Pyrus bretschneideri Rehd.):Genome-wide identification, classification, and expression profiling during fruit development. Frontiers in Plant Science, 7:577. |
| [5] |
Cao Y, Xie L, Ma Y, Ren C, Xing M, Fu Z, Wu X, Yin X, Xu C, Li X. 2019. PpMYB15 and PpMYBF 1 transcription factors are involved in regulating flavonol biosynthesis in peach fruit. Journal of Agricultural and Food Chemistry, 67 (2):644-652.
doi: 10.1021/acs.jafc.8b04810 URL |
| [6] |
Cba B, Srg A, Ap C, Pkta B. 2020. COP1 mediates light-dependent regulation of flavonol biosynthesis through HY5 in Arabidopsis. Plant Science, 303:110760.
doi: 10.1016/j.plantsci.2020.110760 URL |
| [7] |
Chen C J, Chen H, Zhang Y, Thomas H R, Frank M H, He Y H, Xia R. 2020a. TBtools:an integrative toolkit developed for interactive analyses of big biological data. Molecular Plant, 13 (8):1194-1202.
doi: 10.1016/j.molp.2020.06.009 URL |
| [8] |
Chen G, He W, Guo X, Pan J. 2021. Genome-wide identification, classification and expression analysis of the MYB transcription factor family in Petunia. International Journal of Molecular Sciences, 22 (9):4838.
doi: 10.3390/ijms22094838 URL |
| [9] |
Chen G, Xu P, Pan J, Li Y, Zhou J, Kuang H, Lian H. 2020b. Inhibition of FvMYB 10 transcriptional activity promotes color loss in strawberry fruit. Plant Science, 298:110578.
doi: 10.1016/j.plantsci.2020.110578 URL |
| [10] |
Du H, Yang S S, Liang Z, Feng B R, Liu L, Huang Y B, Tang Y X. 2012. Genome-wide analysis of the MYB transcription factor superfamily in soybean. BMC Plant Biology, 12:106.
doi: 10.1186/1471-2229-12-106 pmid: 22776508 |
| [11] |
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L. 2010. MYB transcription factors in Arabidopsis. Trends in Plant Science, 15 (10):573-581.
doi: 10.1016/j.tplants.2010.06.005 URL |
| [12] |
Fan H, Cui M, Li N, Li X, Lin Y. 2020. Genome-wide identification and expression analyses of R2R3-MYB transcription factor genes from two orchid species. PeerJ, 8:e9781.
doi: 10.7717/peerj.9781 URL |
| [13] | Fornalé S, Lopez E, Salazar-Henao J E, Fernández-Nohales P, Rigau J, Caparros-Ruiz D. 2014. AtMYB7,a new player in the regulation of UV-sunscreens in Arabidopsis thaliana. Plant & Cell Physiology, 55 (3):507-516. |
| [14] |
Gatica-Arias A, Farag M A, Stanke M, Matoušek J, Wessjohann L, Weber G. 2012. Flavonoid production in transgenic hop(Humulus lupulus L.) altered by PAP1/MYB75 from Arabidopsis thaliana. Plant Cell Reports, 31 (1):111-119.
doi: 10.1007/s00299-011-1144-5 pmid: 21912858 |
| [15] |
Golz J F, Allen P J, Li S F, Parish R W, Jayawardana N U, Bacic A, Doblin M S. 2018. Layers of regulation - Insights into the role of transcription factors controlling mucilage production in the Arabidopsis seed coat. Plant Science, 272:179-192.
doi: 10.1016/j.plantsci.2018.04.021 URL |
| [16] |
Han Y, Yu J, Zhao T, Cheng T, Wang J, Yang W, Pan H, Zhang Q. 2019. Dissecting the genome-wide evolution and function of R2R3-MYB transcription factor family in Rosa chinensis. Genes, 10 (10):823.
doi: 10.3390/genes10100823 URL |
| [17] |
He Z, Zhang H, Gao S, Lercher M J, Chen W H, Hu S. 2016. Evolview v2:an online visualization and management tool for customized and annotated phylogenetic trees. Nucleic Acids Research, 44 (W1):W236-W241.
doi: 10.1093/nar/gkw370 URL |
| [18] |
Hegvold A B, Gabrielsen O S. 1996. The importance of the linker connecting the repeats of the c-Myb oncoprotein may be due to a positioning function. Nucleic Acids Research, 24 (20):3990-3995.
pmid: 8918802 |
| [19] |
Jeff V, Cahid C, Eunseog Y, Chen J, Cazzonelli C I, Georg S. 2012. Transgene silencing and transgene-derived siRNA production in tobacco plants homozygous for an introduced AtMYB90 construct. PLoS ONE, 7 (2):e30141.
doi: 10.1371/journal.pone.0030141 URL |
| [20] |
Jiang C, Gu J, Chopra S, Gu X, Peterson T. 2004. Ordered origin of the typical two- and three-repeat Myb genes. Gene, 326:13-22.
pmid: 14729259 |
| [21] | Leighton F M. 1965. The genus Agapanthus L'Héritier. South African Journal of Botany, 4:1-50. |
| [22] |
Li H Y, Yue Y Z, Ding W J, Chen G W, Li L, Li Y L, Shi T T, Yang X L, Wang L G. 2020. Genome-wide identification, classification, and expression profiling reveals R2R3-MYB transcription factors related to monoterpenoid biosynthesis in Osmanthus fragrans. Genes, 11 (4):353.
doi: 10.3390/genes11040353 URL |
| [23] |
Li J, Liu H, Yang C, Wang J, Xu L. 2019. Genome-wide identification of MYB genes and expression analysis under different biotic and abiotic stresses in Helianthus annuus L. Industrial Crops and Products, 143:111924.
doi: 10.1016/j.indcrop.2019.111924 URL |
| [24] |
Li Maofu, Yang Yuan, Wang Hua, Fan Youwei, Sun Pei, Jin Wanmei. 2022. Analysis the function of R2R3 MYB transcription factor RhMYB113c on regulating anthocyanin synthesis in Rosa hybrida. Acta Horticulturae Sinica, 49 (9):1957-1966. (in Chinese)
doi: 10.16420/j.issn.0513-353x.2021-0630 URL |
|
李茂福, 杨媛, 王华, 范又维, 孙佩, 金万梅. 2022. 月季中R2R3-MYB基因RhMYB113c调控花青素苷合成. 园艺学报, 49 (9):1957-1966.
doi: 10.16420/j.issn.0513-353x.2021-0630 URL |
|
| [25] | Li Qing-yun, Tang Qian-wen, Chen Guan-qun, Shen Xiao-hui. 2023. Extraction, identification and physical-chemical stability of anthocyanins from two hydrangea varieties. Guihaia, 43 (4):765-776. (in Chinese) |
| 李清韵, 唐倩雯, 陈冠群, 申晓辉. 2023. 两个八仙花品种花色苷的提取、鉴定和理化稳定性. 广西植物, 43 (4):765-776. | |
| [26] | Li Xiang, Duan Jing-jing, Luo Xiao-ning, Zhang Yan-long, Niu Li-xin, Shi Qian-qian. 2019. Formation mechanism of different tree peony flower colors by anatomy and biochemistry. Journal of Northeast Forestry University, 47 (3):38-43. (in Chinese) |
| 李想, 段晶晶, 罗小宁, 张延龙, 牛立新, 史倩倩. 2019. 依据理化性质分析牡丹花色形成的影响因素. 东北林业大学学报, 47 (3):38-43. | |
| [27] |
Li Z, Peng R, Tian Y, Han H, Xu J, Yao Q. 2016. Genome-wide identification and analysis of the MYB transcription factor superfamily in Solanum lycopersicum. Plant Cell Physiology, 57 (8):1657-1677.
doi: 10.1093/pcp/pcw091 URL |
| [28] |
Mori S, Otani M, Kobayashi H, Nakano M. 2014. Isolation and characterization of the dihydroflavonol 4-reductase gene in the monocotyledonous ornamental Agapanthus praecox ssp. orientalis(Leighton)Leighton. Scientia Horticulturae, 166:24-30.
doi: 10.1016/j.scienta.2013.12.009 URL |
| [29] |
Nemie-Feyissa D, Heidari B, Blaise M, Lillo C. 2015. Analysis of interactions between heterologously produced bHLH and MYB proteins that regulate anthocyanin biosynthesis:quantitative interaction kinetics by Microscale Thermophoresis. Phytochemistry, 111:21-26.
doi: 10.1016/j.phytochem.2015.01.004 pmid: 25659750 |
| [30] | Nozawa M, Kawahara Y, Nei M. 2007. Genomic drift and copy number variation of sensory receptor genes in humans. Proceedings of the National Academy of Sciences of the United States of America, 104 (51):20421-20426. |
| [31] |
Ogata K, Morikawa S, Nakamura H, Sekikawa A, Inoue T, Kanai H, Sarai A, Ishii S, Nishimura Y. 1994. Solution structure of a specific DNA complex of the Myb DNA-binding domain with cooperative recognition helices. Cell, 79 (4):639-648.
doi: 10.1016/0092-8674(94)90549-5 pmid: 7954830 |
| [32] |
Paz-Ares J, Ghosal D, Wienand U, Peterson P A, Saedler H. 1987. The regulatory c1 locus of Zea mays encodes a protein with homology to MYB proto-oncogene products and with structural similarities to transcriptional activators. The EMBO Journal, 6 (12):3553-3558.
doi: 10.1002/embj.1987.6.issue-12 URL |
| [33] |
Rosinski J A, Atchley W R. 1998. Molecular evolution of the Myb family of transcription factors:evidence for polyphyletic origin. Journal of Molecular Evolution, 46 (1):74-83.
doi: 10.1007/pl00006285 pmid: 9419227 |
| [34] |
Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B. 2007. Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant Journal, 50 (4):660-677.
doi: 10.1111/j.1365-313X.2007.03078.x pmid: 17419845 |
| [35] |
Stracke R, Werber M, Weisshaar B. 2001. The R2R3-MYB gene family in Arabidopsis thaliana. Current Opinion in Plant Biology, 4 (5):447-456.
doi: 10.1016/s1369-5266(00)00199-0 pmid: 11597504 |
| [36] |
Wang L, Albert N W, Zhang H, Arathoon S, Boase M R, Ngo H, Schwinn K E, Davies K M, Lewis D H. 2014. Temporal and spatial regulation of anthocyanin biosynthesis provide diverse flower colour intensities and patterning in Cymbidium orchid. Planta, 240 (5):983-1002.
doi: 10.1007/s00425-014-2152-9 pmid: 25183255 |
| [37] |
Wang X, Niu Y, Zheng Y. 2021. Multiple functions of MYB transcription factors in abiotic stress responses. International Journal of Molecular Sciences, 22 (11):6125.
doi: 10.3390/ijms22116125 URL |
| [38] |
Wang X C, Wu J, Guan M L, Zhao C H, Geng P, Zhao Q. 2020. Arabidopsis MYB 4 plays dual roles in flavonoid biosynthesis. The Plant Journal, 101 (3):637-652.
doi: 10.1111/tpj.v101.3 URL |
| [39] |
Xu Q, He J, Dong J, Hou X, Zhang X. 2018. Genomic survey and expression profiling of the MYB gene family in watermelon. Horticultural Plant Journal, 4 (1):1-15.
doi: 10.1016/j.hpj.2017.12.001 URL |
| [40] |
Yabuya T, Nakamura M, Iwashina T, Yamaguchi M, Takehara T. 1997. Anthocyanin-flavone copigmentation in bluish purple flowers of Japanese garden iris(Iris ensata Thunb.). Euphytica, 98:163-167.
doi: 10.1023/A:1003152813333 URL |
| [41] |
Ye Zimao, Shen Wanxia, Liu Mengyu, Wang Tong, Zhang Xiaonan, Yu Xin, Liu Xiaofeng, Zhao Xiaochun. 2023. Effect of R2R3-MYB transcription factor CitMYB 21 on flavonoids biosynthesis in citrus. Acta Horticulturae Sinica, 50 (2):250-264. (in Chinese)
doi: 10.16420/j.issn.0513-353x.2021-1188 |
|
叶子茂, 申晚霞, 刘梦雨, 王彤, 张晓楠, 余歆, 刘小丰, 赵晓春. 2023. R2R3-MYB转录因子CitMYB21对柑橘类黄酮生物合成的影响. 园艺学报, 50 (2):250-264.
doi: 10.16420/j.issn.0513-353x.2021-1188 URL |
|
| [42] | Zhang Bao-zhi. 2013. Study on the composition of anthocyanins and physiological characteristics of flowering process in Jiangnan Peony[M. D. Dissertation]. Shanghai: Shanghai Jiao Tong University. (in Chinese) |
| 张宝智. 2013. 江南牡丹花色素组成与开花过程生理特征研究[硕士论文]. 上海: 上海交通大学. | |
| [43] | Zhang Ning, Wang Di. 2004. Study on the efficient genetic transformation system of tobacco mediated by Agrobacterium. Gansu Agricultural Science and Technology,(9):11-13. (in Chinese) |
| 张宁, 王蒂. 2004. 农杆菌介导的烟草高效遗传转化体系研究. 甘肃农业科技,(9):11-13. | |
| [44] |
Zhou M, Zhang K, Sun Z, Yan M, Wu Y. 2017. LNK1 and LNK 2 corepressors interact with the MYB3 transcription factor in phenylpropanoid biosynthesis. Plant Physiology, 174 (3):1348-1358.
doi: 10.1104/pp.17.00160 URL |
| [1] | 王同欢, 吴雨馨, 武艺圆, 李鑫鑫, 刘梦阳, 杨莲莲, 李佳鹏, 张忠山, 曹访, 仲雪婷, 王占旗. 菊花脑GRAS家族鉴定及其低温胁迫响应表达分析[J]. 园艺学报, 2023, 50(4): 815-830. |
| [2] | 郑嘉瑞, 杨晓燕, 叶家保, 廖咏玲, 许锋. MYC2转录因子在植物中的功能研究进展[J]. 园艺学报, 2023, 50(4): 896-908. |
| [3] | 刘语诺, 曹亚, 王帅, 杜美霞, 郑林, 陈善春, 邹修平. 柑橘CsMYB41和CsMYB63响应溃疡病菌侵染的表达[J]. 园艺学报, 2023, 50(3): 495-507. |
| [4] | 毛可欣, 安淼, 王海荣, 王世金, 吕巍, 郭盈添, 李健, 李国田. 猕猴桃MYB家族成员鉴定及其低温表达分析[J]. 园艺学报, 2023, 50(3): 534-548. |
| [5] | 叶子茂, 申晚霞, 刘梦雨, 王彤, 张晓楠, 余歆, 刘小丰, 赵晓春. R2R3-MYB转录因子CitMYB21对柑橘类黄酮生物合成的影响[J]. 园艺学报, 2023, 50(2): 250-264. |
| [6] | 宋艳红, 陈亚铎, 张晓玉, 宋盼, 刘丽锋, 李刚, 赵霞, 周厚成. 森林草莓FvbHLH130转录因子调控植株提前开花[J]. 园艺学报, 2023, 50(2): 295-306. |
| [7] | 韩睿, 钟雄辉, 陈登辉, 崔建, 乐祥庆, 颉建明, 康俊根. 黑腐病菌效应因子XopR的甘蓝靶标基因BobHLH34的克隆及功能分析[J]. 园艺学报, 2023, 50(2): 319-330. |
| [8] | 田明康, 徐智祥, 刘秀群, 眭顺照, 李名扬, 李志能. 蜡梅AP2亚家族转录因子鉴定及CpAP2-L11功能研究[J]. 园艺学报, 2023, 50(2): 382-396. |
| [9] | 蔺海娇, 梁雨晨, 李玲, 马军, 张璐, 兰振颖, 苑泽宁. 薰衣草CBF途径相关耐寒基因挖掘与调控网络分析[J]. 园艺学报, 2023, 50(1): 131-144. |
| [10] | 贾鑫, 曾臻, 陈月, 冯慧, 吕英民, 赵世伟. 月季‘月月粉’RcDREB2A的克隆与表达分析[J]. 园艺学报, 2022, 49(9): 1945-1956. |
| [11] | 许海峰, 王中堂, 陈新, 刘志国, 王利虎, 刘平, 刘孟军, 张琼. 冬枣果皮着色相关类黄酮靶向代谢组学及潜在MYB转录因子分析[J]. 园艺学报, 2022, 49(8): 1761-1771. |
| [12] | 郑林, 王帅, 刘语诺, 杜美霞, 彭爱红, 何永睿, 陈善春, 邹修平. 柑橘响应黄龙病菌侵染的NAC基因的克隆及表达分析[J]. 园艺学报, 2022, 49(7): 1441-1457. |
| [13] | 钱婕妤, 蒋玲莉, 郑钢, 陈佳红, 赖吴浩, 许梦晗, 付建新, 张超. 百日草花青素苷合成相关MYB转录因子筛选及ZeMYB9功能研究[J]. 园艺学报, 2022, 49(7): 1505-1518. |
| [14] | 陈道宗, 刘镒, 沈文杰, 朱博, 谭晨. 白菜、甘蓝和甘蓝型油菜PAP1/2同源基因的鉴定及分析[J]. 园艺学报, 2022, 49(6): 1301-1312. |
| [15] | 王妍, 孙政, 冯珊, 袁心怡, 仲林林, 曾云流, 傅小鹏, 程运江, 包满珠, 张帆. 香石竹DcERF-1转录因子对切花衰老的负调控作用[J]. 园艺学报, 2022, 49(6): 1313-1326. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
版权所有 © 2012 《园艺学报》编辑部 京ICP备10030308号-2 国际联网备案号 11010802023439
编辑部地址: 北京市海淀区中关村南大街12号中国农业科学院蔬菜花卉研究所 邮编: 100081
电话: 010-82109523 E-Mail: yuanyixuebao@126.com
技术支持:北京玛格泰克科技发展有限公司