
园艺学报 ›› 2023, Vol. 50 ›› Issue (4): 802-814.doi: 10.16420/j.issn.0513-353x.2022-0023
收稿日期:2022-04-21
修回日期:2022-12-11
出版日期:2023-04-25
发布日期:2023-04-27
通讯作者:
**(E-mail:liang_sun@cau.edu.cn)
作者简介:第一联系人:*共同第一作者
基金资助:
XU Yue,*, LI Zixiong,*, CHEN Jie, SUN Liang( )
)
Received:2022-04-21
Revised:2022-12-11
Online:2023-04-25
Published:2023-04-27
Contact:
**(E-mail:liang_sun@cau.edu.cn)
摘要:
番茄冬春季设施生产中为提高坐果率而采用的2,4-D蘸花措施会导致畸形果率增加,其转录层面的机理尚不明确。本研究中对以醋栗番茄LA1589(Solanum pimpinellifolium accession LA1589)为背景的果形近等基因系WT、sun和ovate花蕾喷施2,4-D,而后对花蕾转录组与成熟果实果形进行分析。结果发现,3个近等基因系在2,4-D处理3周后所形成的子房及其发育而来的成熟果实的果梗端均伸长,并最终导致子房与果实的形状指数增大,而在此过程中两个果形位点与2,4-D间存在互作。转录组分析发现,2,4-D处理可能通过激活生长素信号通路,促进生长素极性运输与细胞分裂素合成以及增加ATHB和bHLH转录因子的积累来改变果形;sun与ovate位点通过与2,4-D互作进一步增强了生长素积累、信号转导与极性运输,扩大了受影响的转录因子范围,最终强化了2,4-D对果形的影响;同时该互作还在sun与ovate近等基因系中分别上调了BRX与CKX5基因的表达来进一步增强上述过程;最后,ovate与2,4-D的互作导致两个SWEET基因的表达下调并可能影响糖类向子房中的运输。
中图分类号:
徐悦, 李子雄, 陈婕, 孙亮. 2,4-D调控番茄果形的转录基础[J]. 园艺学报, 2023, 50(4): 802-814.
XU Yue, LI Zixiong, CHEN Jie, SUN Liang. Transcriptional Bases of the Regulation of 2,4-D on Tomato Fruit Shape[J]. Acta Horticulturae Sinica, 2023, 50(4): 802-814.
| 2,4-D处理/(mmol · L-1) 2,4-D treatment | 最大宽度/cm Maximum width | 最大长度/cm Maximum height | 果形指数 Fruit shape index | 果梗端比例 Proximal end ratio | ||
|---|---|---|---|---|---|---|
| 基因型 Genotype | WT | 0 | 1.20 ± 0.01 a | 1.23 ± 0.01 d | 1.02 ± 0.01 d | 0.13 ± 0.01 d |
| 270 | 0.95 ± 0.03 b | 1.35 ± 0.02 cd | 1.43 ± 0.03 c | 0.31 ± 0.03 bc | ||
| sun | 0 | 1.14 ± 0.04 a | 1.65 ± 0.03 abc | 1.45 ± 0.03 c | 0.15 ± 0.03 d | |
| 270 | 1.65 ± 0.03 b | 1.92 ± 0.03 a | 2.48 ± 0.27 a | 0.36 ± 0.03 b | ||
| ovate | 0 | 1.19 ± 0.03 a | 1.52 ± 0.12 bcd | 1.28 ± 0.13 cd | 0.21 ± 0.03 cd | |
| 270 | 0.92 ± 0.12 b | 1.80 ± 0.25 ab | 1.96 ± 0.05 b | 0.49 ± 0.08 a | ||
| 因素效应 Factorial effects (P-value) | 基因型Genotype | 0.02 | 1.93e-05 | 1.03e-06 | 2.44e-4 | |
| 处理Treatment | 4.56e-07 | 1.28e-03 | 4.24e-08 | 5.36e-08 | ||
| 基因型与处理互作 Genotype × Treatment | 0.31 | 0.45 | 2.94e-03 | 0.17 | ||
表1 2,4-D处理后果实表型分析
Table 1 Morphological analysis of fruits after 2,4-D treatment
| 2,4-D处理/(mmol · L-1) 2,4-D treatment | 最大宽度/cm Maximum width | 最大长度/cm Maximum height | 果形指数 Fruit shape index | 果梗端比例 Proximal end ratio | ||
|---|---|---|---|---|---|---|
| 基因型 Genotype | WT | 0 | 1.20 ± 0.01 a | 1.23 ± 0.01 d | 1.02 ± 0.01 d | 0.13 ± 0.01 d |
| 270 | 0.95 ± 0.03 b | 1.35 ± 0.02 cd | 1.43 ± 0.03 c | 0.31 ± 0.03 bc | ||
| sun | 0 | 1.14 ± 0.04 a | 1.65 ± 0.03 abc | 1.45 ± 0.03 c | 0.15 ± 0.03 d | |
| 270 | 1.65 ± 0.03 b | 1.92 ± 0.03 a | 2.48 ± 0.27 a | 0.36 ± 0.03 b | ||
| ovate | 0 | 1.19 ± 0.03 a | 1.52 ± 0.12 bcd | 1.28 ± 0.13 cd | 0.21 ± 0.03 cd | |
| 270 | 0.92 ± 0.12 b | 1.80 ± 0.25 ab | 1.96 ± 0.05 b | 0.49 ± 0.08 a | ||
| 因素效应 Factorial effects (P-value) | 基因型Genotype | 0.02 | 1.93e-05 | 1.03e-06 | 2.44e-4 | |
| 处理Treatment | 4.56e-07 | 1.28e-03 | 4.24e-08 | 5.36e-08 | ||
| 基因型与处理互作 Genotype × Treatment | 0.31 | 0.45 | 2.94e-03 | 0.17 | ||
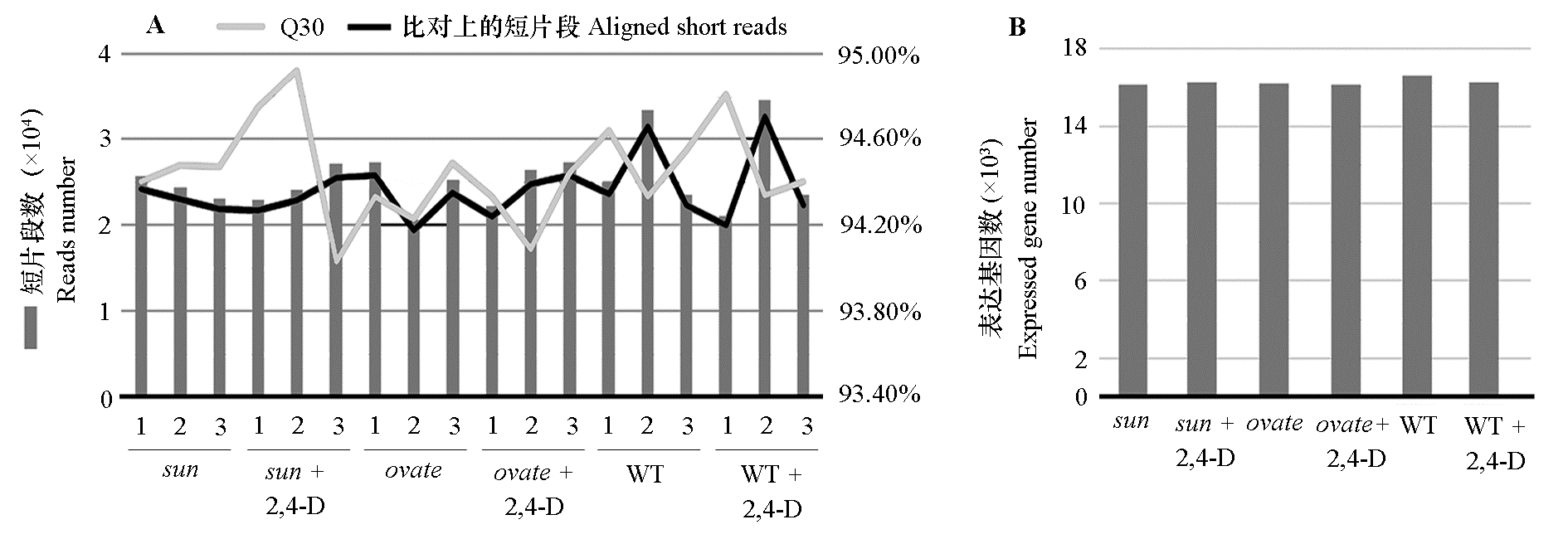
图2 2,4-D处理后各近等基因系花蕾RNA-seq质控(A)和表达基因数(B)
Fig. 2 RNA-seq sequencing quality of the anthesis ovaries after 2,4-D treatment(A)and number of the expressed genes(B)
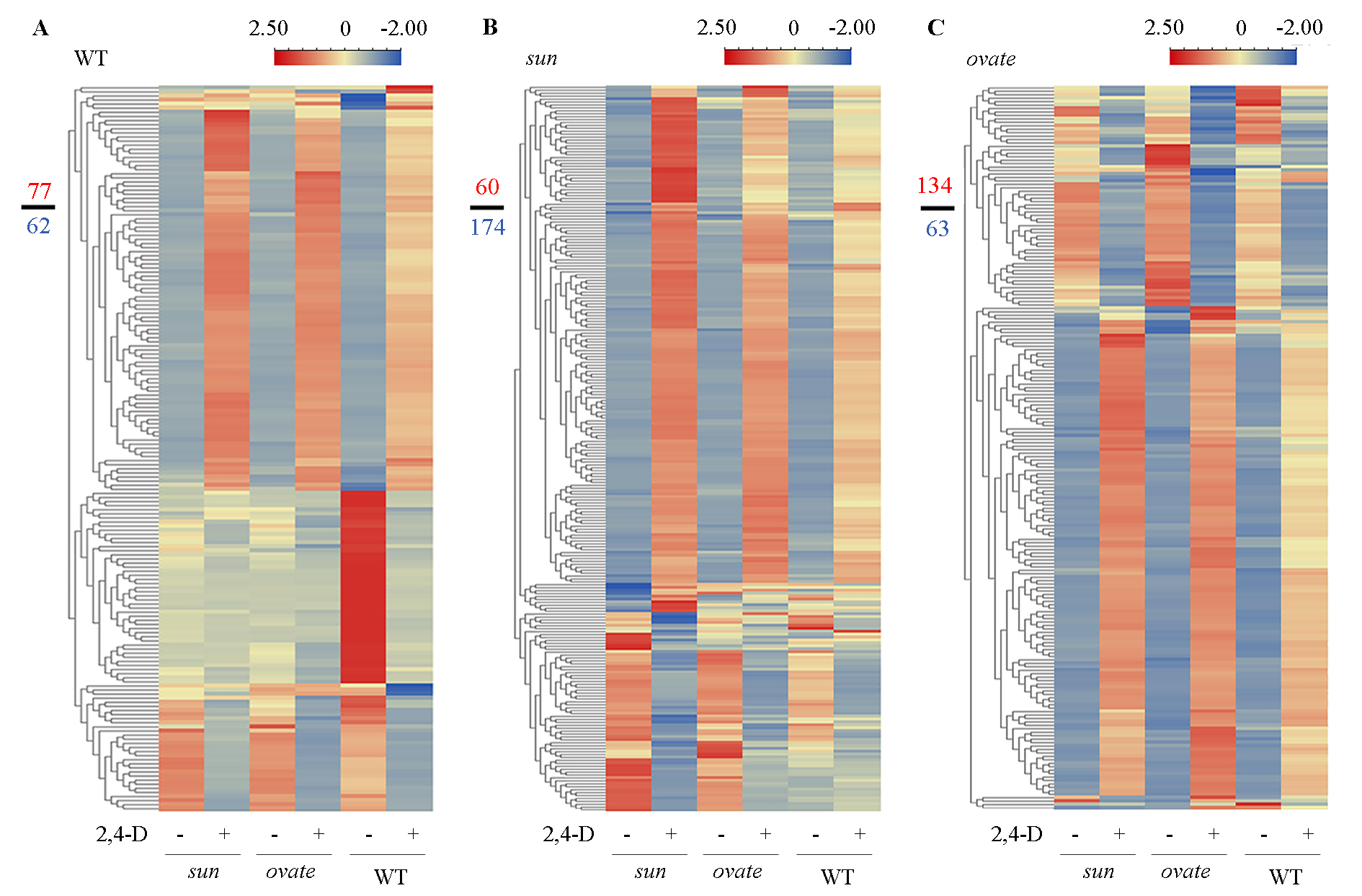
图3 2,4-D处理后WT(A)、sun(B)和ovate(C)近等基因系花蕾中差异表达基因热图 左侧分数中分子与分母分别表示热图中的上调与下调基因数。
Fig. 3 Heatmaps of the differentially expressed genes(DEGs)in the anthesis ovaries of WT(A),sun(B)and ovate(C)NILs after 2,4-D treatment Numerator and denominator in the left represent the number of the up-regulated and down-regulated genes,respectively.
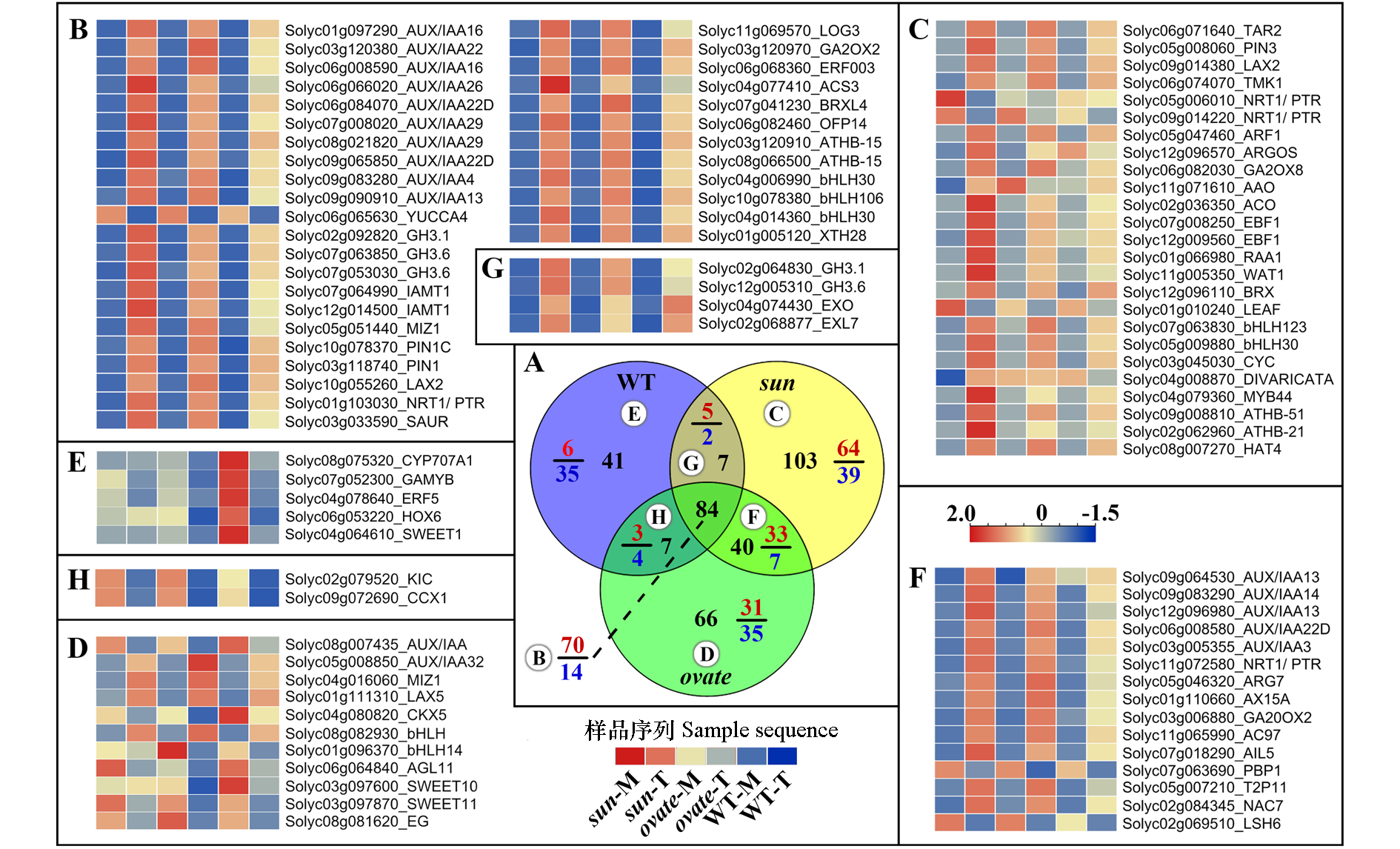
图4 2,4-D处理后各近等基因系花蕾差异表达基因韦恩图分析及感兴趣基因表达热图 A:WT、sun和ovate差异表达基因韦恩图分析,每一部分中黑色数字表示差异表达基因总数,分数的分子与分母分别表示上调与下调基因数,带有圆圈的字母表示该部分基因对应的热图;B:3个近等基因系共有差异表达基因中感兴趣基因热图;C ~ E:分别表示WT、sun和ovate中独有的差异表达基因中感兴趣基因热图;F:表示sun与ovate共有的差异表达基因中感兴趣基因热图;G:表示sun与WT共有的差异表达基因中感兴趣基因热图;H:表示ovate与WT共有的差异表达基因中感兴趣基因热图。
Fig. 4 Venn diagram and heatmap of the interested differentially expressed genes(DEGs)in the anthesis ovaries of WT,sun and ovate NILs after 2,4-D treatment A:Venn diagram of DEGs in WT,sun and ovate NILs after 2,4-D treatment. The black number in each part represents the total number of DEGs,and the numerator and denominator of the fraction represent the number of up-regulated and down-regulated genes,respectively. Letters with circles represent the heatmap corresponding to these genes. B:Heatmap of the interested DEGs shared by the 3 NILs;C-E:heatmaps of the interested genes that only differentially expressed in WT,sun and ovate NILs,respectively. F:Interested DEGs shared by sun and ovate. G:DEGs shared by sun and WT. H:DEGs shared by ovate and WT.

图5 2,4-D处理后各近等基因系花蕾差异表达基因的String分析 A:3个近等基因系共有差异表达基因分析;B:sun 独有的差异表达基因分析;C:ovate独有的差异表达基因分析;D:WT独有的差异表达基因分析;E:sun与ovate共有的差异表达基因分析;F:WT与ovate 共有的差异表达基因分析。
Fig. 5 String analysis of the differentially expressed genes(DEGs)caused by the 2,4-D treatment A:Analysis of DEGs that shared by the three NILs. B:Analysis of DEGs that were only in sun NIL. C:Analysis of DEGs that were only in ovate NIL. D:Analysis of DEGs that were only in WT NIL. E:Analysis of DEGs that were shared by sun and ovate NILs. F:Analysis of DEGs that were shared by WT and ovate NILs.
| [1] |
Azimzadeh J, Nacry P, Christodoulidou A, Drevensek S, Camilleri C, Amiour N, Parcy F, Pastuglia M, Bouchez D. 2008. Arabidopsis TONNEAU1 proteins are essential for preprophase band formation and interact with centrin. Plant Cell, 20 (8):2146-2159.
doi: 10.1105/tpc.107.056812 pmid: 18757558 |
| [2] |
Baima S, Nobili F, Sessa G, Lucchetti S, Ruberti I, Morelli G. 1995. The expression of the Athb-8 homeobox gene is restricted to provascular cells in Arabidopsis thaliana. Development, 121 (12):4171-4182.
doi: 10.1242/dev.121.12.4171 pmid: 8575317 |
| [3] |
Breia R, Conde A, Badim H, Fortes A M, Geros H, Granell A. 2021. Plant SWEETs:from sugar transport to plant-pathogen interaction and more unexpected physiological roles. Plant Physiol, 186 (2):836-852.
doi: 10.1093/plphys/kiab127 URL |
| [4] |
Burstenbinder K, Moller B, Plotner R, Stamm G, Hause G, Mitra D, Abel S. 2017. The IQD family of calmodulin-binding proteins links calcium signaling to microtubules,membrane subdomains,and the nucleus. Plant Physiol, 173 (3):1692-1708.
doi: 10.1104/pp.16.01743 URL |
| [5] |
Burstenbinder K, Savchenko T, Muller J, Adamson A W, Stamm G, Kwong R, Zipp B J, Dinesh D C, Abel S. 2013. Arabidopsis calmodulin-binding protein IQ67-domain 1 localizes to microtubules and interacts with kinesin light chain-related protein-1. Journal of Biological Chemistry, 288 (3):1871-1882.
doi: 10.1074/jbc.M112.396200 URL |
| [6] |
Chen S, Wang X J, Tan G F, Zhou W Q, Wang G L. 2020. Gibberellin and the plant growth retardant Paclobutrazol altered fruit shape and ripening in tomato. Protoplasma, 257 (3):853-861.
doi: 10.1007/s00709-019-01471-2 pmid: 31863170 |
| [7] | Chu Y H, Jang J C, Huang Z, van der Knaap E. 2019. Tomato locule number and fruit size controlled by natural alleles of lc and fas. Plant Direct, 3 (7):e00142. |
| [8] |
Damodharan S, Zhao D, Arazi T. 2016. A common miRNA160-based mechanism regulates ovary patterning,floral organ abscission and lamina outgrowth in tomato. The Plant Journal, 86 (6):458-471.
doi: 10.1111/tpj.13127 pmid: 26800988 |
| [9] |
de Jong M, Wolters-Arts M, Garcia-Martinez J L, Mariani C, Vriezen W H. 2011. The Solanum lycopersicum AUXIN RESPONSE FACTOR 7(SlARF7)mediates cross-talk between auxin and gibberellin signalling during tomato fruit set and development. Journal of Experimental Botany, 62 (2):617-626.
doi: 10.1093/jxb/erq293 URL |
| [10] |
Drevensek S, Goussot M, Duroc Y, Christodoulidou A, Steyaert S, Schaefer E, Duvernois E, Grandjean O, Vantard M, Bouchez D, Pastuglia M. 2012. The Arabidopsis TRM1-TON1 interaction reveals a recruitment network common to plant cortical microtubule arrays and eukaryotic centrosomes. Plant Cell, 24 (1):178-191.
doi: 10.1105/tpc.111.089748 URL |
| [11] | Eyer L, Vain T, Parizkova B, Oklestkova J, Barbez E, Kozubikova H, Pospisil T, Wierzbicka R, Kleine-Vehn J, Franek M, Strnad M, Robert S, Novak O. 2016. 2,4-D and IAA Amino acid conjugates show distinct metabolism in Arabidopsis. PLoS ONE, 11 (7):e0159269. |
| [12] |
Feller A, Machemer K, Braun E L, Grotewold E. 2011. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. The Plant Journal, 66 (1):94-116.
doi: 10.1111/j.1365-313X.2010.04459.x pmid: 21443626 |
| [13] |
Hao Y, Hu G, Breitel D, Liu M, Mila I, Frasse P, Fu Y, Aharoni A, Bouzayen M, Zouine M. 2015. Auxin response factor SlARF 2 is an essential component of the regulatory mechanism controlling fruit ripening in tomato. PLoS Genetics, 11 (12):e1005649.
doi: 10.1371/journal.pgen.1005649 URL |
| [14] |
Hedden P, Thomas S G. 2012. Gibberellin biosynthesis and its regulation. Biochemical Journal, 444 (1):11-25.
doi: 10.1042/BJ20120245 pmid: 22533671 |
| [15] |
Huang Z, van Houten J, Gonzalez G, Xiao H, van der Knaap E. 2013. Genome-wide identification, phylogeny and expression analysis of SUN,OFP and YABBY gene family in tomato. Molecular Genetics Genomics, 288 (3-4):111-129.
doi: 10.1007/s00438-013-0733-0 URL |
| [16] |
Hur Y S, Um J H, Kim S, Kim K, Park H J, Lim J S, Kim W Y, Jun S E, Yoon E K, Lim J, Ohme-Takagi M, Kim D, Park J, Kim G T, Cheon C I. 2015. Arabidopsis thaliana homeobox 12(ATHB12),a homeodomain-leucine zipper protein,regulates leaf growth by promoting cell expansion and endoreduplication. New Phytologist, 205 (1):316-328.
doi: 10.1111/nph.12998 pmid: 25187356 |
| [17] |
Korasick D A, Enders T A, Strader L C. 2013. Auxin biosynthesis and storage forms. Journal of Experimental Botany, 64 (9):2541-55.
doi: 10.1093/jxb/ert080 pmid: 23580748 |
| [18] |
Kurakawa T, Ueda N, Maekawa M, Kobayashi K, Kojima M, Nagato Y, Sakakibara H, Kyozuka J. 2007. Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature, 445 (7128):652-655.
doi: 10.1038/nature05504 |
| [19] |
Lazzaro M D, Wu S, Snouffer A, Wang Y, van der Knaap E. 2018. Plant organ shapes are regulated by protein interactions and associations with microtubules. Frontiers of Plant Science, 9:1766.
doi: 10.3389/fpls.2018.01766 URL |
| [20] | Liu J, van Eck J, Cong B, Tanksley S D. 2002. A new class of regulatory genes underlying the cause of pear-shaped tomato fruit. Proceedings of the National Academy of Science of United States of America, 99 (20):13302-13306. |
| [21] |
Mouchel C F, Briggs G C, Hardtke C S. 2004. Natural genetic variation in Arabidopsis identifies BREVIS RADIX,a novel regulator of cell proliferation and elongation in the root. Genes & Development, 18 (6):700-714.
doi: 10.1101/gad.1187704 URL |
| [22] | Perrot-Rechenmann C. 2010. Cellular responses to auxin:division versus expansion. Cold Spring Harb Perspect Biol, 2 (5):a001446. |
| [23] |
Schmulling T, Werner T, Riefler M, Krupkova E, Bartrina Y, Manns I. 2003. Structure and function of cytokinin oxidase/dehydrogenase genes of maize,rice,Arabidopsis and other species. Journal of Plant Research, 116 (3):241-252.
doi: 10.1007/s10265-003-0096-4 URL |
| [24] |
Snouffer A, Kraus C, van der Knaap E. 2020. The shape of things to come:ovate family proteins regulate plant organ shape. Current Opinion in Plant Biology, 53:98-105.
doi: S1369-5266(19)30096-2 pmid: 31837627 |
| [25] |
Spinner L, Pastuglia M, Belcram K, Pegoraro M, Goussot M, Bouchez D, Schaefer D G. 2010. The function of TONNEAU1 in moss reveals ancient mechanisms of division plane specification and cell elongation in land plants. Development, 137 (16):2733-2742.
doi: 10.1242/dev.043810 pmid: 20663817 |
| [26] |
Sun L, Rodriguez G R, Clevenger J P, Illa-Berenguer E, Lin J, Blakeslee J J, Liu W, Fei Z, Wijeratne A, Meulia T, van der Knaap E. 2015. Candidate gene selection and detailed morphological evaluations of fs8.1,a quantitative trait locus controlling tomato fruit shape. Journal of Experimental Botany, 66 (20):6471-6482.
doi: 10.1093/jxb/erv361 URL |
| [27] |
Vaddepalli P, de Zeeuw T, Strauss S, Burstenbinder K, Liao C Y, Ramalho J J, Smith R S, Weijers D. 2021. Auxin-dependent control of cytoskeleton and cell shape regulates division orientation in the Arabidopsis embryo. Current Biology, 31 (22):4946-4955.
doi: 10.1016/j.cub.2021.09.019 URL |
| [28] | van der Knaap E, Chakrabarti M, Chu Y H, Clevenger J P, Illa-Berenguer E, Huang Z, Keyhaninejad N, Mu Q, Sun L, Wang Y, Wu S. 2014. What lies beyond the eye:the molecular mechanisms regulating tomato fruit weight and shape. Frontiers of Plant Science, 5:227. |
| [29] | van der Knaap E, Østergaard L. 2018. Shaping a fruit:developmental pathways that impact growth patterns. Seminars in Cell & Developmental Biology, 79:27-36. |
| [30] |
Wang Y, Clevenger J P, Illa-Berenguer E, Meulia T, van der Knaap E, Sun L. 2019. A comparison of sun,ovate,fs8.1 and auxin application on tomato fruit shape and gene expression. Plant and Cell Physiology, 60 (5):1067-1081.
doi: 10.1093/pcp/pcz024 URL |
| [31] |
Wu S, Clevenger J P, Sun L, Visa S, Kamiya Y, Jikumaru Y, Blakeslee J, van der Knaap E. 2015. The control of tomato fruit elongation orchestrated by sun,ovate and fs8.1 in a wild relative of tomato. Plant Science, 238:95-104.
doi: 10.1016/j.plantsci.2015.05.019 URL |
| [32] |
Wu S, Xiao H, Cabrera A, Meulia T, van der Knaap E. 2011. SUN regulates vegetative and reproductive organ shape by changing cell division patterns. Plant Physiology, 157 (3):1175-1186.
doi: 10.1104/pp.111.181065 URL |
| [33] |
Wu S, Zhang B, Keyhaninejad N, Rodriguez G R, Kim H J, Chakrabarti M, Illa-Berenguer E, Taitano N K, Gonzalo M J, Diaz A, Pan Y, Leisner C P, Halterman D, Buell C R, Weng Y, Jansky S H, van Eck H, Willemsen J, Monforte A J, Meulia T, van der Knaap E. 2018. A common genetic mechanism underlies morphological diversity in fruits and other plant organs. Nature Communications, 9 (1):4734.
doi: 10.1038/s41467-018-07216-8 pmid: 30413711 |
| [34] |
Xiao H, Jiang N, Schaffner E, Stockinger E J, van der Knaap E. 2008. A retrotransposon-mediated gene duplication underlies morphological variation of tomato fruit. Science, 319 (5869):1527-1530.
doi: 10.1126/science.1153040 pmid: 18339939 |
| [35] |
Xiao H, Radovich C, Welty N, Hsu J, Li D, Meulia T, van der Knaap E. 2009. Integration of tomato reproductive developmental landmarks and expression profiles,and the effect of SUN on fruit shape. BMC Plant Biology, 9:49.
doi: 10.1186/1471-2229-9-49 pmid: 19422692 |
| [36] |
Zhang C, Li X Y, Yan H F, Ullah I, Zuo Z Y, Li L L, Yu J J. 2020. Effects of irrigation quantity and biochar on soil physical properties,growth characteristics,yield and quality of greenhouse tomato. Agricultural Water Management, 241:106263.
doi: 10.1016/j.agwat.2020.106263 URL |
| [37] |
Zhang S L, Gu X, Shao J C, Hu Z F, Yang W C, Wang L P, Su H Y, Zhu L Y. 2021. Auxin metabolism is involved in fruit set and early fruit development in the parthenocarpic tomato“R35-P”. Frontiers in Plant Science, 12:671713.
doi: 10.3389/fpls.2021.671713 URL |
| [38] |
Zhao Q, Wu Y, Gao L, Ma J, Li C Y, Xiang C B. 2014. Sulfur nutrient availability regulates root elongation by affecting root indole-3-acetic acid levels and the stem cell niche. Journal of Integrative Plant Biology, 56 (12):1151-1163.
doi: 10.1111/jipb.12217 |
| [1] | 郑锦荣, 聂俊, 李艳红, 谭德龙, 谢玉明, 张长远. 樱桃番茄新品种‘粤科达101’[J]. 园艺学报, 2023, 50(4): 909-910. |
| [2] | 饶智雄, 安玉艳, 曹荣祥, 唐泉, 汪良驹. 外源ALA缓解ABA抑制草莓根系伸长生长的机理研究[J]. 园艺学报, 2023, 50(3): 461-474. |
| [3] | 刘玉菡, 陶宁, 王庆国, 李清清. 番茄中ABC转运蛋白SlABCG23调控茉莉酸信号途径[J]. 园艺学报, 2023, 50(3): 559-568. |
| [4] | 王泽涵, 于文涛, 王鹏杰, 刘财国, 樊晓静, 谷梦雅, 蔡春平, 王攀, 叶乃兴. 茶树秃房与茸房种质花器官差异表达基因的WGCNA分析[J]. 园艺学报, 2023, 50(3): 620-634. |
| [5] | 蒋彧, 涂勋良, 何俊蓉. 国兰叶色突变体叶片差异表达基因分析[J]. 园艺学报, 2023, 50(2): 371-381. |
| [6] | 于婷婷, 李欢, 宁源生, 宋建飞, 彭璐琳, 贾竣淇, 张玮玮, 杨洪强. 苹果GRAS全基因组鉴定及其对生长素的响应分析[J]. 园艺学报, 2023, 50(2): 397-409. |
| [7] | 郑清波, 鲍泽洋, 蓝青青, 周钰雯, 周雨菲, 郑彩霞, 李旭. 童性与生长素对不定根发生的影响研究进展[J]. 园艺学报, 2023, 50(2): 441-450. |
| [8] | 史洪丽, 李腊, 郭翠梅, 余婷婷, 简伟, 杨星勇. 番茄灰霉病生防菌株TL1的分离、鉴定及其生防能力分析[J]. 园艺学报, 2023, 50(1): 79-90. |
| [9] | 蔺海娇, 梁雨晨, 李玲, 马军, 张璐, 兰振颖, 苑泽宁. 薰衣草CBF途径相关耐寒基因挖掘与调控网络分析[J]. 园艺学报, 2023, 50(1): 131-144. |
| [10] | 忽靖宇, 阙开娟, 缪田丽, 吴少政, 王田田, 张磊, 董鲜, 季鹏章, 董家红. 侵染鸢尾的番茄斑萎病毒鉴定[J]. 园艺学报, 2023, 50(1): 170-176. |
| [11] | 赵雪艳, 王琪, 王莉, 王方圆, 王庆, 李艳. 基于比较转录组的延胡索组织差异性表达分析[J]. 园艺学报, 2023, 50(1): 177-187. |
| [12] | 郑积荣, 王同林, 胡松申. 高品质番茄新品种‘杭杂603’[J]. 园艺学报, 2022, 49(S2): 103-104. |
| [13] | 郑积荣, 王同林. 番茄新品种‘杭杂601’[J]. 园艺学报, 2022, 49(S2): 105-106. |
| [14] | 郑积荣, 王同林. 樱桃番茄新品种‘杭杂503’[J]. 园艺学报, 2022, 49(S2): 107-108. |
| [15] | 黄婷婷, 刘淑芹, 张永志, 李 平, 张志焕, 宋立波. 樱桃番茄新品种‘樱莎红4号’[J]. 园艺学报, 2022, 49(S2): 109-110. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
版权所有 © 2012 《园艺学报》编辑部 京ICP备10030308号-2 国际联网备案号 11010802023439
编辑部地址: 北京市海淀区中关村南大街12号中国农业科学院蔬菜花卉研究所 邮编: 100081
电话: 010-82109523 E-Mail: yuanyixuebao@126.com
技术支持:北京玛格泰克科技发展有限公司