
Acta Horticulturae Sinica ›› 2024, Vol. 51 ›› Issue (10): 2267-2280.doi: 10.16420/j.issn.0513-353x.2023-0723
• Genetic & Breeding·Germplasm Resources·Molecular Biology • Previous Articles Next Articles
ZHANG Shuwen1, LIANG Senmiao1, YU Zheping1, SUN Li1, YE Haiping3, HU Xiaojin4, YAN Liju6, WU Yuyong5, YING Zhengzheng5, ZHENG Xiliang1, QI Xingjiang1,2,*( )
)
Received:2024-05-28
Revised:2024-08-08
Online:2024-10-25
Published:2024-10-21
Contact:
QI Xingjiang
ZHANG Shuwen, LIANG Senmiao, YU Zheping, SUN Li, YE Haiping, HU Xiaojin, YAN Liju, WU Yuyong, YING Zhengzheng, ZHENG Xiliang, QI Xingjiang. Population Structure Analysis of Old Germplasms of Chinese Bayberry Based on WGS[J]. Acta Horticulturae Sinica, 2024, 51(10): 2267-2280.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2023-0723
| 采样地点 Sampling location | 经度Longitude | 纬度 Latitude | 海拔/m Altitude | 品种 Cultivar | 种质编号 Number | 估计树龄/a Tree-age(estimated) |
|---|---|---|---|---|---|---|
| 浙江黄岩 Huangyan,Zhejiang | 121.17° | 28.36° | 30 ~ 60 | 东魁 Dongkui | HY-1,HY-2,HY-4 | 70 |
| HY-5 | 180 | |||||
| HY-9 | 40 | |||||
| 浙江仙居 Xianju,Zhejiang | 120.47° | 28.76° | 150 ~ 170 | 水梅 Shuimei | XJ-1,XJ-3,XJ-5, XJ-7,XJ-9 | 100 |
| 浙江慈溪 Cixi,Zhejiang | 121.15° | 30.56° | 30 | 荸荠种 Biqizhong | CX-1,CX-2,CX-3,CX-4,CX-5 | 120 |
| 湖南靖州 Jingzhou,Hunan | 109.63° | 26.66° | 370 | 木洞大叶 Mudong Daye | HN-1 | 120 |
| HN-2 | 130 | |||||
| 上冲梅 Shangchongmei | HN-3,HN-7 | 110 | ||||
| HN-4 | 120 | |||||
| 湖北来凤 Laifeng,Hubei | 109.29° | 29.61° | 580 ~ 695 | 红梅 Hongmei | HB-1,HB-2 | 150 |
| HB-4,HB-5 | 100 | |||||
| 乌梅 Wumei | HB-6 | 100 | ||||
| 云南腾冲 Tengchong,Yunnan | 98.34° | 25.19° | 1 550 ~ 1 600 | 野生种质 Wild germplasm | YN-1,YN-6 | 190 |
| YN-2 | 130 | |||||
| YN-3,YN-4 | 200 | |||||
| YN-5 | 300 | |||||
| 浙江临海 (原产地福建) Linhai,Zhejiang(Origin is Fujian Province) | 119.28° | 26.08° | 130 | 硬丝 Yingsi | FJ-1 | 30 |
| 软丝 Ruansi | FJ-2 | |||||
| 浮宫一号Fugong 1 | FJ-3 | |||||
| 安海王子Anhai Wangzi | FJ-4 | |||||
| 安海早 Anhaizao | FJ-5 |
Table 1 The information of Chinese bayberry old germplasm resources
| 采样地点 Sampling location | 经度Longitude | 纬度 Latitude | 海拔/m Altitude | 品种 Cultivar | 种质编号 Number | 估计树龄/a Tree-age(estimated) |
|---|---|---|---|---|---|---|
| 浙江黄岩 Huangyan,Zhejiang | 121.17° | 28.36° | 30 ~ 60 | 东魁 Dongkui | HY-1,HY-2,HY-4 | 70 |
| HY-5 | 180 | |||||
| HY-9 | 40 | |||||
| 浙江仙居 Xianju,Zhejiang | 120.47° | 28.76° | 150 ~ 170 | 水梅 Shuimei | XJ-1,XJ-3,XJ-5, XJ-7,XJ-9 | 100 |
| 浙江慈溪 Cixi,Zhejiang | 121.15° | 30.56° | 30 | 荸荠种 Biqizhong | CX-1,CX-2,CX-3,CX-4,CX-5 | 120 |
| 湖南靖州 Jingzhou,Hunan | 109.63° | 26.66° | 370 | 木洞大叶 Mudong Daye | HN-1 | 120 |
| HN-2 | 130 | |||||
| 上冲梅 Shangchongmei | HN-3,HN-7 | 110 | ||||
| HN-4 | 120 | |||||
| 湖北来凤 Laifeng,Hubei | 109.29° | 29.61° | 580 ~ 695 | 红梅 Hongmei | HB-1,HB-2 | 150 |
| HB-4,HB-5 | 100 | |||||
| 乌梅 Wumei | HB-6 | 100 | ||||
| 云南腾冲 Tengchong,Yunnan | 98.34° | 25.19° | 1 550 ~ 1 600 | 野生种质 Wild germplasm | YN-1,YN-6 | 190 |
| YN-2 | 130 | |||||
| YN-3,YN-4 | 200 | |||||
| YN-5 | 300 | |||||
| 浙江临海 (原产地福建) Linhai,Zhejiang(Origin is Fujian Province) | 119.28° | 26.08° | 130 | 硬丝 Yingsi | FJ-1 | 30 |
| 软丝 Ruansi | FJ-2 | |||||
| 浮宫一号Fugong 1 | FJ-3 | |||||
| 安海王子Anhai Wangzi | FJ-4 | |||||
| 安海早 Anhaizao | FJ-5 |
| SNP位置SNP ID | F1 | F2 | R |
|---|---|---|---|
| Chr1:9855573 | TTGTCCACATCAATGCCGCAT | TTGTCCACATCAATGCCGCAC | GTTGGATTAGATACTTATGACATTGCTACG |
| Chr1:24875516 | AGGCACTCAGCCTTGCTGCTC | AGGCACTCAGCCTTGCTGCTG | GCGACTTAACTGTTATTATCACAGC |
| Chr2:1096506 | CTGAGTGTAGATAGAATAGAGCTCC | CTGAGTGTAGATAGAATAGAGCTCT | GGGATATGCACTTACGGATTGC |
| Chr2:34942573 | GCTATGTAAAAATACGTATCGCCAG | GCTATGTAAAAATACGTATCGCCAT | GGCAATCTCTACCAGCATAATG |
| Chr3:3035320 | TACTGCTTTTCTTTTATC | TACTGCTTTTCTTTTATT | TGAGCCCCAAAGATATTGCAAGAACATAAC |
| Chr3:25557841 | TCTAGGATGAGATTTTGTG | TCTAGGATGAGATTTTGTT | TTCAAAGACAAGAATGCAGCATG |
| Chr4:2150997 | GAAGGTGACCAAGTTCATGCTGTACTCAATTTTTGACTT | GAAGGTCGGAGTCAACGGATTGTACTCAATTTTTGACTC | TTCAAAGCCACATCCAACATCA |
| Chr4:30168356 | CTCATCACCAAGAGTCTTCCG | CTCATCACCAAGAGTCTTCCA | CAGTTGACACTGACAGAGCTGG |
| Chr5:6997703 | CAGGATATGATCTAGGAAGTTGTGC | CAGGATATGATCTAGGAAGTTGTGT | TGGCGCAATCACCCCAATTTTAACTGGA |
| Chr5:19892053 | GGTCGATTTGTGGAGGTTTTTCAGT | GGTCGATTTGTGGAGGTTTTTCAGG | AGGCCTCGATGACCACTTTGCCACCCAC |
| Chr6:4338019 | CTCTTTGCAAAGCTCGCTC | CTCTTTGCAAAGCTCGCTT | TTTCTCAATCATAAGAGCGGCCTCTT |
| Chr6:17174448 | TGTTGATGGACCATCCACGGAGTAA | TGTTGATGGACCATCCACGGAGTAC | TTAATTTGTTTTTTATTTATCACC |
| Chr6:20374169 | GTTAAGTCATCCCCTCAAT | GTTAAGTCATCCCCTCAAC | AAGCGACCTCTTCCTCACACAA |
| Chr6:28747343 | TGCTGAAATGAGGCTCAGTGG | TGCTGAAATGAGGCTCAGTGA | AATGATCAGGCTTCATAAGCTC |
| Chr6:32048517 | CCTCCTAGAATTTCAACGCTCTTAA | CCTCCTAGAATTTCAACGCTCTTAT | AACATGACTCGTTTGGTTCAAAT |
| Chr7:7268907 | GCAACAATCTTCCTCGCTTTT | GCAACAATCTTCCTCGCTTTC | GAAAGAGAAGCCCTCCTCAAATTCAGTCAG |
| Chr7:31486783 | CCTCGAAATGTGGAGGTCCCAGC | CCTCGAAATGTGGAGGTCCCAGT | CATGGCGCAATCACCAAATCCCATACTG |
| Chr8:7003270 | CCAACAAATGATAGCGCTTTAG | CCAACAAATGATAGCGCTTTAA | GCTAAAAGTCTTGAGGGTATTA |
| Chr8:18551161 | GGTTGTGAGAGGATTAAAAGTA | GGTTGTGAGAGGATTAAAAGTG | AAATTCCAATTGCATGAGGTTTTCGG |
| Chr8:28476059 | TTCTGCGTTCTCCGACGTTCTTCCC | TTCTGCGTTCTCCGACGTTCTTCCT | TGCATGTGTTATGAACGGATGT |
Table 2 The KASP primer sequence
| SNP位置SNP ID | F1 | F2 | R |
|---|---|---|---|
| Chr1:9855573 | TTGTCCACATCAATGCCGCAT | TTGTCCACATCAATGCCGCAC | GTTGGATTAGATACTTATGACATTGCTACG |
| Chr1:24875516 | AGGCACTCAGCCTTGCTGCTC | AGGCACTCAGCCTTGCTGCTG | GCGACTTAACTGTTATTATCACAGC |
| Chr2:1096506 | CTGAGTGTAGATAGAATAGAGCTCC | CTGAGTGTAGATAGAATAGAGCTCT | GGGATATGCACTTACGGATTGC |
| Chr2:34942573 | GCTATGTAAAAATACGTATCGCCAG | GCTATGTAAAAATACGTATCGCCAT | GGCAATCTCTACCAGCATAATG |
| Chr3:3035320 | TACTGCTTTTCTTTTATC | TACTGCTTTTCTTTTATT | TGAGCCCCAAAGATATTGCAAGAACATAAC |
| Chr3:25557841 | TCTAGGATGAGATTTTGTG | TCTAGGATGAGATTTTGTT | TTCAAAGACAAGAATGCAGCATG |
| Chr4:2150997 | GAAGGTGACCAAGTTCATGCTGTACTCAATTTTTGACTT | GAAGGTCGGAGTCAACGGATTGTACTCAATTTTTGACTC | TTCAAAGCCACATCCAACATCA |
| Chr4:30168356 | CTCATCACCAAGAGTCTTCCG | CTCATCACCAAGAGTCTTCCA | CAGTTGACACTGACAGAGCTGG |
| Chr5:6997703 | CAGGATATGATCTAGGAAGTTGTGC | CAGGATATGATCTAGGAAGTTGTGT | TGGCGCAATCACCCCAATTTTAACTGGA |
| Chr5:19892053 | GGTCGATTTGTGGAGGTTTTTCAGT | GGTCGATTTGTGGAGGTTTTTCAGG | AGGCCTCGATGACCACTTTGCCACCCAC |
| Chr6:4338019 | CTCTTTGCAAAGCTCGCTC | CTCTTTGCAAAGCTCGCTT | TTTCTCAATCATAAGAGCGGCCTCTT |
| Chr6:17174448 | TGTTGATGGACCATCCACGGAGTAA | TGTTGATGGACCATCCACGGAGTAC | TTAATTTGTTTTTTATTTATCACC |
| Chr6:20374169 | GTTAAGTCATCCCCTCAAT | GTTAAGTCATCCCCTCAAC | AAGCGACCTCTTCCTCACACAA |
| Chr6:28747343 | TGCTGAAATGAGGCTCAGTGG | TGCTGAAATGAGGCTCAGTGA | AATGATCAGGCTTCATAAGCTC |
| Chr6:32048517 | CCTCCTAGAATTTCAACGCTCTTAA | CCTCCTAGAATTTCAACGCTCTTAT | AACATGACTCGTTTGGTTCAAAT |
| Chr7:7268907 | GCAACAATCTTCCTCGCTTTT | GCAACAATCTTCCTCGCTTTC | GAAAGAGAAGCCCTCCTCAAATTCAGTCAG |
| Chr7:31486783 | CCTCGAAATGTGGAGGTCCCAGC | CCTCGAAATGTGGAGGTCCCAGT | CATGGCGCAATCACCAAATCCCATACTG |
| Chr8:7003270 | CCAACAAATGATAGCGCTTTAG | CCAACAAATGATAGCGCTTTAA | GCTAAAAGTCTTGAGGGTATTA |
| Chr8:18551161 | GGTTGTGAGAGGATTAAAAGTA | GGTTGTGAGAGGATTAAAAGTG | AAATTCCAATTGCATGAGGTTTTCGG |
| Chr8:28476059 | TTCTGCGTTCTCCGACGTTCTTCCC | TTCTGCGTTCTCCGACGTTCTTCCT | TGCATGTGTTATGAACGGATGT |
| 种质编号 Sample code | 有效数据量/Gb Clean base | GC含量/% GC content | Q20/% | Q30/% | 重复率/% Duplication rate | 测序深度/× Average depth |
|---|---|---|---|---|---|---|
| HY-1 | 10.69 | 38.18 | 99.96 | 97.61 | 35.40 | 36.09 |
| HY-2 | 10.36 | 38.00 | 99.96 | 97.63 | 36.83 | 34.97 |
| HY-4 | 10.45 | 39.43 | 99.96 | 97.72 | 39.03 | 35.27 |
| HY-5 | 12.17 | 39.51 | 99.96 | 97.72 | 35.50 | 41.09 |
| HY-9 | 11.58 | 38.34 | 99.96 | 97.72 | 35.30 | 39.07 |
| XJ-1 | 11.38 | 38.09 | 99.89 | 97.53 | 35.40 | 38.42 |
| XJ-3 | 10.52 | 39.18 | 99.88 | 97.66 | 33.84 | 35.50 |
| XJ-5 | 10.75 | 39.07 | 99.88 | 97.63 | 36.80 | 36.29 |
| XJ-7 | 10.40 | 38.05 | 99.93 | 97.69 | 34.34 | 35.12 |
| XJ-9 | 11.72 | 38.43 | 99.94 | 97.57 | 36.30 | 39.55 |
| CX-1 | 10.32 | 38.16 | 99.92 | 97.52 | 29.90 | 34.84 |
| CX-2 | 10.99 | 38.08 | 99.93 | 97.66 | 36.70 | 37.11 |
| CX-3 | 11.11 | 38.24 | 99.93 | 97.51 | 35.91 | 37.51 |
| CX-4 | 10.60 | 38.00 | 99.95 | 97.58 | 34.70 | 35.77 |
| CX-5 | 10.24 | 39.14 | 99.95 | 97.70 | 30.24 | 34.57 |
| HN-1 | 12.86 | 39.00 | 99.97 | 97.62 | 37.40 | 43.40 |
| HN-2 | 11.12 | 39.13 | 99.96 | 97.64 | 34.70 | 37.54 |
| HN-3 | 11.20 | 39.11 | 99.96 | 97.62 | 38.10 | 37.81 |
| HN-4 | 10.06 | 38.00 | 99.96 | 97.66 | 35.30 | 33.96 |
| HN-7 | 10.90 | 39.24 | 99.96 | 97.62 | 36.27 | 36.80 |
| HB-1 | 10.22 | 40.04 | 99.93 | 97.68 | 35.20 | 34.48 |
| HB-2 | 11.88 | 39.00 | 99.94 | 97.56 | 38.44 | 40.11 |
| HB-4 | 11.14 | 40.30 | 99.94 | 97.64 | 36.80 | 37.62 |
| HB-5 | 10.75 | 38.00 | 99.96 | 97.79 | 36.24 | 36.30 |
| HB-6 | 11.42 | 38.27 | 99.96 | 97.59 | 35.15 | 38.54 |
| FJ-1 | 10.01 | 38.14 | 99.99 | 97.00 | 27.05 | 33.80 |
| FJ-2 | 16.08 | 38.42 | 99.99 | 97.62 | 27.70 | 54.28 |
| FJ-3 | 10.35 | 38.00 | 99.99 | 96.94 | 25.20 | 34.93 |
| FJ-4 | 11.59 | 38.28 | 99.99 | 97.06 | 25.33 | 39.13 |
| FJ-5 | 11.75 | 38.12 | 99.99 | 97.63 | 25.42 | 39.68 |
| YN-1 | 10.63 | 38.51 | 99.93 | 97.72 | 30.77 | 35.88 |
| YN-2 | 10.81 | 38.34 | 99.93 | 97.44 | 31.30 | 36.51 |
| YN-3 | 10.78 | 38.07 | 99.93 | 97.46 | 31.04 | 36.40 |
| YN-4 | 10.54 | 38.16 | 99.93 | 97.48 | 29.22 | 35.57 |
| YN-5 | 10.50 | 38.31 | 99.93 | 97.36 | 30.62 | 35.46 |
| YN-6 | 10.37 | 38.16 | 99.93 | 97.49 | 28.15 | 35.01 |
| 平均Average | 11.06 | 38.57 | 99.95 | 97.56 | 33.38 | 37.34 |
Table 3 Sample re-sequencing information
| 种质编号 Sample code | 有效数据量/Gb Clean base | GC含量/% GC content | Q20/% | Q30/% | 重复率/% Duplication rate | 测序深度/× Average depth |
|---|---|---|---|---|---|---|
| HY-1 | 10.69 | 38.18 | 99.96 | 97.61 | 35.40 | 36.09 |
| HY-2 | 10.36 | 38.00 | 99.96 | 97.63 | 36.83 | 34.97 |
| HY-4 | 10.45 | 39.43 | 99.96 | 97.72 | 39.03 | 35.27 |
| HY-5 | 12.17 | 39.51 | 99.96 | 97.72 | 35.50 | 41.09 |
| HY-9 | 11.58 | 38.34 | 99.96 | 97.72 | 35.30 | 39.07 |
| XJ-1 | 11.38 | 38.09 | 99.89 | 97.53 | 35.40 | 38.42 |
| XJ-3 | 10.52 | 39.18 | 99.88 | 97.66 | 33.84 | 35.50 |
| XJ-5 | 10.75 | 39.07 | 99.88 | 97.63 | 36.80 | 36.29 |
| XJ-7 | 10.40 | 38.05 | 99.93 | 97.69 | 34.34 | 35.12 |
| XJ-9 | 11.72 | 38.43 | 99.94 | 97.57 | 36.30 | 39.55 |
| CX-1 | 10.32 | 38.16 | 99.92 | 97.52 | 29.90 | 34.84 |
| CX-2 | 10.99 | 38.08 | 99.93 | 97.66 | 36.70 | 37.11 |
| CX-3 | 11.11 | 38.24 | 99.93 | 97.51 | 35.91 | 37.51 |
| CX-4 | 10.60 | 38.00 | 99.95 | 97.58 | 34.70 | 35.77 |
| CX-5 | 10.24 | 39.14 | 99.95 | 97.70 | 30.24 | 34.57 |
| HN-1 | 12.86 | 39.00 | 99.97 | 97.62 | 37.40 | 43.40 |
| HN-2 | 11.12 | 39.13 | 99.96 | 97.64 | 34.70 | 37.54 |
| HN-3 | 11.20 | 39.11 | 99.96 | 97.62 | 38.10 | 37.81 |
| HN-4 | 10.06 | 38.00 | 99.96 | 97.66 | 35.30 | 33.96 |
| HN-7 | 10.90 | 39.24 | 99.96 | 97.62 | 36.27 | 36.80 |
| HB-1 | 10.22 | 40.04 | 99.93 | 97.68 | 35.20 | 34.48 |
| HB-2 | 11.88 | 39.00 | 99.94 | 97.56 | 38.44 | 40.11 |
| HB-4 | 11.14 | 40.30 | 99.94 | 97.64 | 36.80 | 37.62 |
| HB-5 | 10.75 | 38.00 | 99.96 | 97.79 | 36.24 | 36.30 |
| HB-6 | 11.42 | 38.27 | 99.96 | 97.59 | 35.15 | 38.54 |
| FJ-1 | 10.01 | 38.14 | 99.99 | 97.00 | 27.05 | 33.80 |
| FJ-2 | 16.08 | 38.42 | 99.99 | 97.62 | 27.70 | 54.28 |
| FJ-3 | 10.35 | 38.00 | 99.99 | 96.94 | 25.20 | 34.93 |
| FJ-4 | 11.59 | 38.28 | 99.99 | 97.06 | 25.33 | 39.13 |
| FJ-5 | 11.75 | 38.12 | 99.99 | 97.63 | 25.42 | 39.68 |
| YN-1 | 10.63 | 38.51 | 99.93 | 97.72 | 30.77 | 35.88 |
| YN-2 | 10.81 | 38.34 | 99.93 | 97.44 | 31.30 | 36.51 |
| YN-3 | 10.78 | 38.07 | 99.93 | 97.46 | 31.04 | 36.40 |
| YN-4 | 10.54 | 38.16 | 99.93 | 97.48 | 29.22 | 35.57 |
| YN-5 | 10.50 | 38.31 | 99.93 | 97.36 | 30.62 | 35.46 |
| YN-6 | 10.37 | 38.16 | 99.93 | 97.49 | 28.15 | 35.01 |
| 平均Average | 11.06 | 38.57 | 99.95 | 97.56 | 33.38 | 37.34 |
| 样本 Sample | SNP数量 Number of SNPs | 转换位点 Number of transition | 颠换位点 Number of transversion | 转换/颠换 Transition/ transversion | 杂合变异位点 Number of heterozygosity | 纯合变异位点 Number of homozygosity | 杂合率/% Heterozygosity rate |
|---|---|---|---|---|---|---|---|
| HY-1 | 2 050 938 | 1 411 884 | 639 054 | 2.21 | 1 724 369 | 326 569 | 84.08 |
| HY-2 | 2 048 642 | 1 410 307 | 638 335 | 2.21 | 1 721 860 | 326 782 | 84.05 |
| HY-4 | 2 052 202 | 1 412 718 | 639 484 | 2.21 | 1 724 844 | 327 358 | 84.05 |
| HY-5 | 2 052 967 | 1 413 221 | 639 746 | 2.21 | 1 723 585 | 329 382 | 83.96 |
| HY-9 | 2 052 974 | 1 413 453 | 639 521 | 2.21 | 1 725 777 | 327 197 | 84.06 |
| XJ-1 | 2 020 543 | 1 392 832 | 627 711 | 2.22 | 1 794 440 | 226 103 | 88.81 |
| XJ-3 | 2 015 957 | 1 389 488 | 626 469 | 2.22 | 1 788 704 | 227 253 | 88.73 |
| XJ-5 | 2 014 444 | 1 388 425 | 626 019 | 2.22 | 1 785 481 | 228 963 | 88.63 |
| XJ-7 | 2 019 041 | 1 391 447 | 627 594 | 2.22 | 1 792 393 | 226 648 | 88.77 |
| XJ-9 | 2 053 334 | 1 413 369 | 639 965 | 2.21 | 1 726 162 | 327 172 | 84.07 |
| CX-1 | 1 726 871 | 1 187 340 | 539 531 | 2.20 | 1 563 406 | 163 465 | 90.53 |
| CX-2 | 2 355 419 | 1 621 539 | 733 880 | 2.21 | 2 094 979 | 260 440 | 88.94 |
| CX-3 | 1 726 465 | 1 187 243 | 539 222 | 2.20 | 1 562 720 | 163 745 | 90.52 |
| CX-4 | 1 730 185 | 1 189 641 | 540 544 | 2.20 | 1 565 862 | 164 323 | 90.50 |
| CX-5 | 1 726 906 | 1 187 304 | 539 602 | 2.20 | 1 561 793 | 165 113 | 90.44 |
| HN-1 | 2 001 119 | 1 380 042 | 621 077 | 2.22 | 1 513 540 | 487 579 | 75.63 |
| HN-2 | 2 159 194 | 1 494 361 | 664 833 | 2.25 | 1 779 964 | 379 230 | 82.44 |
| HN-3 | 2 063 061 | 1 423 165 | 639 896 | 2.22 | 1 570 871 | 492 190 | 76.14 |
| HN-4 | 2 063 833 | 1 423 804 | 640 029 | 2.22 | 1 574 460 | 489 373 | 76.29 |
| HN-7 | 2 062 759 | 1 422 958 | 639 801 | 2.22 | 1 571 669 | 491 090 | 76.19 |
| HB-1 | 2 023 343 | 1 399 019 | 624 324 | 2.24 | 1 519 121 | 504 222 | 75.08 |
| HB-2 | 2 022 311 | 1 394 607 | 627 704 | 2.22 | 1 476 575 | 545 736 | 73.01 |
| HB-4 | 2 014 613 | 1 389 561 | 625 052 | 2.22 | 1 466 369 | 548 244 | 72.79 |
| HB-5 | 2 058 455 | 1 421 689 | 636 766 | 2.23 | 1 566 655 | 491 800 | 76.11 |
| HB-6 | 1 893 434 | 1 305 944 | 587 490 | 2.22 | 1 222 637 | 670 797 | 64.57 |
| FJ-1 | 2 049 201 | 1 417 215 | 631 986 | 2.24 | 1 636 785 | 412 416 | 79.87 |
| FJ-2 | 2 135 600 | 1 477 909 | 657 691 | 2.25 | 1 751 915 | 383 685 | 82.03 |
| FJ-3 | 2 059 344 | 1 422 225 | 637 119 | 2.23 | 1 645 218 | 414 126 | 79.89 |
| FJ-4 | 2 144 618 | 1 486 049 | 658 569 | 2.26 | 1 738 952 | 405 666 | 81.08 |
| FJ-5 | 2 109 931 | 1 460 493 | 649 438 | 2.25 | 1 725 461 | 384 470 | 81.78 |
| YN-1 | 3 517 942 | 2 430 232 | 1 087 710 | 2.23 | 966 536 | 2 551 406 | 27.47 |
| YN-2 | 3 508 535 | 2 422 505 | 1 086 030 | 2.23 | 913 854 | 2 594 681 | 26.05 |
| YN-3 | 3 512 213 | 2 426 292 | 1 085 921 | 2.23 | 952 693 | 2 559 520 | 27.13 |
| YN-4 | 3 505 810 | 2 421 699 | 1 084 111 | 2.23 | 943 055 | 2 562 755 | 26.90 |
| YN-5 | 3 499 575 | 2 416 870 | 1 082 705 | 2.23 | 884 303 | 2 615 272 | 25.27 |
| YN-6 | 3 504 092 | 2 418 865 | 1 085 227 | 2.23 | 857 631 | 2 646 461 | 24.48 |
Table 4 The SNP information detected in the 36 samples
| 样本 Sample | SNP数量 Number of SNPs | 转换位点 Number of transition | 颠换位点 Number of transversion | 转换/颠换 Transition/ transversion | 杂合变异位点 Number of heterozygosity | 纯合变异位点 Number of homozygosity | 杂合率/% Heterozygosity rate |
|---|---|---|---|---|---|---|---|
| HY-1 | 2 050 938 | 1 411 884 | 639 054 | 2.21 | 1 724 369 | 326 569 | 84.08 |
| HY-2 | 2 048 642 | 1 410 307 | 638 335 | 2.21 | 1 721 860 | 326 782 | 84.05 |
| HY-4 | 2 052 202 | 1 412 718 | 639 484 | 2.21 | 1 724 844 | 327 358 | 84.05 |
| HY-5 | 2 052 967 | 1 413 221 | 639 746 | 2.21 | 1 723 585 | 329 382 | 83.96 |
| HY-9 | 2 052 974 | 1 413 453 | 639 521 | 2.21 | 1 725 777 | 327 197 | 84.06 |
| XJ-1 | 2 020 543 | 1 392 832 | 627 711 | 2.22 | 1 794 440 | 226 103 | 88.81 |
| XJ-3 | 2 015 957 | 1 389 488 | 626 469 | 2.22 | 1 788 704 | 227 253 | 88.73 |
| XJ-5 | 2 014 444 | 1 388 425 | 626 019 | 2.22 | 1 785 481 | 228 963 | 88.63 |
| XJ-7 | 2 019 041 | 1 391 447 | 627 594 | 2.22 | 1 792 393 | 226 648 | 88.77 |
| XJ-9 | 2 053 334 | 1 413 369 | 639 965 | 2.21 | 1 726 162 | 327 172 | 84.07 |
| CX-1 | 1 726 871 | 1 187 340 | 539 531 | 2.20 | 1 563 406 | 163 465 | 90.53 |
| CX-2 | 2 355 419 | 1 621 539 | 733 880 | 2.21 | 2 094 979 | 260 440 | 88.94 |
| CX-3 | 1 726 465 | 1 187 243 | 539 222 | 2.20 | 1 562 720 | 163 745 | 90.52 |
| CX-4 | 1 730 185 | 1 189 641 | 540 544 | 2.20 | 1 565 862 | 164 323 | 90.50 |
| CX-5 | 1 726 906 | 1 187 304 | 539 602 | 2.20 | 1 561 793 | 165 113 | 90.44 |
| HN-1 | 2 001 119 | 1 380 042 | 621 077 | 2.22 | 1 513 540 | 487 579 | 75.63 |
| HN-2 | 2 159 194 | 1 494 361 | 664 833 | 2.25 | 1 779 964 | 379 230 | 82.44 |
| HN-3 | 2 063 061 | 1 423 165 | 639 896 | 2.22 | 1 570 871 | 492 190 | 76.14 |
| HN-4 | 2 063 833 | 1 423 804 | 640 029 | 2.22 | 1 574 460 | 489 373 | 76.29 |
| HN-7 | 2 062 759 | 1 422 958 | 639 801 | 2.22 | 1 571 669 | 491 090 | 76.19 |
| HB-1 | 2 023 343 | 1 399 019 | 624 324 | 2.24 | 1 519 121 | 504 222 | 75.08 |
| HB-2 | 2 022 311 | 1 394 607 | 627 704 | 2.22 | 1 476 575 | 545 736 | 73.01 |
| HB-4 | 2 014 613 | 1 389 561 | 625 052 | 2.22 | 1 466 369 | 548 244 | 72.79 |
| HB-5 | 2 058 455 | 1 421 689 | 636 766 | 2.23 | 1 566 655 | 491 800 | 76.11 |
| HB-6 | 1 893 434 | 1 305 944 | 587 490 | 2.22 | 1 222 637 | 670 797 | 64.57 |
| FJ-1 | 2 049 201 | 1 417 215 | 631 986 | 2.24 | 1 636 785 | 412 416 | 79.87 |
| FJ-2 | 2 135 600 | 1 477 909 | 657 691 | 2.25 | 1 751 915 | 383 685 | 82.03 |
| FJ-3 | 2 059 344 | 1 422 225 | 637 119 | 2.23 | 1 645 218 | 414 126 | 79.89 |
| FJ-4 | 2 144 618 | 1 486 049 | 658 569 | 2.26 | 1 738 952 | 405 666 | 81.08 |
| FJ-5 | 2 109 931 | 1 460 493 | 649 438 | 2.25 | 1 725 461 | 384 470 | 81.78 |
| YN-1 | 3 517 942 | 2 430 232 | 1 087 710 | 2.23 | 966 536 | 2 551 406 | 27.47 |
| YN-2 | 3 508 535 | 2 422 505 | 1 086 030 | 2.23 | 913 854 | 2 594 681 | 26.05 |
| YN-3 | 3 512 213 | 2 426 292 | 1 085 921 | 2.23 | 952 693 | 2 559 520 | 27.13 |
| YN-4 | 3 505 810 | 2 421 699 | 1 084 111 | 2.23 | 943 055 | 2 562 755 | 26.90 |
| YN-5 | 3 499 575 | 2 416 870 | 1 082 705 | 2.23 | 884 303 | 2 615 272 | 25.27 |
| YN-6 | 3 504 092 | 2 418 865 | 1 085 227 | 2.23 | 857 631 | 2 646 461 | 24.48 |
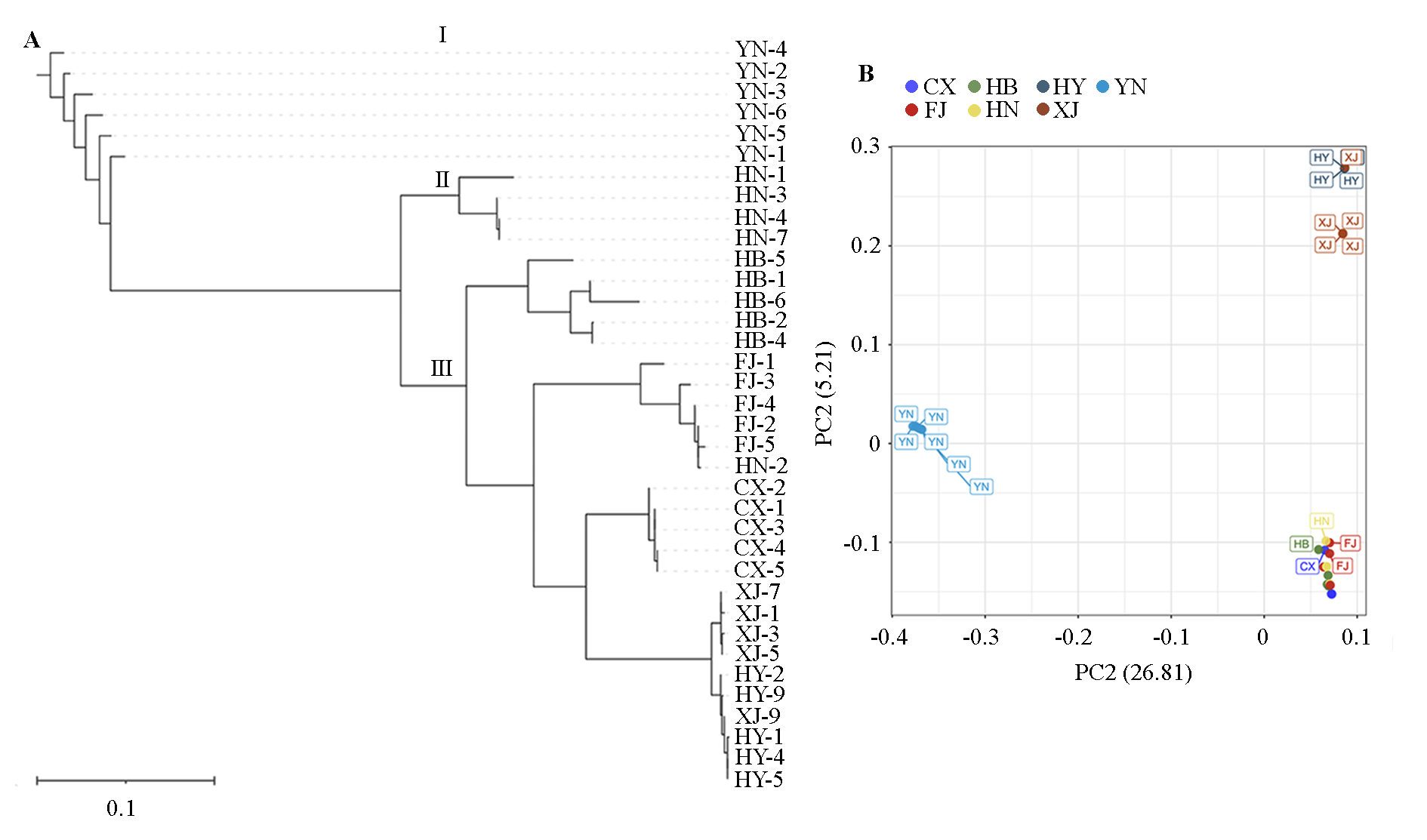
Fig. 2 Population evolution tree(A)and PCA analysis(B)of old germplasms of Chinese bayberry The information of 36 germplasms was detailed in Table 1. FJ:Fujian;CX:Cixi,Zhejiang;HB:Laifeng,Hubei;HN:Jingzhou,Hunan;XJ:Xianju,Zhejiang;HY:Huangyan,Zhejiang;YN:Tengchong,Yunnan.
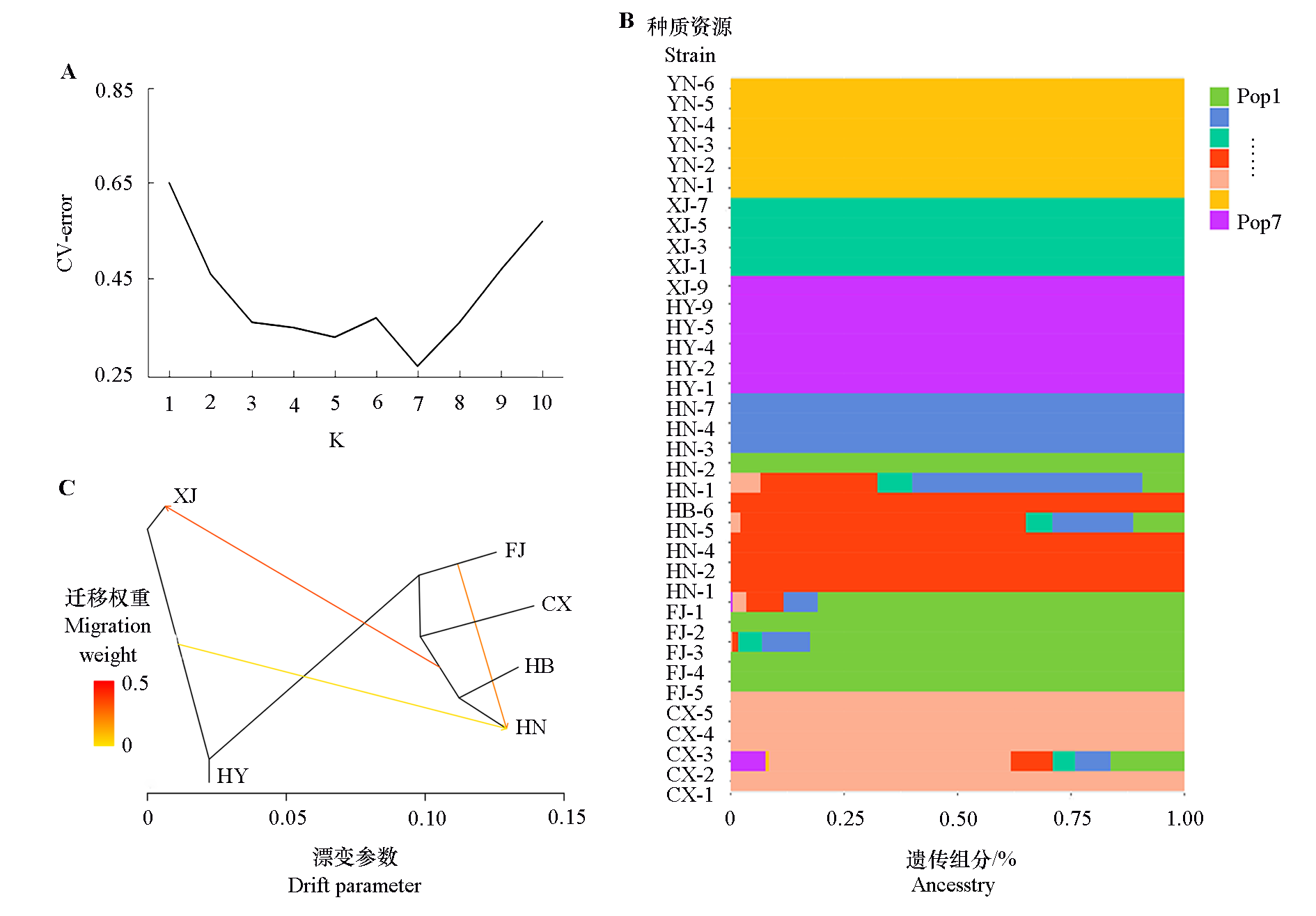
Fig. 3 Cross validation(CV)error rate of K(A),population structure relationship(B)and signals of introgression among different populations(C) The information of 36 germplasms was detailed in Table 1. FJ:Fujian;CX:Cixi,Zhejiang;HB:Laifeng,Hubei;HN:Jingzhou,Hunan;XJ:Xianju,Zhejiang;HY:Huangyan,Zhejiang;YN:Tengchong,Yunnan.

Fig. 4 LD attenuation(A)and Fst analysis(B)between Huangyan and Cixi old germplasm population of Zhejiang The information of 36 germplasms was detailed in Table 1. FJ:Fujian;CX:Cixi,Zhejiang;HB:Laifeng,Hubei;HN:Jingzhou,Hunan;XJ:Xianju,Zhejiang;HY:Huangyan,Zhejiang;YN:Tengchong,Yunnan.
| [1] |
|
| [2] |
|
| [3] |
|
| [4] |
|
| [5] |
|
| [6] |
doi: 10.1093/bioinformatics/btr330 pmid: 21653522 |
| [7] |
|
| [8] |
|
| [9] |
|
| [10] |
|
| [11] |
doi: 10.16420/j.issn.0513-353x.2022-0521 |
|
侯炳豪, 高婷, 魏月德, 高水练, 蔡银笔, 陈志明, 金珊, 张兴坦, 王鹏杰, 叶乃兴. 2023. 基于高深度基因组重测序的‘铁观音’茶树无性繁殖后代遗传变异研究. 园艺学报, 50 (7):1505-1517.
|
|
| [12] |
|
| [13] |
|
| [14] |
|
| [15] |
|
| [16] |
|
| [17] |
|
| [18] |
doi: 10.1093/bioinformatics/btp352 pmid: 19505943 |
| [19] |
|
| [20] |
|
| [21] |
|
| [22] |
|
| [23] |
|
| [24] |
|
| [25] |
|
| [26] |
|
| [27] |
|
| [28] |
|
| [29] |
|
| [30] |
|
|
张瑞, 张夏燚, 赵婷, 王双成, 张仲兴, 刘博, 张德, 王延秀. 2022. 基于转录组分析垂丝海棠响应盐碱胁迫的分子机制. 园艺学报, 49 (2):237-251.
doi: 10.16420/j.issn.0513-353x.2021-0077 |
|
| [31] |
|
| [32] |
|
| [33] |
|
| [34] |
doi: 10.16420/j.issn.0513-353x.2018-0233 |
|
张淑文, 梁森苗, 郑锡良, 任海英, 朱婷婷, 戚行江. 2019. 杨梅基因组SSR引物的开发与应用. 园艺学报, 46 (1):149-156.
doi: 10.16420/j.issn.0513-353x.2018-0233 |
|
| [35] |
|
|
张淑文, 俞浙萍, 孙鹂, 武祥琪, 梁森苗, 郑锡良, 戚行江. 2022. 基于重测序的杨梅InDel标记开发与果实性状关联分析. 分子植物育种, 20 (6):1890-1900.
|
| [1] | QIN Yu, GUO Rongkun, NONG Huilan, WANG Yan, CUI Kai, DONG Ningguang. Using the Fluorescent Labeled SSR Markers to Establish Molecular Identity of Hawthorn [J]. Acta Horticulturae Sinica, 2024, 51(6): 1227-1240. |
| [2] | ZHANG Wenjing, XU Dayong, WU Qianlin, YANG Fo, XIN Bingyue, ZENG Xin, LI Feng. Genome Analysis of Bacillus velezensis XDY66,an Antagonist of Tomato Botrytis cinerea [J]. Acta Horticulturae Sinica, 2024, 51(6): 1413-1425. |
| [3] | WU Xiangqi, SUN Li, YU Zheping, YU Qinpei, LIANG Senmiao, ZHENG Xiliang, QI Xingjiang, ZHANG Shuwen. Functional Study of MrSPL4 Gene in Response to Drought and Low Temperature Stress in Chinese Bayberry [J]. Acta Horticulturae Sinica, 2024, 51(5): 927-938. |
| [4] | YUAN Na, XU Qinyuan, XU Zhaolong, ZHOU Ling, LIU Xiaoqing, CHEN Xin, DU Jianchang. The Development and Verification of SNP Lquid Chips for Common Bean Based on Targeted Sequencing Technology [J]. Acta Horticulturae Sinica, 2024, 51(5): 1017-1032. |
| [5] | JIANG Shuang, WANG Xiaoqing, SHI Chunhui, LI Shuigen, LUO Jun. Detection of SLAF Tag Polymorphism Associated with Pear Russet Peel by PCR Product Pool Resequencing [J]. Acta Horticulturae Sinica, 2024, 51(4): 707-714. |
| [6] | YUAN Qingyun, HAN Yu, HE Wei, SU Hui, BAN Qiuyan, WU Chunlai, ZHOU Qiongqiong, XU Wenjing, WANG Liyuan, ZHANG Fen. Cloning and ABA Transport Function Analysis of CsNPF6.1/6.3 in Tea Plants [J]. Acta Horticulturae Sinica, 2024, 51(12): 2817-2828. |
| [7] | YAO Yuan, DENG Lijun, HU Juan, TANG Xiaoyu, WANG Tie, LI Hang, SUN Guochao, XIONG Bo, LIAO Ling, WANG Zhihui. Whole Genome Resequencing Analysis of‘Cuihongli’Plum and Its Early-Ripening Bud Sport Mutation [J]. Acta Horticulturae Sinica, 2024, 51(10): 2255-2266. |
| [8] | NIE Xinghua, ZHANG Yu, LIU Song, YANG Jiabin, HAO Yaqiong, LIU Yang, QIN Ling, XING Yu. Study on Genetic Characteristics and Taxonomic Status of Wild Chinese Chestnut Based on Genome Re-sequencing [J]. Acta Horticulturae Sinica, 2023, 50(8): 1622-1636. |
| [9] | WANG Rui, HONG Wenjuan, LUO Hua, ZHAO Lina, CHEN Ying, WANG Jun. Construction of SSR Fingerprints of Pomegranate Cultivars and Male Parent Identification of Hybrids [J]. Acta Horticulturae Sinica, 2023, 50(2): 265-278. |
| [10] | LIU Chenglang, FENG Di, CAO Zonghong, YAN Suyun, ZHANG Yongqing, XU Xiangzeng, GAO Shide, YE Junli, CHAI Lijun, XIE Zongzhou, DENG Xiuxin. Comparative Analysis of Genetic Background of‘Dongshi Zaoyou’,‘Shatianyou’and‘Shuijingyou’Pummelos [J]. Acta Horticulturae Sinica, 2023, 50(11): 2337-2349. |
| [11] | XIE Qian, ZHANG Shiyan, JIANG Lai, DING Mingyue, LIU Lingling, WU Rujian, CHEN Qingxi. Analysis of Canarium album Transcriptome SSR Information and Molecular Marker Development and Application [J]. Acta Horticulturae Sinica, 2023, 50(11): 2350-2364. |
| [12] | CHEN Meichun, DENG Yingjie, CHEN Zheng, ZHU Yujing, LI Chunniu, WANG Jieping, LIU Bo. Morphological Characteristics and Volatile Component Diversity of the Jasminum Germplasm Resources [J]. Acta Horticulturae Sinica, 2023, 50(11): 2435-2452. |
| [13] | WU Guijin, CHEN Longqing, WEI Qiuyu, HU Yunchong, XU Jian, LI Dongxue, CHEN Shengtong, WANG Zhonglang, GENG Fang. Statistics and Phenotypic Traits Analysis of Camellia reticulata Registered Cultivars [J]. Acta Horticulturae Sinica, 2023, 50(10): 2157-2170. |
| [14] | YU Zheping, YAN Xiujuan, ZHANG Shuwen, ZHENG Xiliang, SUN Li, REN Haiying, LIANG Senmiao, QI Xingjiang. Preliminary Study on the Mechanism of Improvement of Resistance to Chinese Bayberry Twig Blight Disease by Brassinolide and Oligosaccharin [J]. Acta Horticulturae Sinica, 2023, 50(10): 2229-2241. |
| [15] | LUO Hailin, YUAN Lei, WENG Hua, YAN Jiahui, GUO Qingyun, WANG Wenqing, MA Xinming. Identification and Analysis of Complete Genomic Sequence of Broad Bean Wilt Virus 2 Pepper Isolate in Qinghai Province [J]. Acta Horticulturae Sinica, 2023, 50(1): 161-169. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd