
Acta Horticulturae Sinica ›› 2022, Vol. 49 ›› Issue (1): 100-116.doi: 10.16420/j.issn.0513-353x.2020-0850
• Research Papers • Previous Articles Next Articles
XIE Siyi1,3, ZHOU Chengzhe1,2,3, ZHU Chen1,2,3, ZHAN Dongmei1,3, CHEN Lan1,3, WU Zuchun1,3, LAI Zhongxiong1,2, GUO Yuqiong1,3,*( )
)
Received:2021-04-23
Revised:2021-07-26
Online:2022-01-25
Published:2022-01-24
Contact:
GUO Yuqiong
E-mail:guoyq828@163.com
CLC Number:
XIE Siyi, ZHOU Chengzhe, ZHU Chen, ZHAN Dongmei, CHEN Lan, WU Zuchun, LAI Zhongxiong, GUO Yuqiong. Genome-wide Identification and Expression Analysis of CsTIFY Transcription Factor Family Under Abiotic Stress and Hormone Treatments in Camellia sinensis[J]. Acta Horticulturae Sinica, 2022, 49(1): 100-116.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2020-0850
| 用途 Purpose | 基因 Gene | 上游引物序列(5′-3′) Forward primer | 下游引物序列(5′-3′) Reverse primer |
|---|---|---|---|
| qRT-PCR | CsTIFY1 | TGAGTAACCGATTCGCTGG | GCTTGTGGAAACCCATCTCTG |
| CsTIFY2 | TCAACGAGGAAGAGGCTATGG | GGGTCCTCATCCTCAACCATT | |
| CsTIFY3 | GCCTCTGTTGGTGGTTCTTC | CGGTTATTCATCTCAGTCCCT | |
| CsTIFY4 | TCTCCGCCACTTCTGATCTG | GCGAATCGGTTACTCATCTCAG | |
| CsPPD1 | CCTACCAGATGTGCGAGAATG | TCGCCTTCATCCAAGCTGAT | |
| CsPPD2 | GCAAATTCTCCATTGCTCGC | TCCACGACGGTCTTCTCATC | |
| CsJAZ1 | TCCTCCAAACCTCTTCCCAAT | CACAGAGCCAGAAGCAATCAC | |
| CsJAZ2 | TCATTGATCGCCGAAGAAGC | GAATAGAGAGGCAAAGCAGG | |
| CsJAZ4 | TGACCCACTTGCTGCTTCT | GAACGCCACCTTGTCCAT | |
| CsJAZ5 | TGTCGAGTTCGTCTGGTTCT | CCGAATGTGCCTTTCTTCTC | |
| CsJAZ6 | CGGTGGCTGTCAAAGAAGGTATT | CTCTAATCGCCATTTCAGTCCCT | |
| CsJAZ7 | TCATTGATCGCCGAAGAAGC | GAATAGAGAGGCAAAGCAGGAG | |
| CsJAZ8 | CGGATTACAACCACCACCAC | TCGTTAGCGGCTGTTCCTGT | |
| CsZML1 | GGAGGGAGGAATCTTTGTTTGG | CTGCCAGTTTCTCCTGTTACAAC | |
| CsZML2 | CTAAACGTTCCCAACCACTTCAC | GTCGTTGAGAGAGTGAGTTTCGT | |
| CsZML3 | TGCAGCACATGAGAAATGG | GAACACATAAACCTGACCCTGG | |
| CsZML4 | CCAGAGACGAATCACCAATCC | CTCACCACCCACAACTTCATC | |
| CsZML5 | CCGATTCCACTTCAAGCAAG | TCCTCCTCCTCCGTCTTCAT | |
| CsZML6 | TCGATGAAGACGGAGGAGG | GGGTTGTTATTCACAGGGAGG | |
| CsZML7 | CGAATCCACAGCCACTTCAAG | TCCGCCTCATCCATAGCATC | |
| CsZML8 | CTGAGACCAACCACGAAGCT | TCGAATCGGATCTGAGGGTT | |
| 内参基因 | β-ACTNI | GCCATCTTTGATTGGAATGG | GGTGCCACAACCTTGATCTT |
| Reference gene | SAND | CCAATTGCCCCCTTAATGAC | GCAATCATTTCCTTCGTGGAG |
| 亚细胞定位 | CsTIFY1-1302 | ACTCTTGACCATGGTAGATCTATGATAGATAGACAGGTTGCAAATCAT | ATCTGATCCACCTTTACTAGTCTAAAAAATTGCCTTGTTCGGG |
| Subcellular | |||
| localization | CsJAZ1-1302 | ACTCTTGACCATGGTAGATCTATGGCATCGAGATCAGCTGTTG | ATCTGATCCACCTTTACTAGTCTAATCTTCATATCTTGATGTATTCTTCCC |
| CsJAZ5-1302 | ACTCTTGACCATGGTAGATCTATGTCGAGTTCGTCTGGTTCTGC | ATCTGATCCACCTTTACTAGTCTACAAATGGTGCTCAAACTGCA |
Table 1 CsTIFY gene family primer sequences
| 用途 Purpose | 基因 Gene | 上游引物序列(5′-3′) Forward primer | 下游引物序列(5′-3′) Reverse primer |
|---|---|---|---|
| qRT-PCR | CsTIFY1 | TGAGTAACCGATTCGCTGG | GCTTGTGGAAACCCATCTCTG |
| CsTIFY2 | TCAACGAGGAAGAGGCTATGG | GGGTCCTCATCCTCAACCATT | |
| CsTIFY3 | GCCTCTGTTGGTGGTTCTTC | CGGTTATTCATCTCAGTCCCT | |
| CsTIFY4 | TCTCCGCCACTTCTGATCTG | GCGAATCGGTTACTCATCTCAG | |
| CsPPD1 | CCTACCAGATGTGCGAGAATG | TCGCCTTCATCCAAGCTGAT | |
| CsPPD2 | GCAAATTCTCCATTGCTCGC | TCCACGACGGTCTTCTCATC | |
| CsJAZ1 | TCCTCCAAACCTCTTCCCAAT | CACAGAGCCAGAAGCAATCAC | |
| CsJAZ2 | TCATTGATCGCCGAAGAAGC | GAATAGAGAGGCAAAGCAGG | |
| CsJAZ4 | TGACCCACTTGCTGCTTCT | GAACGCCACCTTGTCCAT | |
| CsJAZ5 | TGTCGAGTTCGTCTGGTTCT | CCGAATGTGCCTTTCTTCTC | |
| CsJAZ6 | CGGTGGCTGTCAAAGAAGGTATT | CTCTAATCGCCATTTCAGTCCCT | |
| CsJAZ7 | TCATTGATCGCCGAAGAAGC | GAATAGAGAGGCAAAGCAGGAG | |
| CsJAZ8 | CGGATTACAACCACCACCAC | TCGTTAGCGGCTGTTCCTGT | |
| CsZML1 | GGAGGGAGGAATCTTTGTTTGG | CTGCCAGTTTCTCCTGTTACAAC | |
| CsZML2 | CTAAACGTTCCCAACCACTTCAC | GTCGTTGAGAGAGTGAGTTTCGT | |
| CsZML3 | TGCAGCACATGAGAAATGG | GAACACATAAACCTGACCCTGG | |
| CsZML4 | CCAGAGACGAATCACCAATCC | CTCACCACCCACAACTTCATC | |
| CsZML5 | CCGATTCCACTTCAAGCAAG | TCCTCCTCCTCCGTCTTCAT | |
| CsZML6 | TCGATGAAGACGGAGGAGG | GGGTTGTTATTCACAGGGAGG | |
| CsZML7 | CGAATCCACAGCCACTTCAAG | TCCGCCTCATCCATAGCATC | |
| CsZML8 | CTGAGACCAACCACGAAGCT | TCGAATCGGATCTGAGGGTT | |
| 内参基因 | β-ACTNI | GCCATCTTTGATTGGAATGG | GGTGCCACAACCTTGATCTT |
| Reference gene | SAND | CCAATTGCCCCCTTAATGAC | GCAATCATTTCCTTCGTGGAG |
| 亚细胞定位 | CsTIFY1-1302 | ACTCTTGACCATGGTAGATCTATGATAGATAGACAGGTTGCAAATCAT | ATCTGATCCACCTTTACTAGTCTAAAAAATTGCCTTGTTCGGG |
| Subcellular | |||
| localization | CsJAZ1-1302 | ACTCTTGACCATGGTAGATCTATGGCATCGAGATCAGCTGTTG | ATCTGATCCACCTTTACTAGTCTAATCTTCATATCTTGATGTATTCTTCCC |
| CsJAZ5-1302 | ACTCTTGACCATGGTAGATCTATGTCGAGTTCGTCTGGTTCTGC | ATCTGATCCACCTTTACTAGTCTACAAATGGTGCTCAAACTGCA |
| 基因名称 Gene name | ID | 氨基酸 Amino acid | 分子量/kD Molecular weight | 等电点 Isoelectric point | 内含子 Intron | 外显子 Exons | 不稳定指数 Instability index | 脂肪族指数 Aliphatic index | 平均疏水性 GRAVY | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|---|---|---|
| CsTIFY1 | TEA007552.1 | 250 | 26.77771 | 9.90 | 1 | 2 | 52.44 | 55.04 | -0.694 | 细胞核 |
| Cell nucleus | ||||||||||
| CsTTFY2 | TEA012041.1 | 934 | 103.1778 | 5.57 | 11 | 12 | 49.30 | 84.03 | -0.140 | 叶绿体 |
| Chloroplast | ||||||||||
| CsTIFY3 | TEA032457.1 | 352 | 38.90351 | 10.25 | 4 | 5 | 53.41 | 72.81 | -0.461 | 液泡 Vacuole |
| CsTIFY4 | TEA009327.1 | 393 | 41.65471 | 9.40 | 5 | 6 | 46.36 | 65.29 | -0.437 | 细胞核 |
| Cell nucleus | ||||||||||
| CsPPD1 | TEA005947.1 | 1 820 | 204.7527 | 6.14 | 16 | 17 | 41.51 | 92.61 | -0.120 | 胞间连丝 |
| Plasmodesma | ||||||||||
| CsPPD2 | TEA011549.1 | 1 114 | 125.3216 | 8.23 | 12 | 13 | 50.60 | 88.93 | -0.299 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ1 | TEA027049.1 | 212 | 23.87791 | 9.15 | 5 | 6 | 55.42 | 67.08 | -0.632 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ2 | TEA030826.1 | 148 | 16.39844 | 7.84 | 0 | 1 | 48.12 | 58.65 | -0.564 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ3 | TEA001501.1 | 261 | 29.26615 | 9.17 | 4 | 5 | 57.65 | 70.69 | -0.666 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ4 | TEA001681.1 | 410 | 42.84663 | 9.34 | 7 | 8 | 35.36 | 66.44 | -0.310 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ5 | TEA030190.1 | 294 | 31.53145 | 9.46 | 4 | 5 | 55.85 | 70.37 | -0.395 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ6 | TEA001821.1 | 436 | 46.20707 | 9.58 | 7 | 8 | 48.77 | 71.40 | -0.343 | 细胞质 Cytoplasm |
| CsJAZ7 | TEA032228.1 | 220 | 24.45353 | 9.18 | 5 | 6 | 55.48 | 67.36 | -0.557 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ8 | TEA004474.1 | 136 | 15.60498 | 9.93 | 2 | 3 | 84.54 | 75.29 | -0.721 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML1 | TEA033832.1 | 385 | 42.00050 | 4.88 | 10 | 11 | 42.88 | 70.08 | -0.625 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML2 | TEA033836.1 | 271 | 30.51872 | 4.57 | 4 | 5 | 46.48 | 78.04 | -0.601 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML3 | TEA013465.1 | 303 | 33.49678 | 6.19 | 6 | 7 | 31.87 | 60.79 | -0.916 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML4 | TEA014550.1 | 234 | 25.20185 | 5.00 | 3 | 4 | 50.61 | 71.24 | -0.550 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML5 | TEA014541.1 | 352 | 38.21245 | 4.85 | 10 | 11 | 50.11 | 72.30 | -0.574 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML6 | TEA011504.1 | 352 | 38.21245 | 4.85 | 10 | 11 | 50.11 | 72.30 | -0.574 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML7 | TEA002032.1 | 383 | 41.64484 | 4.76 | 10 | 11 | 54.89 | 63.92 | -0.795 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML8 | TEA001414.1 | 461 | 49.38947 | 5.75 | 11 | 12 | 41.39 | 73.19 | -0.375 | 细胞核 |
| Cell nucleus |
Table 2 Physicochemical properties of CsTIFY family proteins of tea tree
| 基因名称 Gene name | ID | 氨基酸 Amino acid | 分子量/kD Molecular weight | 等电点 Isoelectric point | 内含子 Intron | 外显子 Exons | 不稳定指数 Instability index | 脂肪族指数 Aliphatic index | 平均疏水性 GRAVY | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|---|---|---|
| CsTIFY1 | TEA007552.1 | 250 | 26.77771 | 9.90 | 1 | 2 | 52.44 | 55.04 | -0.694 | 细胞核 |
| Cell nucleus | ||||||||||
| CsTTFY2 | TEA012041.1 | 934 | 103.1778 | 5.57 | 11 | 12 | 49.30 | 84.03 | -0.140 | 叶绿体 |
| Chloroplast | ||||||||||
| CsTIFY3 | TEA032457.1 | 352 | 38.90351 | 10.25 | 4 | 5 | 53.41 | 72.81 | -0.461 | 液泡 Vacuole |
| CsTIFY4 | TEA009327.1 | 393 | 41.65471 | 9.40 | 5 | 6 | 46.36 | 65.29 | -0.437 | 细胞核 |
| Cell nucleus | ||||||||||
| CsPPD1 | TEA005947.1 | 1 820 | 204.7527 | 6.14 | 16 | 17 | 41.51 | 92.61 | -0.120 | 胞间连丝 |
| Plasmodesma | ||||||||||
| CsPPD2 | TEA011549.1 | 1 114 | 125.3216 | 8.23 | 12 | 13 | 50.60 | 88.93 | -0.299 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ1 | TEA027049.1 | 212 | 23.87791 | 9.15 | 5 | 6 | 55.42 | 67.08 | -0.632 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ2 | TEA030826.1 | 148 | 16.39844 | 7.84 | 0 | 1 | 48.12 | 58.65 | -0.564 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ3 | TEA001501.1 | 261 | 29.26615 | 9.17 | 4 | 5 | 57.65 | 70.69 | -0.666 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ4 | TEA001681.1 | 410 | 42.84663 | 9.34 | 7 | 8 | 35.36 | 66.44 | -0.310 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ5 | TEA030190.1 | 294 | 31.53145 | 9.46 | 4 | 5 | 55.85 | 70.37 | -0.395 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ6 | TEA001821.1 | 436 | 46.20707 | 9.58 | 7 | 8 | 48.77 | 71.40 | -0.343 | 细胞质 Cytoplasm |
| CsJAZ7 | TEA032228.1 | 220 | 24.45353 | 9.18 | 5 | 6 | 55.48 | 67.36 | -0.557 | 细胞核 |
| Cell nucleus | ||||||||||
| CsJAZ8 | TEA004474.1 | 136 | 15.60498 | 9.93 | 2 | 3 | 84.54 | 75.29 | -0.721 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML1 | TEA033832.1 | 385 | 42.00050 | 4.88 | 10 | 11 | 42.88 | 70.08 | -0.625 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML2 | TEA033836.1 | 271 | 30.51872 | 4.57 | 4 | 5 | 46.48 | 78.04 | -0.601 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML3 | TEA013465.1 | 303 | 33.49678 | 6.19 | 6 | 7 | 31.87 | 60.79 | -0.916 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML4 | TEA014550.1 | 234 | 25.20185 | 5.00 | 3 | 4 | 50.61 | 71.24 | -0.550 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML5 | TEA014541.1 | 352 | 38.21245 | 4.85 | 10 | 11 | 50.11 | 72.30 | -0.574 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML6 | TEA011504.1 | 352 | 38.21245 | 4.85 | 10 | 11 | 50.11 | 72.30 | -0.574 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML7 | TEA002032.1 | 383 | 41.64484 | 4.76 | 10 | 11 | 54.89 | 63.92 | -0.795 | 细胞核 |
| Cell nucleus | ||||||||||
| CsZML8 | TEA001414.1 | 461 | 49.38947 | 5.75 | 11 | 12 | 41.39 | 73.19 | -0.375 | 细胞核 |
| Cell nucleus |
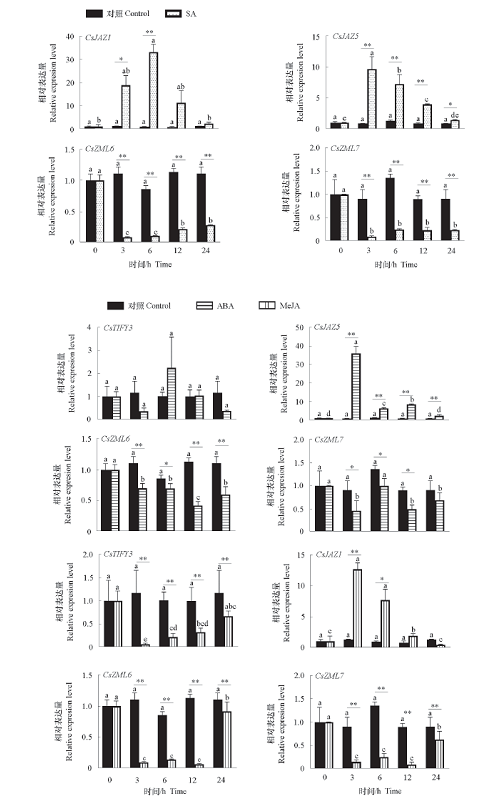
Fig. 5 Expression of CsTIFY genes in tea plants under exogenous SA,ABA,MeJA stresses * indicates significant differences at the same time point(α = 0.05),** indicates extremely significant(α = 0.01);Different lowercase letters indicate significant differences at different points in time(P < 0.05). The same below.
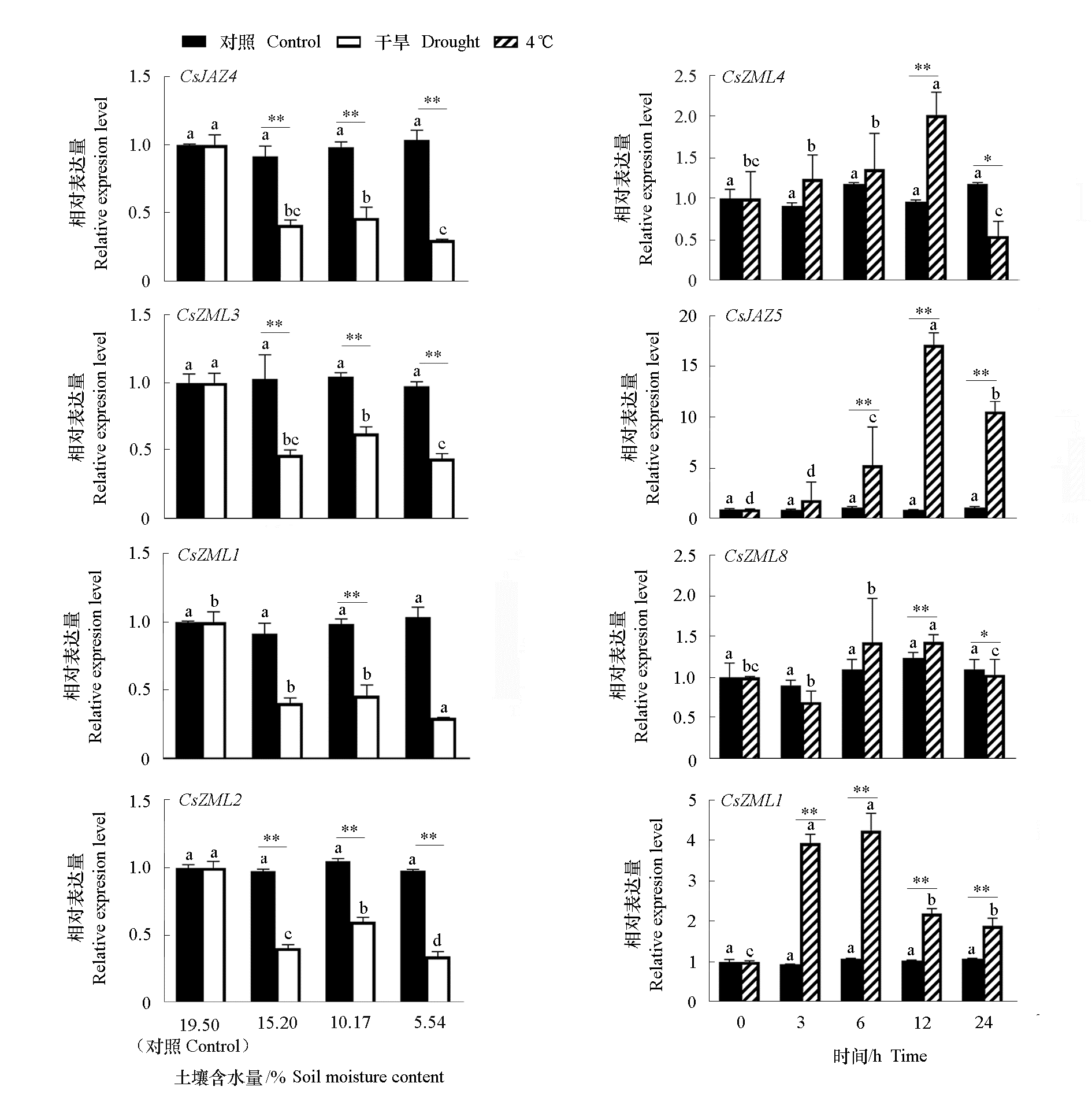
Fig. 6 CsTIFY gene expression of tea under abiotic stress drought and 4 ℃ treatments CsTIFY gene expression of tea under abiotic stress drought and 4 ℃ treatments
| [1] | Bartel V, Wim G, Alex B, akayuki K, Gheysen, Godelieve. 2007. The tify family previously known as ZIM. Trends in Plant Science, 12 (6):239-244. |
| [2] | Ding Y, Wang Y, Qiu C, Qian W, Xie H, Ding Z. 2020. Alternative splicing in tea plants was extensively triggered by drought,heat and their combined stresses. PeerJ:e8258. |
| [3] | Dong Taoxing, Cai Kunzheng, Zhang Jingxin, Rong Hui, Xie Guozheng, Zeng Rensen. 2007. The physiological roles of methyl jasmonate(MeJA) in drought resistance of rice seedlings. Ecology and Environmental Sciences,(4):1261-1265. (in Chinese) |
| 董桃杏, 蔡昆争, 张景欣, 荣辉, 谢国政, 曾任森. 2007. 茉莉酸甲酯(MeJA)对水稻幼苗的抗旱生理效应. 生态环境,(4):1261-1265. | |
| [4] |
Ebel C, BenFeki A, Hanin M, Solano R, Chini A. 2018. Characterization of wheat(Triticum aestivum)TIFY family and role of Triticum durum TdTIFY11a in salt stress tolerance. PLoS ONE, 13 (7):e0200566.
doi: 10.1371/journal.pone.0200566 URL |
| [5] |
Guo Y, Zhao S, Zhu C, Chang X, Yue C, Wang Z, Lin Y, Lai Z. 2017. Identification of drought-responsive miRNAs and physiological characterization of tea plant(Camellia sinensis L.)under drought stress. BMC Plant Biology, 17 (1):211.
doi: 10.1186/s12870-017-1172-6 URL |
| [6] |
Hause B, Wasternack C. 2013. Jasmonates:biosynthesis,perception,signal transduction and action in plant stress response,growth and development. An update to the 2007 review in Annals of Botany. Annals of Botany,(6):1021-1058.
doi: 10.1093/aob/mct067 pmid: 23558912 |
| [7] | Hong G, Xue X, Mao Y. 2012. Arabidopsis MYC 2 interacts with DELLA proteins in regulating sesquiterpene synthase gene expression. Plant Cell, (6):2635-2648. |
| [8] | Huang Ying. 2017. Analysis of TIFY gene family and its interaction protein screening in Salvia miltiorrhiza[Ph. D. Dissertation]. Xi'an: Shaanxi Normal University. (in Chinese) |
| 黄英. 2017. 丹参TIFY基因家族分析及其互作蛋白筛选[博士论文]. 西安: 陕西师范大学. | |
| [9] | Huang Z, Jin S, Guo H, Zhong X, He J, Li X, Jiang M, Yu X, Long H, Ma M, Chen Q. 2016. Genome-wide identification and characterization of TIFY family genes in Moso bamboo(Phyllostachys edulis)and expression profiling analysis under dehydration and cold stresses. PeerJ:4. |
| [10] |
Li N, Yue C, Cao H, Qian W, Hao X, Wang Y, Wang L, Ding C, Wang X, Yang Y. 2018. Transcriptome sequencing dissection of the mechanisms underlying differential cold sensitivity in young and mature leaves of the tea plan(Camellia sinensis). Journal of Plant Physiology, 224-225:144-155.
doi: 10.1016/j.jplph.2018.03.017 URL |
| [11] | Li Shuyu, Li Chuanyou. 2016. Developmental plasticity of plant roots. China Basic Science, 18 (2):14-21. (in Chinese) |
| 李淑钰, 李传友. 2016. 植物根系可塑性发育的研究进展与展望. 中国基础科学, 18 (2):14-21. | |
| [12] | Li X, Yin X, Wang H, Li J, Guo C, Gao H, Zheng Y, Fan C, Wang X. 2015. Genome-wide identification and analysis of the apple(Malus × domestica Borkh.)TIFY gene family. Tree Genetics & Genomes,(1):1-13. |
| [13] | Mei Chuang, Zhang Xiaoyan, Yan Peng, Aisajan Mamat, Feng Beibei, Ma Kai, Han Liqun, Dong Lianxin, Wang Jixun. 2021. Identification of TIFY family in apple and their expression analysis under insect stress. Acta Horticulturae Sinica, 48 (2):233-242. (in Chinese) |
| 梅闯, 张小燕, 闫鹏, 艾沙江 · 买买提, 冯贝贝, 马凯, 韩立群, 董连新, 王继勋. 2021. 苹果TIFY 家族基因鉴定及其在虫害胁迫下的表达分析. 园艺学报, 48 (2):233-242. | |
| [14] |
Meng L, Zhang T, Geng S, Scott P B, Li H, Chen S. 2019. Comparative proteomics and metabolomics of JAZ7-mediated drought tolerance in Arabidopsis. Journal of Proteomics: 196:81-91.
doi: S1874-3919(19)30036-3 pmid: 30731210 |
| [15] | Nishii A, Takemura M, Fujita H, Shikata M, Yokota A, Kohchi T. 2000. Characterization of a novel gene encoding a putative single Zinc-finger protein,ZIM,expressed during the reproductive phase in Arabidopsis thaliana. Bioscience,Biotechnology,and Biochemistry,(7):1402-1409. |
| [16] | Qing Z, Muchen C, Xiaomin Y, Lishan W, Chunfang G, Ray M, Jisen Z. 2017. Transcriptome dynamics of Camellia sinensis in response to continuous salinity and drought stress. Tree Genetics & Genomes,(4):1-17. |
| [17] |
Sarfraz H S, Akhtar K M, Muhammad A. 2011. Transcription factors as tools to engineer enhanced drought stress tolerance in plants. Biotechnology Progress, 27 (2):297-306.
doi: 10.1002/btpr.514 pmid: 21302367 |
| [18] |
Shikata M, Matsuda Y, Ando K, Nishii A, Takemura M, Yokota A, Kohchi T. 2004. Characterization of Arabidopsis ZIM,a member of a novel plant-specific GATA factor gene family. Journal of Experimental Botany, 55 (397):631-639.
doi: 10.1093/jxb/erh078 URL |
| [19] | Shuichi Y. 1998. Transcription factors in plants:physiological functions and regulation of expression. Journal of Plant Research,(1103):363-371. |
| [20] | Sirhindi G A, Sharma P A, Arya P B C, Goel P B C, Kumar G B C, Acharya V B C, Singh A K B C. 2016. Genome-wide characterization and expression profiling of TIFY gene family in pigeonpea[Cajanus cajan(L.)Mill sp.] under copper stress(Article). Journal of Plant Biochemistry and Biotechnology,(3):301-310. |
| [21] | Song Yun, Li Linxuan, Zhuo Fengping, Zhang Xueyan, Ren Maozhi, Li Fuguang. 2015. Progress on jasmonic acid signaling in plant stress resistant. Journal of Agricultural Science and Technology, 17 (2):17-24. (in Chinese) |
| 宋云, 李林宣, 卓凤萍, 张雪妍, 任茂智, 李付广. 2015. 茉莉酸信号传导在植物抗逆性方面研究进展. 中国农业科技导报, 17 (2): 17-24. | |
| [22] | Su Wenbing, Jiang Yuanyuan, Bai Yunlu, Gan Xiaoqing, Liu Yuexue, Lin Shunquan. 2019. Journal of Agricultural Biotechnology, 27 (5):919-926. (in Chinese) |
| 苏文炳, 蒋园园, 白昀鹭, 甘小清, 刘月学, 林顺权. 2019. 转录因子调控植物萜类化合物生物合成研究进展. 农业生物技术学报, 27 (5): 919-926. | |
| [23] | Sun Kaiwen. 2019. Regulation of the plant hormone abscisic acid on the defense response mediated by jasmonic acid[Ph. D. Dissertation]. Nanjing: Nanjing Normal University. (in Chinese) |
| 孙凯文. 2019. 植物激素脱落酸对茉莉酸介导的防御反应的调控[博士论文]. 南京: 南京师范大学. | |
| [24] |
Vanholme B, Grunewald W, Bateman A, Kohchi T, Gheysen G. 2007. The TIFY family previously known as ZIM. Trends in Plant Science, 12 (6):239-244..
pmid: 17499004 |
| [25] | Wang Linhua, Liang Shurong, Lü Shumin, Zhao Huijie, Qu Xiaofei, Liu Weiwei. 2010. Protective effect of SA on photosynthetic apparatus of wheat leaves under heat and high irradiance stress. Journal of Henan Agricultural Sciences,(4):21-25. (in Chinese) |
| 王林华, 梁书荣, 吕淑敏, 赵会杰, 曲小菲, 刘魏魏. 2010. 水杨酸对高温强光胁迫下小麦叶片光合机构的保护作用. 河南农业科学,(4):21-25. | |
| [26] | Wang Niyan, Jiang De’an. 2002. Jasmonic acid and its methyl esters and plant induced disease resistance. Plant Physiology Journal,(3):279-284. (in Chinese) |
| 王妮妍, 蒋德安. 2002. 茉莉酸及其甲酯与植物诱导抗病性. 植物生理学通讯,(3):279-284. | |
| [27] |
Wang Y, Pan F, Chen D, Chu W, Liu H, Xiang Y. 2017. Genome-wide identification and analysis of the Populus trichocarpa TIFY gene family. Plant Physiology and Biochemistry, 115:360-371.
doi: S0981-9428(17)30135-3 pmid: 28431355 |
| [28] | White D W R. 2006. PEAPOD regulates lamina size and curvature in Arabidopsis. Proceedings of the National Academy of Sciences of the United States of America, 103 (35):13238-13243. |
| [29] | Wu Yanyan. 2019. Effects of exogenous ABA on physiological characteristics of Machilus machinei seedlings under drought stress[Ph. D. Dissertation]. Chongqing: Southwest University. (in Chinese) |
| 吴琰琰. 2019. 外源ABA对干旱胁迫下桢楠幼苗生理特性的影响[博士论文]. 重庆: 西南大学. | |
| [30] | Xia E H, Li F D, Tong W, Li P H, Wu Q, Zhao H J, Ge R H, Li R P, Li Y Y, Zhang Z Z, Wei C L, Wan X C. 2019. Tea plant information archive:a comprehensive genomics and bioinformatics platform for tea plant. Plant Biotechnology Journal,(10):1938-1953. |
| [31] | Xu G, Guo C, Shan H. 2012. Divergence of duplicate genes in exon-intron structure. Proceedings of The National Academy of Sciences of The United States of America,(4):1187-1192. |
| [32] |
Yang Ruijia, Zhang Zhongbao, Wu Zhongyi. 2020. Progress of the structural and functional analysis of plant transcription factor TIFY protein family. Biotechnology Bulletin, 36 (12):121-128. (in Chinese)
doi: 10.13560/j.cnki.biotech.bull.1985.2020-0307 |
|
杨锐佳, 张中保, 吴忠义. 2020. 植物转录因子TIFY家族蛋白结构和功能的研究进展. 生物技术通报, 36 (12):121-128.
doi: 10.13560/j.cnki.biotech.bull.1985.2020-0307 |
|
| [33] | Yang Y, Ahammed G J, Wan C, Liu H, Chen R, Zhou Y. 2019. Comprehensive analysis of TIFY transcription factors and their expression profiles under jasmonic acid and abiotic stresses in watermelon. International Journal of Genomics, 10.1155/2019/6813086. |
| [34] | Ye H, Du H, Tang N, Li X, Xiong L. 2009. Identification and expression profiling analysis of TIFY family genes involved in stress and phytohormone responses in rice. Plant Molecular Biology,(3):291-305. |
| [35] | Yin Chao, Li Mei, Liu Yuhui. 2013. The role of the nucleus in plant defense response. Current Biotechnology, 3 (1):12-17. (in Chinese) |
| 尹超, 李梅, 刘昱辉. 2013. 细胞核在植物防御反应中的作用. 生物技术进展, 3 (1):12-17. | |
| [36] |
Yu Hong, Wang Xianyun, Yang Xiaohui, Li Haitao, Xiao Shuguang, Du Huatang, Hou Zhenghui. 2009. Relationship between microclimatic factors and yield of spring tea Camellia sinensis(L.)O. Kuntze in Henan Jigongshan Nature Reserve. Chinese Journal of Eco-Agriculture, 17 (1):94-99. (in Chinese)
doi: 10.3724/SP.J.1011.2009.00094 URL |
| 喻泓, 王贤赟, 杨晓晖, 李海涛, 肖曙光, 杜化堂, 侯正辉. 2009. 河南鸡公山茶园春茶产量与小气候关系研究. 中国生态农业学报, 17 (1):94-99. | |
| [37] | Zhang L, You J, Chan Z. 2015. Identification and characterization of TIFY family genes in Brachypodium distachyon. Journal of Plant Research,(6):995-1005. |
| [38] | Zhang Y, Gao M, Singer S D, Fei Z, Wang H, Wang X. 2012. Genome-wide identification and analysis of the TIFY gene family in grape. PLoS ONE,(9):1-13. |
| [39] | Zhao Xiaoxiao, Xie Kunliang, Zhang Shumeng, Zhang Chao, Xi Yajun, Sun Fengli. 2019. Identification and analysis of TIFY gene family in Switchgrass. Acta Agrestia Sinica, 27 (5):1126-1137. (in Chinese) |
| 赵晓晓, 谢坤良, 张舒梦, 张超, 奚亚军, 孙风丽. 2019. 柳枝稷TIFY基因家族的鉴定与分析. 草地学报, 27 (5):1126-1137. | |
| [40] | Zheng Y, Chen X, Wang P, Sun Y, Yue C, Ye N. 2020. Genome-wide and expression pattern analysis of JAZ family involved in stress responses and postharvest processing treatments in Camellia sinensis. Scientific Reports,(1):2792. |
| [41] |
Zhou C, Zhu C, Fu H, Li X, Chen L, Lin Y, Lai Z, Guo Y. 2019. Genome-wide investigation of superoxide dismutase(SOD)gene family and their regulatory miRNAs reveal the involvement in abiotic stress and hormone response in tea plant(Camellia sinensis). PLoS ONE,doi: 10.1371/journal.pone.0223609.
doi: 10.1371/journal.pone.0223609 |
| [42] |
Zhu D, Bai X, Chen C, Chen Q, Cai H, Li Y, Ji W, Zhai H, Lv D, Luo X, Zhu Y. 2011. GsTIFY10,a novel positive regulator of plant tolerance to bicarbonate stress and a repressor of jasmonate signaling. Plant Molecular Biology, 77 (3):285-297.
doi: 10.1007/s11103-011-9810-0 URL |
| [1] | ZHANG Xin, QI Yanxiang, ZENG Fanyun, WANG Yanwei, XIE Peilan, XIE Yixian, and PENG Jun. Functional Analysis of Dicer-like Genes in Fusarium oxysporum f. sp. cubense Race 4 [J]. Acta Horticulturae Sinica, 2023, 50(2): 279-294. |
| [2] | LIANG Jiali, WU Qisong, CHEN Guangquan, ZHANG Rong, XU Chunxiang, and FENG Shujie, . Identification of the Neopestalotiopsis musae Pathogen of Banana Leaf Spot Disease [J]. Acta Horticulturae Sinica, 2023, 50(2): 410-420. |
| [3] | WANG Xiqing, JIA Yunhe, YAN Wen, FU Yongkai, YOU Haibo, LI Dongyan, and ZHAO Jingchao. A New Watermelon Cultivar‘Longsheng Jiali’with High Resistance to Fusarium Wilt [J]. Acta Horticulturae Sinica, 2023, 50(2): 455-456. |
| [4] | ZHAI Hanhan, ZHAI Yujie, TIAN Yi, ZHANG Ye, YANG Li, WEN Zhiliang, CHEN Haijiang. Genome-wide Identification of Peach SAUR Gene Family and Characterization of PpSAUR5 Gene [J]. Acta Horticulturae Sinica, 2023, 50(1): 1-14. |
| [5] | YUAN Xin, XU Yunhe, ZHANG Yupei, SHAN Nan, CHEN Chuying, WAN Chunpeng, KAI Wenbin, ZHAI Xiawan, CHEN Jinyin, GAN Zengyu. Studies on AcAREB1 Regulating the Expression of AcGH3.1 During Postharvest Ripening of Kiwifruit [J]. Acta Horticulturae Sinica, 2023, 50(1): 53-64. |
| [6] | YU Yangjun, SU Tongbing, ZHANG Fenglan, ZHANG Deshuang, ZHAO Xiuyun, YU Shuancang, WANG Weihong, LI Peirong, XIN Xiaoyun, WANG Jiao, and WU Changjian. A New Purple Seedling-Edible Chinese Cabbage F1 Hybrid‘Jingyan Zikuaicai’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 91-92. |
| [7] | LI Zhengli, ZHANG Li, and MA Licang. A New Capsicum frutescens Cultivar‘Huangla Chaotian’with Yellow Fruit [J]. Acta Horticulturae Sinica, 2022, 49(S2): 123-124. |
| [8] | YE Zhiqin and YANG Juan. A New Chrysanthemum Cultivar‘Zibinyun’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 173-174. |
| [9] | ZHAO Yanli, LI Zhanhong, CAO Qin, DAI Miaofei, GAO Mengmeng, SUN Zhenzhu, LI Huikuan, LI Yonghua, BIAN Shuxun, and HUANG Gan. A New Chrysanthemum Cultivar‘Bianjing Qingdianhuang’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 175-176. |
| [10] | CAO Qin, ZHAO Yanli, LI Zhanhong, GAO Mengmeng, DAI Miaofei, LI Yonghua, LI Huikuan, SUN Zhenzhu, BIAN Shuxun, and LI Fei. A New Muitiflora Chrysanthemum Cultivar‘Bianjing Zijingling’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 177-178. |
| [11] | CHU Zhuannan, LI Weiwen, DONG Ling, PENG Xingxing, CUI Guangsheng, TANG Maogui, ZHANG Qinghua, SHEN Wensheng, ZHAO Wei, XIONG Rui, HAN Piao, WU Jiqiu, LI Yulong, YANG Yueying, and ZHANG Qiling. A New Chrysanthemum morifolium Cultivar‘Gongju 1’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 179-180. |
| [12] | XU Tong, LOU Jianhua, SHI Xiaohua, XIA Yiping, LI Danqing, and ZHANG Jiaping, . A New Iris ensata var. hortensis Cultivar‘Summer Power’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 203-204. |
| [13] | GAO Xiaoyu, QIN Xinsheng, XIAO Guankang, and FENG Zhijian. A New Hibiscus rosa-sinensis Cultivar‘Bohea Marchito’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 261-262. |
| [14] | SUN Simiao, WANG Kun, GAO Yuan, WANG Dajiang, and LI Lianwen. A New Ornamental Crabapple Cultivar‘Zichen’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 267-268. |
| [15] | LIANG Guichan, LIANG Jian, LIU Shihan, XI Ruchun, DAI Seping, LI Xiaodong, and DENG Xiaomei, . A New Lagerstroemia speciosa Cultivar‘Zijuan’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 169-170. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd