
Acta Horticulturae Sinica ›› 2023, Vol. 50 ›› Issue (5): 985-999.doi: 10.16420/j.issn.0513-353x.2022-0273
• Genetic & Breeding·Germplasm Resources·Molecular Biology • Previous Articles Next Articles
LI Guobin, CAI Liangyu, XIAO Licheng, WANG Jiafa, ZHANG Dedi, ZHANG Junhong( )
)
Received:2022-11-27
Revised:2023-01-16
Online:2023-05-25
Published:2023-05-31
Contact:
ZHANG Junhong
E-mail:zhangjunhng@hzau.edu.cn
CLC Number:
LI Guobin, CAI Liangyu, XIAO Licheng, WANG Jiafa, ZHANG Dedi, ZHANG Junhong. Creating Pink Fruit and Fruit Without Green Shoulder Using CRISPR/ Cas9 Technology in Tomato[J]. Acta Horticulturae Sinica, 2023, 50(5): 985-999.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2022-0273
| 基因 Gene | sgRNA 靶标序列 Target sequence of sgRNA |
|---|---|
| SlGLK2-sgRNA1 | TGGTGTCGATTAAATTCCCA |
| SlGLK2-sgRNA2 | TGAATAGCAGGTTCTTCACA |
| SlMYB12-sgRNA1 | AACACCTTGTTGTGAAAAA |
| SlMYB12-sgRNA2 | CCAAGAGCCTTCTCCATTAG |
Table 1 Target sites of SlGLK2 and SlMYB12 gene
| 基因 Gene | sgRNA 靶标序列 Target sequence of sgRNA |
|---|---|
| SlGLK2-sgRNA1 | TGGTGTCGATTAAATTCCCA |
| SlGLK2-sgRNA2 | TGAATAGCAGGTTCTTCACA |
| SlMYB12-sgRNA1 | AACACCTTGTTGTGAAAAA |
| SlMYB12-sgRNA2 | CCAAGAGCCTTCTCCATTAG |
| 引物 Primer | 用途 Purpose | 序列(5′-3′) Sequence |
|---|---|---|
| SlGLK2-CRISPR | 构建载体Construction vector | F:GAATCTAACAGTGTAGTTTGTGGTGTCGATTAAATTCCCAGTTT TAGAGC TAGAAATAG |
| R:GCTATTTCTAGCTCTAAAACTGAATAGCAGGTTCTTCACACAA ACTACACTG TTAGATT | ||
| SlMYB12-CRISPR | 构建载体Construction vector | F:GAATCTAACAGTGTAGTTTGAACACCTTGTTGTGAAAAAGGTT TTAGAGCT AGAAATAG |
| R:GCTATTTCTAGCTCTAAAACCCAAGAGCCTTCTCCATTAGCAA ACTACACTG TTAGATT | ||
| SlGLK2 | 检测靶点Detection target | F:GATGCTGGCATTTGTGTGATTC |
| R:TTAATTACGACTAGAACCGTCAACG | ||
| SlMYB12 | 检测靶点Detection target | F:TAACTAGACTAAAGTGAAGAGCGGG |
| R:CTAATTACCTGTTACCCAAAGTTGC |
Table 2 Primer sequence used for construct CRISPR vector
| 引物 Primer | 用途 Purpose | 序列(5′-3′) Sequence |
|---|---|---|
| SlGLK2-CRISPR | 构建载体Construction vector | F:GAATCTAACAGTGTAGTTTGTGGTGTCGATTAAATTCCCAGTTT TAGAGC TAGAAATAG |
| R:GCTATTTCTAGCTCTAAAACTGAATAGCAGGTTCTTCACACAA ACTACACTG TTAGATT | ||
| SlMYB12-CRISPR | 构建载体Construction vector | F:GAATCTAACAGTGTAGTTTGAACACCTTGTTGTGAAAAAGGTT TTAGAGCT AGAAATAG |
| R:GCTATTTCTAGCTCTAAAACCCAAGAGCCTTCTCCATTAGCAA ACTACACTG TTAGATT | ||
| SlGLK2 | 检测靶点Detection target | F:GATGCTGGCATTTGTGTGATTC |
| R:TTAATTACGACTAGAACCGTCAACG | ||
| SlMYB12 | 检测靶点Detection target | F:TAACTAGACTAAAGTGAAGAGCGGG |
| R:CTAATTACCTGTTACCCAAAGTTGC |

Fig. 3 Targeted mutagenesis of SlGLK2 gene in gene-edited tomato The red marker is the target. The bold font is PAM. The blue plus and the horizontal lines are the base insertions and deletions respectively. The green markers are the base substitutions. The same below.

Fig. 5 Amino acid sequence analysis of the SlGLK2 and SlMYB12 in gene editing transgenic plants of AC,B5 and B9 tomatos The red marker is the target. Blue represents the mutated amino acid. The dash indicates the stop codon.
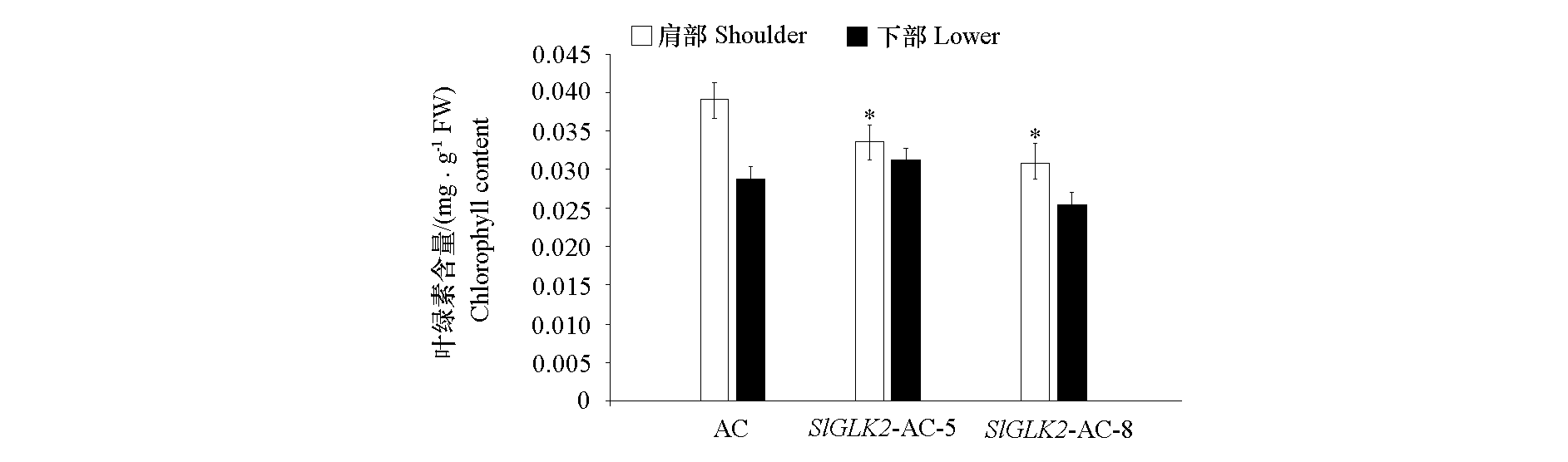
Fig. 8 The chlorophyll content of fruits in SlGLK2-AC gene-edited plants * indicates that there is a significant difference in the values shown of white histogram as the control AC(P < 0.05).
| 材料 Material | 单果质量/g Fruit weight | 果实纵径/mm Fruit length | 果实横径/mm Fruit diameter | 果形指数 Index of fruit shape | 果实硬度/(× 105 Pa) Fruit hardness |
|---|---|---|---|---|---|
| AC | 28.95 ± 0.83 b | 34.51 ± 0.37 c | 38.02 ± 0.77 c | 0.90 ± 0.03 a | 8.14 ± 0.37 b |
| SlMYB12-AC-3 | 28.82 ± 0.75 b | 34.45 ± 0.76 c | 37.79 ± 0.64 c | 0.91 ± 0.02 a | 7.86 ± 0.52 b |
| SlMYB12-AC-4 | 29.46 ± 1.02 b | 34.04 ± 0.25 c | 38.03 ± 0.92 c | 0.89 ± 0.02 a | 8.32 ± 0.41 b |
| SlMYB12-AC-7 | 28.03 ± 0.62 b | 34.26 ± 0.43 c | 37.65 ± 0.43 c | 0.91 ± 0.01 a | 8.22 ± 0.26 b |
| SlGLK2-AC-3 | 27.97 ± 0.69 b | 33.52 ± 0.21 c | 37.63 ± 0.26 c | 0.89 ± 0.03 a | 8.47 ± 0.67 b |
| SlGLK2-AC-4 | 28.84 ± 1.25 b | 33.96 ± 0.64 c | 38.12 ± 0.71 c | 0.89 ± 0.02 a | 7.92 ± 0.85 b |
| SlGLK2-AC-5 | 28.88 ± 0.48 b | 34.62 ± 0.41 c | 38.46 ± 0.65 c | 0.90 ± 0.01 a | 8.08 ± 0.56 b |
| B5 | 80.82 ± 0.67 a | 41.42 ± 1.21 ab | 53.96 ± 1.84 ab | 0.76 ± 0.04 b | 11.33 ± 1.08 a |
| SlGLK2-B5-1 | 79.94 ± 1.23 a | 39.61 ± 1.73 b | 52.75 ± 2.17 b | 0.75 ± 0.06 b | 10.76 ± 0.84 a |
| SlGLK2-B5-2 | 81.89 ± 1.06 a | 43.30 ± 1.55 a | 55.53 ± 1.49 a | 0.77 ± 0.03 b | 11.59 ± 0.86 a |
| SlGLK2-B5-3 | 81.03 ± 1.35 a | 41.42 ± 1.83 ab | 55.02 ± 2.27 a | 0.75 ± 0.05 b | 10.94 ± 1.23 a |
Table 3 Fruit traits of gene-edited plants
| 材料 Material | 单果质量/g Fruit weight | 果实纵径/mm Fruit length | 果实横径/mm Fruit diameter | 果形指数 Index of fruit shape | 果实硬度/(× 105 Pa) Fruit hardness |
|---|---|---|---|---|---|
| AC | 28.95 ± 0.83 b | 34.51 ± 0.37 c | 38.02 ± 0.77 c | 0.90 ± 0.03 a | 8.14 ± 0.37 b |
| SlMYB12-AC-3 | 28.82 ± 0.75 b | 34.45 ± 0.76 c | 37.79 ± 0.64 c | 0.91 ± 0.02 a | 7.86 ± 0.52 b |
| SlMYB12-AC-4 | 29.46 ± 1.02 b | 34.04 ± 0.25 c | 38.03 ± 0.92 c | 0.89 ± 0.02 a | 8.32 ± 0.41 b |
| SlMYB12-AC-7 | 28.03 ± 0.62 b | 34.26 ± 0.43 c | 37.65 ± 0.43 c | 0.91 ± 0.01 a | 8.22 ± 0.26 b |
| SlGLK2-AC-3 | 27.97 ± 0.69 b | 33.52 ± 0.21 c | 37.63 ± 0.26 c | 0.89 ± 0.03 a | 8.47 ± 0.67 b |
| SlGLK2-AC-4 | 28.84 ± 1.25 b | 33.96 ± 0.64 c | 38.12 ± 0.71 c | 0.89 ± 0.02 a | 7.92 ± 0.85 b |
| SlGLK2-AC-5 | 28.88 ± 0.48 b | 34.62 ± 0.41 c | 38.46 ± 0.65 c | 0.90 ± 0.01 a | 8.08 ± 0.56 b |
| B5 | 80.82 ± 0.67 a | 41.42 ± 1.21 ab | 53.96 ± 1.84 ab | 0.76 ± 0.04 b | 11.33 ± 1.08 a |
| SlGLK2-B5-1 | 79.94 ± 1.23 a | 39.61 ± 1.73 b | 52.75 ± 2.17 b | 0.75 ± 0.06 b | 10.76 ± 0.84 a |
| SlGLK2-B5-2 | 81.89 ± 1.06 a | 43.30 ± 1.55 a | 55.53 ± 1.49 a | 0.77 ± 0.03 b | 11.59 ± 0.86 a |
| SlGLK2-B5-3 | 81.03 ± 1.35 a | 41.42 ± 1.83 ab | 55.02 ± 2.27 a | 0.75 ± 0.05 b | 10.94 ± 1.23 a |
| 材料 Material | 可溶性固形物/% Soluble solid | 可滴定酸/% Titrable acid | 固酸比 S/T | 总维生素C/(mg · g-1) Total vitamin C | 还原维生素C/(mg · g-1) Reduced vitamin C |
|---|---|---|---|---|---|
| AC | 5.7 ± 0.34 a | 0.62 ± 0.21 b | 9.19 ± 0.87 a | 0.09 ± 0.01 b | 0.09 ± 0.01 c |
| SlMYB12-AC-3 | 5.3 ± 0.27 b | 0.69 ± 0.17 b | 7.68 ± 1.21 b | 0.08 ± 0.02 b | 0.08 ± 0.01 c |
| SlMYB12-AC-4 | 5.5 ± 0.61 a | 0.64 ± 0.13 b | 8.59 ± 0.96 ab | 0.12 ± 0.02 b | 0.09 ± 0.01 c |
| SlMYB12-AC-7 | 5.8 ± 0.29 a | 0.63 ± 0.24 b | 9.21 ± 1.35 a | 0.10 ± 0.01 b | 0.09 ± 0.01 c |
| SlGLK2-AC-3 | 5.9 ± 0.53 a | 0.68 ± 0.30 b | 8.67 ± 1.07 ab | 0.11 ± 0.01 b | 0.10 ± 0.01 c |
| SlGLK2-AC-4 | 5.6 ± 0.28 a | 0.66 ± 0.27 b | 8.48 ± 0.67 ab | 0.11 ± 0.02 b | 0.09 ± 0.01 c |
| SlGLK2-AC-5 | 5.5 ± 0.36 a | 0.63 ± 0.18 b | 8.73 ± 1.26 a | 0.11 ± 0.02 b | 0.09 ± 0.01 c |
| B5 | 6.0 ± 0.27 a | 1.10 ± 0.22 a | 5.45 ± 0.93 c | 0.20 ± 0.03 a | 0.19 ± 0.02 a |
| SlGLK2-B5-1 | 6.0 ± 0.34 a | 1.04 ± 0.17 a | 5.77 ± 0.72 c | 0.16 ± 0.04 a | 0.15 ± 0.03 b |
| SlGLK2-B5-2 | 5.6 ± 0.51 a | 1.08 ± 0.31 a | 5.18 ± 0.64 c | 0.20 ± 0.03 a | 0.17 ± 0.02 a |
| SlGLK2-B5-3 | 5.8 ± 0.43 a | 1.21 ± 0.37 a | 4.79 ± 1.37 d | 0.18 ± 0.03 a | 0.16 ± 0.03 ab |
Table 4 Fruits quality traits of gene-edited plants
| 材料 Material | 可溶性固形物/% Soluble solid | 可滴定酸/% Titrable acid | 固酸比 S/T | 总维生素C/(mg · g-1) Total vitamin C | 还原维生素C/(mg · g-1) Reduced vitamin C |
|---|---|---|---|---|---|
| AC | 5.7 ± 0.34 a | 0.62 ± 0.21 b | 9.19 ± 0.87 a | 0.09 ± 0.01 b | 0.09 ± 0.01 c |
| SlMYB12-AC-3 | 5.3 ± 0.27 b | 0.69 ± 0.17 b | 7.68 ± 1.21 b | 0.08 ± 0.02 b | 0.08 ± 0.01 c |
| SlMYB12-AC-4 | 5.5 ± 0.61 a | 0.64 ± 0.13 b | 8.59 ± 0.96 ab | 0.12 ± 0.02 b | 0.09 ± 0.01 c |
| SlMYB12-AC-7 | 5.8 ± 0.29 a | 0.63 ± 0.24 b | 9.21 ± 1.35 a | 0.10 ± 0.01 b | 0.09 ± 0.01 c |
| SlGLK2-AC-3 | 5.9 ± 0.53 a | 0.68 ± 0.30 b | 8.67 ± 1.07 ab | 0.11 ± 0.01 b | 0.10 ± 0.01 c |
| SlGLK2-AC-4 | 5.6 ± 0.28 a | 0.66 ± 0.27 b | 8.48 ± 0.67 ab | 0.11 ± 0.02 b | 0.09 ± 0.01 c |
| SlGLK2-AC-5 | 5.5 ± 0.36 a | 0.63 ± 0.18 b | 8.73 ± 1.26 a | 0.11 ± 0.02 b | 0.09 ± 0.01 c |
| B5 | 6.0 ± 0.27 a | 1.10 ± 0.22 a | 5.45 ± 0.93 c | 0.20 ± 0.03 a | 0.19 ± 0.02 a |
| SlGLK2-B5-1 | 6.0 ± 0.34 a | 1.04 ± 0.17 a | 5.77 ± 0.72 c | 0.16 ± 0.04 a | 0.15 ± 0.03 b |
| SlGLK2-B5-2 | 5.6 ± 0.51 a | 1.08 ± 0.31 a | 5.18 ± 0.64 c | 0.20 ± 0.03 a | 0.17 ± 0.02 a |
| SlGLK2-B5-3 | 5.8 ± 0.43 a | 1.21 ± 0.37 a | 4.79 ± 1.37 d | 0.18 ± 0.03 a | 0.16 ± 0.03 ab |
| [1] |
Adato A, Mandel T, Mintz-Oron S, Venger I, Levy D, Yativ M, Dominguez E, Wang Z, de Vos R C, Jetter R, Schreiber L, Heredia A, Rogachev I, Aharoni A. 2009. Fruit-surface flavonoid accumulation in tomato is controlled by a SlMYB12-regulated transcriptional network. PLoS Genetics, 5 (12):e1000777.
doi: 10.1371/journal.pgen.1000777 URL |
| [2] |
Ballester A R, Molthoff J, de Vos R, Hekkert B, Orzaez D, Fernandez-Moreno J P, Tripodi P, Grandillo S, Martin C, Heldens J, Ykema M, Granell A, Bovy A. 2010. Biochemical and molecular analysis of pink tomatoes:deregulated expression of the gene encoding transcription factor SlMYB 12 leads to pink tomato fruit color. Plant Physiology, 152 (1):71-84.
doi: 10.1104/pp.109.147322 pmid: 19906891 |
| [3] |
Brand A, Borovsky Y, Hill T, Rahman K A, Bellalou A, van Deynze A, Paran I. 2014. CaGLK 2 regulates natural variation of chlorophyll content and fruit color in pepper fruit. Theoretical and Applied Genetics, 127 (10):2139-2148.
doi: 10.1007/s00122-014-2367-y URL |
| [4] |
Brooks C, Nekrasov V, Lippman Z B, van Eck J. 2014. Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiology, 166 (3):1292-1297.
doi: 10.1104/pp.114.247577 pmid: 25225186 |
| [5] |
Czerednik A, Busscher M, Angenent G C, de Maagd R A. 2015. The cell size distribution of tomato fruit can be changed by overexpression of CDKA1. Plant Biotechnology Journal, 13 (2):259-268.
doi: 10.1111/pbi.12268 pmid: 25283700 |
| [6] |
Deng L, Wang H, Sun C, Li Q, Jiang H, Du M, Li C B, Li C. 2018. Efficient generation of pink-fruited tomatoes using CRISPR/Cas9 system. Journal of Genetics and Genomics, 45 (1):51-54.
doi: S1673-8527(17)30178-9 pmid: 29157799 |
| [7] |
Filler Hayut S, Melamed Bessudo C, Levy A A. 2017. Targeted recombination between homologous chromosomes for precise breeding in tomato. Nature Communications, 8:15605.
doi: 10.1038/ncomms15605 pmid: 28548094 |
| [8] |
Ilahy R, Tlili I, Siddiqui M W, Hdider C, Lenucci M S. 2019. Inside and beyond color:Comparative overview of functional quality of tomato and watermelon fruits. Frontiers in Plant Science, 10:769.
doi: 10.3389/fpls.2019.00769 URL |
| [9] |
Karkute S G, Singh A K, Gupta O P, Singh P M, Singh B. 2017. CRISPR/Cas 9 mediated genome engineering for improvement of horticultural crops. Frontiers in Plant Science, 8:1635.
doi: 10.3389/fpls.2017.01635 pmid: 28970844 |
| [10] |
Li Guobin, Ai Guo, Wei Jing, Zhang Junhong. 2021. A review of base editing technology and its application in crop genetic improvement. Acta Horticulturae Sinica, 48 (4):719-732. (in Chinese)
doi: 10.16420/j.issn.0513-353x.2020-0540 |
|
李国斌, 艾国, 韦静, 张俊红. 2021. 碱基编辑技术及其在作物遗传改良中的应用综述. 园艺学报, 48 (4):719-732.
doi: 10.16420/j.issn.0513-353x.2020-0540 |
|
| [11] |
Li J, Zhang H, Si X, Tian Y, Chen K, Liu J, Chen H, Gao C. 2017. Generation of thermosensitive male-sterile maize by targeted knockout of the ZmTMS5 gene. Journal of Genetics and Genomics, 44 (9):465-468.
doi: 10.1016/j.jgg.2017.02.002 URL |
| [12] | Li Jun-ming, Xiang Chao-yang, Wang Xiao-xuan, Guo Yan-mei, Huang Ze-jun, Liu Lei, Li Xin, Du Yong-chen. 2021. ‘Thirteenth Five-Year Plan’status and outlook of tomato industry in China. Chinese Vegetables,(2):13-20. (in Chinese) |
| 李君明, 项朝阳, 王孝宣, 国艳梅, 黄泽军, 刘磊, 李鑫, 杜永臣. 2021. “十三五”我国番茄产业现状及展望. 中国蔬菜,(2):13-20. | |
| [13] |
Lin T, Zhu G, Zhang J, Xu X, Yu Q, Zheng Z, Zhang Z, Lun Y, Li S, Wang X, Huang Z, Li J, Zhang C, Wang T, Zhang Y, Wang A, Zhang Y, Lin K, Li C, Xiong G, Xue Y, Mazzucato A, Causse M, Fei Z, Giovannoni J J, Chetelat R T, Zamir D, Stadler T, Li J, Ye Z, Du Y, Huang S. 2014. Genomic analyses provide insights into the history of tomato breeding. Nature Genetics, 46 (11):1220-1226.
doi: 10.1038/ng.3117 pmid: 25305757 |
| [14] |
Liu Y, Zhang C L, Wang X F, Li X M, You C X. 2022. Genome-wide identification and evolutionary analysis of MLO gene family in Rosaceae plants. Horticultural Plant Journal, 8 (4):395-407.
doi: 10.1016/j.hpj.2022.04.007 URL |
| [15] |
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y, Xie Y, Shen R, Chen S, Wang Z, Chen Y, Guo J, Chen L, Zhao X, Dong Z, Liu Y G. 2015. A robust CRISPR/Cas 9 system for convenient,high-efficiency multiplex genome editing in monocot and dicot plants. Molecular Plant, 8 (8):1274-1284.
doi: 10.1016/j.molp.2015.04.007 URL |
| [16] |
Nguyen C V, Vrebalov J T, Gapper N E, Zheng Y, Zhong S, Fei Z, Giovannoni J J. 2014. Tomato GOLDEN2-LIKE transcription factors reveal molecular gradients that function during fruit development and ripening. The Plant Cell, 26 (2):585-601.
doi: 10.1105/tpc.113.118794 pmid: 24510723 |
| [17] | Ouyang Bo. 2002. Study of several disease-course-associated protein genes in transformed tomatoes[Ph. D. Dissertation]. Wuhan: Huazhong Agricultural University. (in Chinese) |
| 欧阳波. 2002. 几种病程相关蛋白基因转化番茄的研究[博士论文]. 武汉: 华中农业大学. | |
| [18] |
Powell A L T, Nguyen C, Hill T, Cheng K L, Figueroa-Balderas R, Aktas H, Ashrafi H, Pons C, Fernández-Muñoz R, Vicente A, Lopez-Baltazer J, Barry C S, Liu Y S, Chetelate R, Granell A, van Deynze A, Giovannoni J J, Bennett A B. 2012. Uniform ripening encodes a Golden 2-like transcription factor regulating tomato fruit chloroplast development. Science, 336:1711-1715.
doi: 10.1126/science.1222218 pmid: 22745430 |
| [19] |
Rodriguez-Leal D, Lemmon Z H, Man J, Bartlett M E, Lippman Z B. 2017. Engineering quantitative trait variation for crop improvement by genome editing. Cell, 171 (2):470-480.
doi: 10.1016/j.cell.2017.08.030 URL |
| [20] |
Tang T, Yu X, Yang H, Gao Q, Ji H, Wang Y, Yan G, Peng Y, Luo H, Liu K, Li X, Ma C, Kang C, Dai C. 2018. Development and validation of an effective CRISPR/Cas 9 vector for efficiently isolating positive transformants and transgene-free mutants in a wide range of plant species. Frontiers in Plant Science, 9:1533.
doi: 10.3389/fpls.2018.01533 URL |
| [21] |
Voytas D F. 2013. Plant genome engineering with sequence-specific nucleases. Annual Review of Plant Biology, 64:327-350.
doi: 10.1146/annurev-arplant-042811-105552 pmid: 23451779 |
| [22] | Wang Dan, Wang Mi, Liu Jun, Zhou Xiaohui, Liu Songyu, Yang Yan, Zhuang Yong. 2022. Cloning of U6 promoters and establishment of CRISPR/Cas 9 mediated gene editing system in eggplant. Acta Horticulturae Sinica, 49 (4):791-800. (in Chinese) |
|
王丹, 王谧, 刘军, 周晓慧, 刘松瑜, 杨艳, 庄勇. 2022. 茄子U6启动子克隆及CRISPR/Cas9介导的基因编辑体系建立. 园艺学报, 49 (4):791-800.
doi: 10.16420/j.issn.0513-353x.2020-0265 |
|
| [23] |
Wang G X, Zong M, Liu D, Wu Y, Tian S W, Han S, Guo N, Duan M M, Miao LM, Liu Fan. 2022. Efficient generation of targeted point mutations in the Brassica oleracea var. botrytis genome via a modified CRISPR/Cas 9 system. Horticultural Plant Journal, 8 (4):527-530.
doi: 10.1016/j.hpj.2022.01.005 URL |
| [24] |
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu J L. 2014. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nature Biotechnology, 32 (9):947-951.
doi: 10.1038/nbt.2969 pmid: 25038773 |
| [25] |
Yu Q H, Wang B, Li N, Tang Y, Yang S, Yang T, Xu J, Guo C, Yan P, Wang Q, Asmutola P. 2017. CRISPR/Cas9-induced targeted mutagenesis and gene replacement to generate long-shelf life tomato lines. Scientific Reports, 7 (1):11874.
doi: 10.1038/s41598-017-12262-1 |
| [26] |
Zhang Aiping, Liu Jiangna, Yan Jianjun, Zhang Xiying, Bai Yunfeng. 2022. Progress and prospect of genome editing in tomato. Acta Horticulturae Sinica, 49 (1):221-232. (in Chinese)
doi: 10.16420/j.issn.0513-353x.2020-0976 |
|
张爱萍, 刘江娜, 闫建俊, 张西英, 白云凤. 2022. 番茄基因编辑研究进展和前景. 园艺学报, 49 (1):221-232.
doi: 10.16420/j.issn.0513-353x.2020-0976 |
|
| [27] |
Zhang H, Zhang J, Wei P, Zhang B, Gou F, Feng Z, Mao Y, Yang L, Zhang H, Xu N, Zhu J K. 2014. The CRISPR/Cas 9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotechnology Journal, 12 (6):797-807.
doi: 10.1111/pbi.12200 pmid: 24854982 |
| [28] |
Zhang J, Zhang H, Botella J R, Zhu J K. 2018. Generation of new glutinous rice by CRISPR_Cas9-targeted mutagenesis of the waxy gene in elite rice varieties. Journal of Integrative Plant Biology, 60:369-375.
doi: 10.1111/jipb.v60.5 URL |
| [29] |
Zuo Xin, Li Mingming, Li Xinrong, Miao Chunyan, Li Yanfang, Yang Xu, Zhang Zhongyi, Wang Fengqing. 2022. CRISPR/Cas 9 technology for RcPDS1 gene editing in Rehmannia chingii. Acta Horticulturae Sinica, 49 (7):1532-1544. (in Chinese)
doi: 10.16420/j.issn.0513-353x.2021-0475 |
|
左鑫, 李铭铭, 李欣容, 苗春妍, 李炎枋, 杨旭, 张重义, 王丰青. 2022. CRISPR/Cas9技术在天目地黄RcPDS1基因编辑中的应用. 园艺学报, 49 (7):1532-1544.
doi: 10.16420/j.issn.0513-353x.2021-0475 |
| [1] | SU Yinling, YANG Zixiang, DAN Zhong, MA Jixian, YANG Long, LI Yirong, TANG Zhengfu, WANG Lingmin, and MU Wanfu. A New Tomato Cultivar‘Yun Tomato 68’ [J]. Acta Horticulturae Sinica, 2023, 50(S1): 63-64. |
| [2] | FAN Juan, SHEN Songzhen, MIAO Qingqing, ZHANG Lehui, YAO Enpeng, and PEI Zhuoqiang . A New Tomato Cultivar‘Weihong 16’ [J]. Acta Horticulturae Sinica, 2023, 50(S1): 65-66. |
| [3] | YANG Mengxia, LIU Xiaolin, CAO Xue, WEI Kai, NING Yu, YANG Pei, LI Shanshan, CHEN Ziyue, WANG Xiaoxuan, GUO Yanmei, DU Yongchen, LI Junming, LIU Lei, LI Xin, HUANG Zejun. Construction and Application of a CRISPR/Cas9 System for Multiplex Gene Editing in Tomato [J]. Acta Horticulturae Sinica, 2023, 50(6): 1215-1229. |
| [4] | ZHANG Hui, ZHU Weimin, ZHU Longying, YANG Xuedong, ZHANG Yingying, TIAN Shoubo, WAN Yanhui, LIU Yahui, YANG Zhijie. A New Cherry Tomato Cultivar‘Huying 9’ [J]. Acta Horticulturae Sinica, 2023, 50(6): 1379-1380. |
| [5] | XU Yue, LI Zixiong, CHEN Jie, SUN Liang. Transcriptional Bases of the Regulation of 2,4-D on Tomato Fruit Shape [J]. Acta Horticulturae Sinica, 2023, 50(4): 802-814. |
| [6] | ZHENG Jinrong, NIE Jun, LI Yanhong, TAN Delong, XIE Yuming, ZHANG Changyuan. A Cherry Tomato Cultivar‘Yuekeda 101’ [J]. Acta Horticulturae Sinica, 2023, 50(4): 909-910. |
| [7] | LIU Yuhan, TAO Ning, WANG Qingguo, LI Qingqing. ABC Transporter SlABCG23 Regulates Jasmonic Acid Signaling Pathway in Tomato [J]. Acta Horticulturae Sinica, 2023, 50(3): 559-568. |
| [8] | SHI Hongli, LI La, GUO Cuimei, YU Tingting, JIAN Wei, YANG Xingyong. Isolation,Identification and Analysis of Biocontrol Ability of Biocontrol Strain TL1 Against Tomato Botrytis cinerea [J]. Acta Horticulturae Sinica, 2023, 50(1): 79-90. |
| [9] | HU Jingyu, QUE Kaijuan, MIAO Tianli, WU Shaozheng, WANG Tiantian, ZHANG Lei, DONG Xian, JI Pengzhang, DONG Jiahong. Identification of Tomato Spotted Wilt Orthotospovirus Infecting Iris tectorum [J]. Acta Horticulturae Sinica, 2023, 50(1): 170-176. |
| [10] | ZHENG Jirong, WANG Tonglin, and HU Songshen. A New Tomato Cultivar‘Hangza 603’with High Quality [J]. Acta Horticulturae Sinica, 2022, 49(S2): 103-104. |
| [11] | ZHENG Jirong and WANG Tonglin. A New Tomato Cultivar‘Hangza 601’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 105-106. |
| [12] | ZHENG Jirong and WANG Tonglin. A New Cherry Tomato Cultivar‘Hangza 503’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 107-108. |
| [13] | HUANG Tingting, LIU Shuqin, ZHANG Yongzhi, LI Ping, ZHANG Zhihuan, and SONG Libo. A New Cherry Tomato Cultivar‘Yingshahong 4’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 109-110. |
| [14] | ZHANG Qianrong, LI Dazhong, QIU Boyin, LIN Hui, MA Huifei, YE Xinru, LIU Jianting, ZHU Haisheng, and WEN Qingfang. A New Tomato Cultivar‘Minnongke 2’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 73-74. |
| [15] | HAN Shuai, WU Jie, ZHANG Heqing, XI Yadong. Identification and Sequence Analysis of Tomato Spotted Wilt Orthotospovirus Infecting Lettuce in Sichuan [J]. Acta Horticulturae Sinica, 2022, 49(9): 2007-2016. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd