
园艺学报 ›› 2021, Vol. 48 ›› Issue (2): 377-388.doi: 10.16420/j.issn.0513-353x.2020-0306
马璐琳1,2,*, 段青1,*, 崔光芬1, 杜文文1, 贾文杰1, 王祥宁1, 王继华1,**( ), 陈发棣2,**(
), 陈发棣2,**( )
)
收稿日期:2020-09-27
修回日期:2020-12-10
出版日期:2021-02-25
发布日期:2021-03-09
通讯作者:
王继华,陈发棣
E-mail:wangjh0505@sohu.com;chenfd@njau.edu.cn
基金资助:
MA Lulin1,2,*, DUAN Qing1,*, CUI Guangfen1, DU Wenwen1, JIA Wenjie1, WANG Xiangning1, WANG Jihua1,**( ), CHEN Fadi2,**(
), CHEN Fadi2,**( )
)
Received:2020-09-27
Revised:2020-12-10
Online:2021-02-25
Published:2021-03-09
Contact:
WANG Jihua,CHEN Fadi
E-mail:wangjh0505@sohu.com;chenfd@njau.edu.cn
摘要:
为了筛选适合于钝裂银莲花类黄酮/花青素合成途径中相关基因qRT-PCR表达分析时的内参基因,根据钝裂银莲花蓝/白不同花色花器官组织的转录组测序结果,选取了多聚泛素酶基因(polyubiquitin,UBQ)、微管蛋白基因(β-tubulin,β-TUB)、水通道蛋白基因(aquaporin,AQP)、肌动蛋白基因(actin,ACT)、甘油醛-3-磷酸-脱氢酶基因(glyceraldehyde-3-phosphate dehydrogenase,GAPDH)、组蛋白基因(histone,HIS)、转录延伸因子基因(elongation factor 1-β,EF-1β)和60S核糖体蛋白基因(60S ribosomal protein L13-1,RPL13)等8个常用内参基因作为候选基因,以钝裂银莲花的叶片、茎杆和蓝/白色花器官等不同组织为试验材料,通过qRT-PCR检测这8个候选内参基因的表达情况,利用geNorm、NormFinder和BestKeeper等软件对其稳定性进行分析评价。结果表明:8个候选内参基因中,UBQ表现最稳定,而β-TUB相对稳定性最差。以最稳定的UBQ为内参对钝裂银莲花类黄酮/花青素合成途径中16个相关基因表达情况进行qRT-PCR分析,结果与前期转录组测序结果一致。UBQ为钝裂银莲花花色素合成途径相关基因表达分析的最适内参基因。
中图分类号:
马璐琳, 段青, 崔光芬, 杜文文, 贾文杰, 王祥宁, 王继华, 陈发棣. 钝裂银莲花花色素合成相关基因qRT-PCR内参基因的筛选[J]. 园艺学报, 2021, 48(2): 377-388.
MA Lulin, DUAN Qing, CUI Guangfen, DU Wenwen, JIA Wenjie, WANG Xiangning, WANG Jihua, CHEN Fadi. Selection and Validation of Reference Genes for qRT-PCR Analysis of the Correlated Genes in Flower Pigments Biosynthesis Pathway of Anemone obtusiloba[J]. Acta Horticulturae Sinica, 2021, 48(2): 377-388.
| 基因ID Gene ID | FPKM | 比对结果 Blast result | 基因 Gene | |||||
|---|---|---|---|---|---|---|---|---|
| 蓝花 Blue flower | 白花 White flower | |||||||
| 1 | 2 | 3 | 1 | 2 | 3 | |||
| DN35855_c0_g1_i10_1 | 354.26 | 385.74 | 422.52 | 336.35 | 354.00 | 390.89 | Glyceraldehyde 3-phosphate dehydrogenase [Gynura bicolor] (GenBank:AB550243.1) | GAPDH |
| DN24449_c0_g1_i2_1 | 314.43 | 331.50 | 366.20 | 272.08 | 300.62 | 327.47 | Elongation factor 1-beta 2-like[Pistacia vera](NCBI reference sequence:XM_031416658.1) | EF-1β |
| DN38564_c0_g2_i1_1 | 434.91 | 450.67 | 464.33 | 473.54 | 517.63 | 522.32 | Actin 7 [Prunus dulcis] (GenBank:BBH05534.1) | ACT |
| DN25341_c0_g2_i1_2 | 153.63 | 173.57 | 173.87 | 143.18 | 160.60 | 185.16 | Tubulin beta chain[Ricinus communis] (NCBI reference sequence: XM_025156136.1) | β-TUB |
| DN24929_c0_g1_i4_2 | 1 016.78 | 975.95 | 933.45 | 1 206.08 | 1 170.53 | 1 067.26 | Polyubiquitin,partial [Wolffia australiana](GenBank:AEZ49160.1) | UBQ |
| DN34836_c0_g2_i6_2 | 1 052.09 | 1 172.91 | 1 294.86 | 1 140.09 | 1 193.46 | 1 301.07 | Aquaporin TIP1-2[Brassica rapa] (NCBI reference sequence: XP_009129499.) | AQP |
| DN24035_c0_g1_i2_1 | 131.24 | 122.16 | 141.00 | 106.95 | 108.04 | 119.61 | Histone H2A variant 1[Phalaenopsis equestris](NCBI reference sequence:XP_020573071.1) | HIS |
| DN25248_c0_g2_i7_1 | 197.63 | 210.20 | 236.21 | 157.42 | 172.19 | 188.42 | 60S ribosomal protein L13-1 [Camellia sinensis](NCBI reference sequence:XP_028086825.1) | RPL13 |
表1 根据钝裂银莲花转录组测序结果筛选到的8个候选内参基因
Table 1 Eight candidate reference genes in RNA-seq libraries of Anemone obtusiloba
| 基因ID Gene ID | FPKM | 比对结果 Blast result | 基因 Gene | |||||
|---|---|---|---|---|---|---|---|---|
| 蓝花 Blue flower | 白花 White flower | |||||||
| 1 | 2 | 3 | 1 | 2 | 3 | |||
| DN35855_c0_g1_i10_1 | 354.26 | 385.74 | 422.52 | 336.35 | 354.00 | 390.89 | Glyceraldehyde 3-phosphate dehydrogenase [Gynura bicolor] (GenBank:AB550243.1) | GAPDH |
| DN24449_c0_g1_i2_1 | 314.43 | 331.50 | 366.20 | 272.08 | 300.62 | 327.47 | Elongation factor 1-beta 2-like[Pistacia vera](NCBI reference sequence:XM_031416658.1) | EF-1β |
| DN38564_c0_g2_i1_1 | 434.91 | 450.67 | 464.33 | 473.54 | 517.63 | 522.32 | Actin 7 [Prunus dulcis] (GenBank:BBH05534.1) | ACT |
| DN25341_c0_g2_i1_2 | 153.63 | 173.57 | 173.87 | 143.18 | 160.60 | 185.16 | Tubulin beta chain[Ricinus communis] (NCBI reference sequence: XM_025156136.1) | β-TUB |
| DN24929_c0_g1_i4_2 | 1 016.78 | 975.95 | 933.45 | 1 206.08 | 1 170.53 | 1 067.26 | Polyubiquitin,partial [Wolffia australiana](GenBank:AEZ49160.1) | UBQ |
| DN34836_c0_g2_i6_2 | 1 052.09 | 1 172.91 | 1 294.86 | 1 140.09 | 1 193.46 | 1 301.07 | Aquaporin TIP1-2[Brassica rapa] (NCBI reference sequence: XP_009129499.) | AQP |
| DN24035_c0_g1_i2_1 | 131.24 | 122.16 | 141.00 | 106.95 | 108.04 | 119.61 | Histone H2A variant 1[Phalaenopsis equestris](NCBI reference sequence:XP_020573071.1) | HIS |
| DN25248_c0_g2_i7_1 | 197.63 | 210.20 | 236.21 | 157.42 | 172.19 | 188.42 | 60S ribosomal protein L13-1 [Camellia sinensis](NCBI reference sequence:XP_028086825.1) | RPL13 |
| KEGG途径 KEGG pathway | 基因表达情况(白花为对照) Gene expression(White flower was the control group) | 基因ID Gene ID | FPKM | |||||
|---|---|---|---|---|---|---|---|---|
| 蓝花 Blue flower | 白花 White flower | |||||||
| 1 | 2 | 3 | 1 | 2 | 3 | |||
| 类黄酮生 物合成 Flavonoid biosynthetic | 上调Up | DN26579_c0_g1_i3_2 | 569.86 | 469.94 | 406.45 | 109.46 | 98.07 | 113.44 |
| 上调Up | DN27520_c0_g1_i2_2 | 341.41 | 278.90 | 233.47 | 38.08 | 29.97 | 27.08 | |
| 上调Up | DN27888_c0_g1_i1_1 | 1 023.47 | 839.09 | 694.98 | 89.89 | 74.08 | 65.41 | |
| 上调Up | DN34928_c0_g1_i4_1 | 237.71 | 182.14 | 158.62 | 19.84 | 21.50 | 18.22 | |
| 上调Up | DN37635_c0_g1_i7_1 | 390.62 | 309.90 | 252.52 | 27.24 | 22.26 | 25.06 | |
| 无差异No-difference | DN29460_c0_g1_i3_1 | 1.66 | 2.73 | 1.54 | 1.08 | 0.85 | 0.93 | |
| 无差异No-difference | DN29383_c0_g1_i2_1 | 16.90 | 15.95 | 20.66 | 9.54 | 11.30 | 13.71 | |
| 无差异No-difference | DN32877_c0_g1_i3_2 | 64.45 | 55.33 | 49.30 | 53.18 | 40.37 | 68.46 | |
| 无差异No-difference | DN32877_c0_g2_i5_2 | 132.45 | 137.58 | 136.64 | 119.78 | 124.61 | 149.95 | |
| 无差异No-difference | DN33447_c0_g1_i2_2 | 99.69 | 101.66 | 84.61 | 93.18 | 87.51 | 108.33 | |
| 无差异No-difference | DN34032_c0_g1_i1_1 | 100.37 | 101.35 | 92.38 | 92.69 | 86.81 | 93.82 | |
| 无差异No-difference | DN34871_c2_g3_i1_2 | 34.07 | 41.26 | 40.60 | 34.36 | 36.08 | 45.69 | |
| 无差异No-difference | DN35887_c0_g1_i1_2 | 133.36 | 129.04 | 121.26 | 96.60 | 102.29 | 115.96 | |
| 无差异No-difference | DN36570_c0_g2_i4_1 | 81.57 | 91.62 | 87.44 | 81.32 | 80.35 | 109.16 | |
| 花青素生 物合成 Anthocyanin biosynthetic | 上调Up | DN36200_c0_g1_i3_1 | 589.72 | 479.41 | 394.60 | 57.97 | 47.95 | 45.82 |
| 无差异No-difference | DN35376_c0_g1_i4_2 | 31.79 | 31.73 | 30.26 | 37.08 | 37.12 | 34.80 | |
表2 钝裂银莲花转录组测序数据中的类黄酮/花青素合成途径中16个相关基因
Table 2 Some genes in the flavonoids/anthocyanins pathways of Anemone obtusiloba
| KEGG途径 KEGG pathway | 基因表达情况(白花为对照) Gene expression(White flower was the control group) | 基因ID Gene ID | FPKM | |||||
|---|---|---|---|---|---|---|---|---|
| 蓝花 Blue flower | 白花 White flower | |||||||
| 1 | 2 | 3 | 1 | 2 | 3 | |||
| 类黄酮生 物合成 Flavonoid biosynthetic | 上调Up | DN26579_c0_g1_i3_2 | 569.86 | 469.94 | 406.45 | 109.46 | 98.07 | 113.44 |
| 上调Up | DN27520_c0_g1_i2_2 | 341.41 | 278.90 | 233.47 | 38.08 | 29.97 | 27.08 | |
| 上调Up | DN27888_c0_g1_i1_1 | 1 023.47 | 839.09 | 694.98 | 89.89 | 74.08 | 65.41 | |
| 上调Up | DN34928_c0_g1_i4_1 | 237.71 | 182.14 | 158.62 | 19.84 | 21.50 | 18.22 | |
| 上调Up | DN37635_c0_g1_i7_1 | 390.62 | 309.90 | 252.52 | 27.24 | 22.26 | 25.06 | |
| 无差异No-difference | DN29460_c0_g1_i3_1 | 1.66 | 2.73 | 1.54 | 1.08 | 0.85 | 0.93 | |
| 无差异No-difference | DN29383_c0_g1_i2_1 | 16.90 | 15.95 | 20.66 | 9.54 | 11.30 | 13.71 | |
| 无差异No-difference | DN32877_c0_g1_i3_2 | 64.45 | 55.33 | 49.30 | 53.18 | 40.37 | 68.46 | |
| 无差异No-difference | DN32877_c0_g2_i5_2 | 132.45 | 137.58 | 136.64 | 119.78 | 124.61 | 149.95 | |
| 无差异No-difference | DN33447_c0_g1_i2_2 | 99.69 | 101.66 | 84.61 | 93.18 | 87.51 | 108.33 | |
| 无差异No-difference | DN34032_c0_g1_i1_1 | 100.37 | 101.35 | 92.38 | 92.69 | 86.81 | 93.82 | |
| 无差异No-difference | DN34871_c2_g3_i1_2 | 34.07 | 41.26 | 40.60 | 34.36 | 36.08 | 45.69 | |
| 无差异No-difference | DN35887_c0_g1_i1_2 | 133.36 | 129.04 | 121.26 | 96.60 | 102.29 | 115.96 | |
| 无差异No-difference | DN36570_c0_g2_i4_1 | 81.57 | 91.62 | 87.44 | 81.32 | 80.35 | 109.16 | |
| 花青素生 物合成 Anthocyanin biosynthetic | 上调Up | DN36200_c0_g1_i3_1 | 589.72 | 479.41 | 394.60 | 57.97 | 47.95 | 45.82 |
| 无差异No-difference | DN35376_c0_g1_i4_2 | 31.79 | 31.73 | 30.26 | 37.08 | 37.12 | 34.80 | |
| 用途 Purpose | 基因 Gene | 引物序列(5′-3′) Primer sequence | 产物长度/bp Product length |
|---|---|---|---|
| 内参基因 Reference gene | GAPDH | F:ATGGGTGTGAATGAGAAGGA;R:CGTGGACGGTTGTCATAAG | 142 |
| EF1-β2 | F:GAAAGAGAGTGGCAAATCCT;R:GCATCTCGATGCTGCGAA | 101 | |
| ACT | F:GCCGTGCTTTCTCTCTATG;R:TTGGGACTGTGTGGCTAAC | 82 | |
| β-TUB | F:ATATCCCACCAACTGGACT;R:ACTGTTCTGAAACTCTGCG | 90 | |
| UBQ | F:CTTTGCCGGGAAACAATTAG;R:GACGGAGCACCAAGTGAA | 86 | |
| AQP | F:CCGGCGGAAAGTCTTATAAT;R:AATGTTTCCACCGAGAAGAG | 135 | |
| HIS | F:AGCTTGATACACTGATCAAAGG;R:GGTAGACTTGTTGATGAGTGAC | 83 | |
| RPL13 | F:TGCAGTTACTCAGCTCACAG;R:CTTTGAAGGCTTTCATCTCCTC | 98 | |
| 鉴定基因 Identification gene | DN26579_c0_g1_i3_2 | F:AAGACGTGGATCACGGTTA;R:CTTGAACCTCCCATTGCT | 90 |
| DN27520_c0_g1_i2_2 | F:GCACCGAAGAGGAATGCAA;R:GTCGATAACTGGAATCTGCG | 86 | |
| DN27888_c0_g1_i1_1 | F:GTGGTGGCGGAGATGTTA;R:CCTTTGGTGAGAAACACTCG | 83 | |
| DN34928_c0_g1_i4_1 | F:CCTAGTGACAAAGGTGCTG;R:TTAGTAAGTCGTCCATGCCG | 95 | |
| DN37635_c0_g1_i7_1 | F:ATGCCTCCTAGCCTGCTAA;R:TACCAGTTGAACCTGTTTCAGA | 84 | |
| DN29460_c0_g1_i3_1 | F:ATCAACCTCTCCCTTATAGACC;R:TTTCGTTCTTCTTCCCTGAGT | 82 | |
| DN29383_c0_g1_i2_1 | F:ACCAAGATCAAGAATGGCG;R:TTGATATTTGTGTCGGTGAGC | 122 | |
| DN32877_c0_g1_i3_2 | F:GAACGGATCTGTTGTCGC;R:GGTAGCTGGCAAATCTCAAT | 126 | |
| DN32877_c0_g2_i5_2 | F:TGGCAATGGACATAAACCG;R:CTTTGAACTCAATCTTGTGACC | 87 | |
| DN33447_c0_g1_i2_2 | F:TGACCACTGGTACAACGC;R:GACTGTAGTCCGATGAAACAC | 115 | |
| DN34032_c0_g1_i1_1 | F:GGAGGAGGAATCATTTGCC;R:TCAACGAAATGTTGCTTAGCG | 117 | |
| DN34871_c2_g3_i1_2 | F:ACCAAGATCAAGAATGGCG;R:TTGATATTTGTGTCGGTGAGC | 122 | |
| DN35887_c0_g1_i1_2 | F:CACCATGATCTTGCCATTAAC;R:CATTAGCTTTCCAGTAGGCG | 80 | |
| DN36570_c0_g2_i4_1 | F:GAACGGATCTGTTGTCGC;R:AGCCAAAGCCTTGTTAAGT | 94 | |
| DN36200_c0_g1_i3_1 | F:CAGGAATGTCATCTGTACGG;R:ATGCAACAAGTTACCCAAGTAG | 80 | |
| DN35376_c0_g1_i4_2 | F:TGCTTCTTCCCTCACCTG;R:TTCTCGTGGCAAATCCCG | 130 |
表3 引物列表
Table 3 Primers in this study
| 用途 Purpose | 基因 Gene | 引物序列(5′-3′) Primer sequence | 产物长度/bp Product length |
|---|---|---|---|
| 内参基因 Reference gene | GAPDH | F:ATGGGTGTGAATGAGAAGGA;R:CGTGGACGGTTGTCATAAG | 142 |
| EF1-β2 | F:GAAAGAGAGTGGCAAATCCT;R:GCATCTCGATGCTGCGAA | 101 | |
| ACT | F:GCCGTGCTTTCTCTCTATG;R:TTGGGACTGTGTGGCTAAC | 82 | |
| β-TUB | F:ATATCCCACCAACTGGACT;R:ACTGTTCTGAAACTCTGCG | 90 | |
| UBQ | F:CTTTGCCGGGAAACAATTAG;R:GACGGAGCACCAAGTGAA | 86 | |
| AQP | F:CCGGCGGAAAGTCTTATAAT;R:AATGTTTCCACCGAGAAGAG | 135 | |
| HIS | F:AGCTTGATACACTGATCAAAGG;R:GGTAGACTTGTTGATGAGTGAC | 83 | |
| RPL13 | F:TGCAGTTACTCAGCTCACAG;R:CTTTGAAGGCTTTCATCTCCTC | 98 | |
| 鉴定基因 Identification gene | DN26579_c0_g1_i3_2 | F:AAGACGTGGATCACGGTTA;R:CTTGAACCTCCCATTGCT | 90 |
| DN27520_c0_g1_i2_2 | F:GCACCGAAGAGGAATGCAA;R:GTCGATAACTGGAATCTGCG | 86 | |
| DN27888_c0_g1_i1_1 | F:GTGGTGGCGGAGATGTTA;R:CCTTTGGTGAGAAACACTCG | 83 | |
| DN34928_c0_g1_i4_1 | F:CCTAGTGACAAAGGTGCTG;R:TTAGTAAGTCGTCCATGCCG | 95 | |
| DN37635_c0_g1_i7_1 | F:ATGCCTCCTAGCCTGCTAA;R:TACCAGTTGAACCTGTTTCAGA | 84 | |
| DN29460_c0_g1_i3_1 | F:ATCAACCTCTCCCTTATAGACC;R:TTTCGTTCTTCTTCCCTGAGT | 82 | |
| DN29383_c0_g1_i2_1 | F:ACCAAGATCAAGAATGGCG;R:TTGATATTTGTGTCGGTGAGC | 122 | |
| DN32877_c0_g1_i3_2 | F:GAACGGATCTGTTGTCGC;R:GGTAGCTGGCAAATCTCAAT | 126 | |
| DN32877_c0_g2_i5_2 | F:TGGCAATGGACATAAACCG;R:CTTTGAACTCAATCTTGTGACC | 87 | |
| DN33447_c0_g1_i2_2 | F:TGACCACTGGTACAACGC;R:GACTGTAGTCCGATGAAACAC | 115 | |
| DN34032_c0_g1_i1_1 | F:GGAGGAGGAATCATTTGCC;R:TCAACGAAATGTTGCTTAGCG | 117 | |
| DN34871_c2_g3_i1_2 | F:ACCAAGATCAAGAATGGCG;R:TTGATATTTGTGTCGGTGAGC | 122 | |
| DN35887_c0_g1_i1_2 | F:CACCATGATCTTGCCATTAAC;R:CATTAGCTTTCCAGTAGGCG | 80 | |
| DN36570_c0_g2_i4_1 | F:GAACGGATCTGTTGTCGC;R:AGCCAAAGCCTTGTTAAGT | 94 | |
| DN36200_c0_g1_i3_1 | F:CAGGAATGTCATCTGTACGG;R:ATGCAACAAGTTACCCAAGTAG | 80 | |
| DN35376_c0_g1_i4_2 | F:TGCTTCTTCCCTCACCTG;R:TTCTCGTGGCAAATCCCG | 130 |
| 内参基因 Reference gene | 最大值 Maximum value | 最小值 Minimum value | 平均值 Mean | 标准差 SD | 变异系数/% CV |
|---|---|---|---|---|---|
| GAPDH | 22.86 | 21.81 | 22.315 | 0.381 | 1.709 |
| EF1-β2 | 31.99 | 30.38 | 31.269 | 0.614 | 1.963 |
| ACT | 21.77 | 20.60 | 21.075 | 0.349 | 1.658 |
| β-TUB | 27.02 | 23.03 | 24.289 | 1.523 | 6.270 |
| UBQ | 22.96 | 21.51 | 22.184 | 0.586 | 2.641 |
| AQP | 22.06 | 20.51 | 21.150 | 0.599 | 2.831 |
| HIS | 27.82 | 26.03 | 27.006 | 0.714 | 2.645 |
| RPL13 | 25.66 | 23.04 | 24.303 | 1.091 | 4.490 |
表4 候选内参基因Ct值分析
Table 4 Analysis of the Ct values of candidate reference genes
| 内参基因 Reference gene | 最大值 Maximum value | 最小值 Minimum value | 平均值 Mean | 标准差 SD | 变异系数/% CV |
|---|---|---|---|---|---|
| GAPDH | 22.86 | 21.81 | 22.315 | 0.381 | 1.709 |
| EF1-β2 | 31.99 | 30.38 | 31.269 | 0.614 | 1.963 |
| ACT | 21.77 | 20.60 | 21.075 | 0.349 | 1.658 |
| β-TUB | 27.02 | 23.03 | 24.289 | 1.523 | 6.270 |
| UBQ | 22.96 | 21.51 | 22.184 | 0.586 | 2.641 |
| AQP | 22.06 | 20.51 | 21.150 | 0.599 | 2.831 |
| HIS | 27.82 | 26.03 | 27.006 | 0.714 | 2.645 |
| RPL13 | 25.66 | 23.04 | 24.303 | 1.091 | 4.490 |
| 排序 Ranking | geNorm | NormFinder | BestKeeper | |||||
|---|---|---|---|---|---|---|---|---|
| 内参基因 Reference gene | M值 M | 内参基因 Reference gene | 稳定值 SV | 内参基因 Reference genes | 标准差 SD | |||
| 1 | UBQ | 0.413 | UBQ | 0.068 | ACT | 0.282 | ||
| 2 | EF1-β2 | 0.425 | AQP | 0.071 | GAPDH | 0.356 | ||
| 3 | AQP | 0.441 | EF1-β2 | 0.113 | AQP | 0.531 | ||
| 4 | HIS | 0.460 | HIS | 0.160 | UBQ | 0.567 | ||
| 5 | GAPDH | 0.535 | ACT | 0.287 | EF1-β2 | 0.574 | ||
| 6 | ACT | 0.543 | GAPDH | 0.311 | HIS | 0.678 | ||
| 7 | RPL13 | 0.644 | RPL13 | 0.325 | RPL13 | 1.055 | ||
| 8 | β-TUB | 1.128 | β-TUB | 0.784 | β-TUB | 1.283 | ||
表5 候选内参基因稳定性分析
Table 5 Stability analysis of candidate reference genes
| 排序 Ranking | geNorm | NormFinder | BestKeeper | |||||
|---|---|---|---|---|---|---|---|---|
| 内参基因 Reference gene | M值 M | 内参基因 Reference gene | 稳定值 SV | 内参基因 Reference genes | 标准差 SD | |||
| 1 | UBQ | 0.413 | UBQ | 0.068 | ACT | 0.282 | ||
| 2 | EF1-β2 | 0.425 | AQP | 0.071 | GAPDH | 0.356 | ||
| 3 | AQP | 0.441 | EF1-β2 | 0.113 | AQP | 0.531 | ||
| 4 | HIS | 0.460 | HIS | 0.160 | UBQ | 0.567 | ||
| 5 | GAPDH | 0.535 | ACT | 0.287 | EF1-β2 | 0.574 | ||
| 6 | ACT | 0.543 | GAPDH | 0.311 | HIS | 0.678 | ||
| 7 | RPL13 | 0.644 | RPL13 | 0.325 | RPL13 | 1.055 | ||
| 8 | β-TUB | 1.128 | β-TUB | 0.784 | β-TUB | 1.283 | ||
| 排序 Ranking | 内参基因 Reference gene | 几何平均数 Geomean of ranking values |
|---|---|---|
| 1 | UBQ | 1.587 |
| 2 | AQP | 2.621 |
| 3 | EF1-β2 | 3.107 |
| 3 | GAPDH | 3.107 |
| 5 | ACT | 3.915 |
| 6 | HIS | 4.579 |
| 7 | RPL13 | 7.000 |
| 8 | β-TUB | 8.000 |
表6 候选内参稳定性综合排序分析
Table 6 The comprehensive ranking of stability of candidate reference genes
| 排序 Ranking | 内参基因 Reference gene | 几何平均数 Geomean of ranking values |
|---|---|---|
| 1 | UBQ | 1.587 |
| 2 | AQP | 2.621 |
| 3 | EF1-β2 | 3.107 |
| 3 | GAPDH | 3.107 |
| 5 | ACT | 3.915 |
| 6 | HIS | 4.579 |
| 7 | RPL13 | 7.000 |
| 8 | β-TUB | 8.000 |
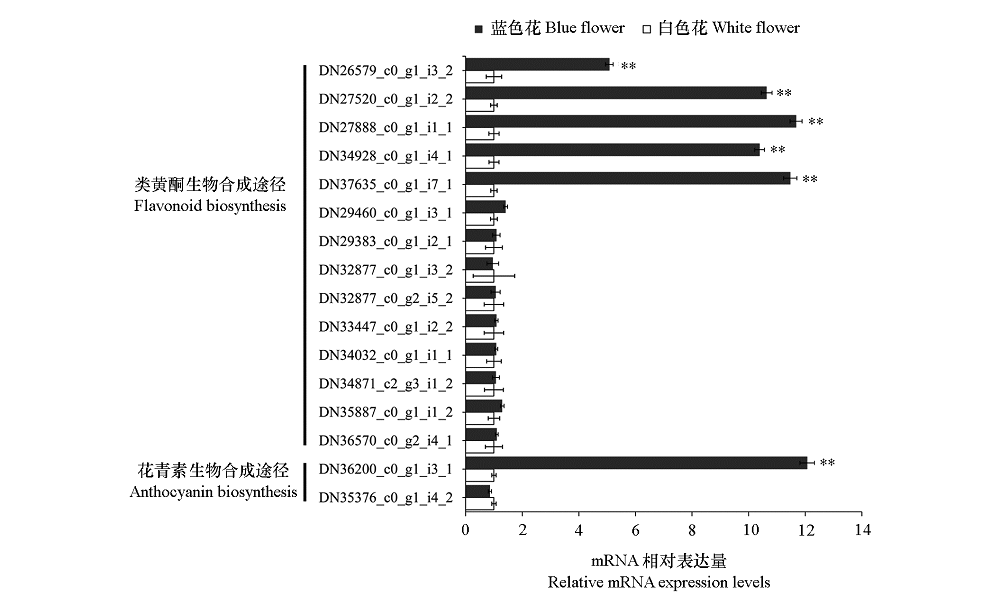
图2 钝裂银莲花类黄酮/花青素合成途径中相关基因的qRT-PCR表达结果
Fig. 2 The qRT-PCR expression pattern of the pigments synthesis related genes in the flavonoids/anthocyanins pathways of Anemone obtusiloba α = 0.01.
| [1] |
Ahn J H, Kim J S, Kim S, Soh H Y, Shin H, Jang H, Ryu J H, Kim A, Yun K Y, Kim S, Kim K S, Choi D, Huh J H. 2015. De novo transcriptome analysis to identify anthocyanin biosynthesis genes responsible for tissue-specific pigmentation in zoysiagrass(Zoysia japonica Steud.). PLoS ONE, 10(4):e0124497
doi: 10.1371/journal.pone.0124497 URL |
| [2] |
Bustin S A. 2002. Quantifification of mRNA using real-time reverse transcription PCR(RT-PCR):trends and problems. Journal of Molecular Endocrinology, 29(1):23-39.
doi: 10.1677/jme.0.0290023 URL |
| [3] |
Chen C B, Wu J Y, Hua Q Z, Tel-Zur N, Xie F F, Zhang Z K, Chen J Y, Zhang R, Hu G B, Zhao J T, Qin Y H. 2019. Identifcation of reliable reference genes for quantitative real-time PCR normalization in pitaya. Plant Methods, 15:70.
doi: 10.1186/s13007-019-0455-3 URL |
| [4] |
Chen D L, Pan X P, Xiao P, Farwell M A, Zhang B H. 2011. Evaluation and identification of reliable reference genes for pharmacogenomics,toxicogenomics,and small RNA expression analysis. Journal of Cellular Physiology, 226(10):2469-2477.
doi: 10.1002/jcp.v226.10 URL |
| [5] |
Debashree L R, Joshua C J. 2014. Validation of reference genes for gene expression analysis in olive(Olea europaea)mesocarp tissue by quantitative real-time RT-PCR. BMC Research Notes, 7:304.
doi: 10.1186/1756-0500-7-304 pmid: 24884716 |
| [6] | Delectis Florae Reipublicae Popularis Sinicae Agendae Academiae Sinicae Edita. 1980. Flora reipublicae popularis sincae. Beijing: Science Press. (in Chinese) |
| 中国科学院中国动植物志编委会. 1980. 中国植物志. 北京: 科学出版社. | |
| [7] |
Forkmann G. 1991. Flavonoids as flower pigments:the formation of the natural spectrum and its extension by genetic engineering. Plant Breeding, 106(1):1-26.
doi: 10.1111/pbr.1991.106.issue-1 URL |
| [8] | Fu Yuan-yuan, Mu Chun-sheng, Gao Hong-wen, Li Jun, Wang Xue-min. 2014. Cloning of 18S rRNA gene and stability evaluation of reference genes in Medicago sativa. Plant Physiology Journal, 50(12):1809-1815. (in Chinese) |
| 付媛媛, 穆春生, 高洪文, 李俊, 王学敏. 2014. 紫花苜蓿18S rRNA基因的克隆及内参基因表达稳定性评价. 植物生理学报, 50(12):1809-1815. | |
| [9] | Hu Chun, Liu Zuo-jun, Li Fu-xiang, Liu Feng-mei. 2013. Detection Methods for pollen viability of Anemone obtusiloba. Bulletin of Botanical Research, 33(5):582-586. |
| 胡春, 刘左军, 李富香, 刘凤梅. 2013. 钝裂银莲花花粉活力测定方法的研究. 植物研究, 33(5):582-586. (in Chinese) | |
| [10] |
Huggett J, Dheda K, Bustin S, Zumla A. 2005. Real-time RT-PCR normalisation;strategies and considerations. Genes and Immunity, 6(4):279-284.
doi: 10.1038/sj.gene.6364190 URL |
| [11] |
Hyun T K, Rim Y, Jang H J, Kim C H, Park J, Kumar R, Lee S, Kim B C, Bhak J, Nguyen-Quoc B, Kim S W, Lee S Y, Kim J Y. 2012. De novo transcriptome sequencing of Momordica cochinchinensis to identify genes involved in the carotenoid biosynthesis. Plant Molecular Biology, 79(4/5):413-427.
doi: 10.1007/s11103-012-9919-9 URL |
| [12] | Jiang Ting-ting, Gao Yan-hui, Tong Zai-kang. 2015. Selection of reference genes for quantitative real-time PCR in Lycoris. Acta Horticulturae Sinica, 42(6):1129-1138. (in Chinese) |
| 蒋婷婷, 高燕会, 童再康. 2015. 石蒜属植物实时荧光定量PCR内参基因的选择. 园艺学报, 42(6):1129-1138. | |
| [13] |
Kou X Y, Zhang L, Yang S Z, Li G H, Ye J L. 2017. Selection and validation of reference genes for quantitative RT-PCR analysis in peach fruit under different experimental conditions. Scientia Horticulturae, 225(1):195-203.
doi: 10.1016/j.scienta.2017.07.004 URL |
| [14] |
Kriangphan N, Vuttipongchaikij S, Kittiwongwattana C, Suttangkakul A, Pinmanee P, Sakulsathaporn A, Suwimon R, Suputtitada S, Chanvivattana Y, Apisitwanich S. 2015. Effects of sequence and expression of eight anthocyanin biosynthesis genes on floral coloration in four Dendrobium hybrids. The Horticulture Journal, 84(1):83-92.
doi: 10.2503/hortj.MI-020 URL |
| [15] | Liu Chuan-jiao, Wang Shun-li, Xue Jing-qi, Zhu Fu-yong, Ren Xiu-xia, Li Ming-yang, Zhang Xiu-xin. 2015. Molecular cloning of ubiquitin protein gene and study on this gene as reference gene in tree peony. Acta Horticulturae Sinica, 42(10):1983-1992. (in Chinese) |
| 刘传娇, 王顺利, 薛璟祺, 朱富勇, 任秀霞, 李名扬, 张秀新. 2015. 牡丹泛素延伸蛋白基因ubiquitin的克隆及其作为内参基因的研究. 园艺学报, 42(10):1983-1992. | |
| [16] |
Liu Q X, Qi X, Yan H D, Huang L K, Nie G, Zhang X Q. 2018. Reference gene selection for quantitative real-time reverse-transcriptase PCR in annual ryegrass(Lolium multiflorum)subjected to various abiotic stresses. Molecules, 23(1):172-184.
doi: 10.3390/molecules23010172 URL |
| [17] | Liu Xiao-ting, Wang Shun-li, Xue Jing-qi, Xue Yu-qian, Lü Ying-min, Zhang Xiu-xin. 2018. Selection of reference genes for quantitative real-time PCR in different tissue and organ of barbadoslily. Acta Horticulturae Sinica, 45(5):919-930. (in Chinese) |
| 刘晓婷, 王顺利, 薛璟祺, 薛玉前, 吕英民, 张秀新. 2018. 朱顶红实时荧光定量PCR中不同组织器官内参基因的筛选. 园艺学报, 45(5):919-930. | |
| [18] |
Lou Q, Liu Y L, Qi Y Y, Jiao S Z, Tian F F, Jiang L, Wang Y J. 2014. Transcriptome sequencing and metabolite analysis reveals the role of delphinidin metabolism in flower colour in grape hyacinth. Journal of Experimental Botany, 65(12):3157-3164.
doi: 10.1093/jxb/eru168 URL |
| [19] | Ma Lu-lin, Cui Guang-fen, Wang Xiang-ning, Jia Wen-jie, Duan Qing, Du Wen-wen, Wang Ji-hua. 2017. Cloning and expression analysis of catalase(Ls-Cat1)gene in Lilium sargentiae Wilson. Journal of Nuclear Agricultural Sciences, 31(9):1700-1707. (in Chinese) |
| 马璐琳, 崔光芬, 王祥宁, 贾文杰, 段青, 杜文文, 王继华. 2017. 泸定百合过氧化氢酶(Ls-Cat1)基因的克隆及表达分析. 核农学报, 31(9):1700-1707. | |
| [20] | Ma Lu-lin, Cui Guang-fen, Wang Xiang-ning, Jia Wen-jie, Duan Qing, Du Wen-wen, Wang Ji-hua, Chen Fa-di. 2019. Selection and validation of reference genes for quantitative real-time PCR analysis in Iris bulleyana during flower color variation. Journal of Nuclear Agricultural Sciences, 33(9):1707-1716. (in Chinese) |
| 马璐琳, 崔光芬, 王祥宁, 贾文杰, 段青, 杜文文, 王继华, 陈发棣. 2019. 西南鸢尾花色变异实时定量PCR内参基因的筛选与验证. 核农学报, 33(9):1707-1716. | |
| [21] |
Ma P Y, Bian X F, Jia Z D, Guo X D, Xie Y Z. 2016. De novo sequencing and comprehensive analysis of the mutant transcriptome from purple sweet potato(Ipomoea batatas L.). Gene, 575(3):641-649.
doi: 10.1016/j.gene.2015.09.056 URL |
| [22] |
Nolan T, Hands R E, Bustin S A. 2006. Quantifification of mRNA using real-time RT-PCR. Nature protocols, 1(3):1559-1582.
doi: 10.1038/nprot.2006.236 URL |
| [23] | Qiao Yong-gang, Wang Yong-fei, Cao Ya-ping, He Jia-xin, Jia Meng-jun, Li Zheng, Zhang Xin-rui, Song Yun. 2020. Reference genes selection and related genes expression analysis under low and high temperature stress in Taraxacum officinale. Acta Horticulturae Sinica, 47(6):1153-1164. (in Chinese) |
| 乔永刚, 王勇飞, 曹亚萍, 贺嘉欣, 贾孟君, 李政, 张鑫瑞, 宋芸. 2020. 药用蒲公英低温和高温胁迫下内参基因筛选与相关基因表达分析. 园艺学报, 47(6):1153-1164. | |
| [24] |
Quackenbush J. 2002. Microarray data normalization and transformation. nature genetics, 32:496-501.
doi: 10.1038/ng1032 URL |
| [25] |
Song S Y, Seo P J, Yang M S, Xiang F N, Park C M. 2008. Exploring valid reference genes for gene expression studies in Brachypodium distachyon by real-time PCR. Bmc Plant Biology, 8(1):112-132.
doi: 10.1186/1471-2229-8-112 URL |
| [26] | Song Xiao-bo, Chang Ying-ying, Liu Hao, Xu Hui-min, Pei Dong. 2019. Reference gene selection and genes expression analysis during adventitious root formation in walnut. Acta Horticulturae Sinica, 46(10):1907-1918. (in Chinese) |
| 宋晓波, 常英英, 刘昊, 徐慧敏, 裴东. 2019. 核桃不定根发生阶段内参基因筛选与关键基因表达分析. 园艺学报, 46(10):1907-1918. | |
| [27] | Sun Mei-lian, Wang Yun-sheng, Yang Dong-qing, Wei Chao-ling, Gao Li-ping, Xia Tao, Shan Yu, Luo Yang. 2010. Reference genes for real-time fluorescence quantitative PCR in Camellia sinensis. Chinese Bulletin of Botany, 45(5):579-587. (in Chinese) |
| 孙美莲, 王云生, 杨冬青, 韦朝领, 高丽萍, 夏涛, 单育, 骆洋. 2010. 茶树实时荧光定量PCR分析中内参基因的选择. 植物学报, 45(5):579-587. | |
| [28] |
Tanaka Y, Ohmiya A. 2008. Seeing is believing:engineering anthocyanin and carotenoid biosynthetic pathways. Current Opinion in Biotechnology, 19(2):190-197.
doi: 10.1016/j.copbio.2008.02.015 URL |
| [29] |
Tanaka Y, Sasaki N, Ohmiya A. 2008. Biosynthesis of plant pigments:anthocyanins, betalains and carotenoids. The Plant Journal, 54(4):733-749.
doi: 10.1111/j.1365-313X.2008.03447.x URL |
| [30] |
Volkov R A, Panchuk I I, Schöffl F. 2003. Heat-stress-dependency and developmental modulation of gene expression:the potential of house-keeping genes as internal standards in mRNA expression profiling using real-time RT-PCR. Journal of Experimental Botany, 54(391):2343-2349.
doi: 10.1093/jxb/erg244 URL |
| [31] | Wang Yan-jie, Chen Ye-qing, Xue Ze-yun, Zhou Hua, Jin Qi-jiang, Xu Ying-chun. 2017. Selection and validation of reference genes for RT-qPCR normalization in lotus(Nelumbo nucifera)during petal coloration. Journal of Nanjing Agricultural University, 40(3):408-415. (in Chinese) |
| 王彦杰, 陈叶清, 薛泽云, 周华, 金奇江, 徐迎春. 2017. 荷花花瓣着色过程实时荧光定量 PCR内参基因的筛选及验证. 南京农业大学学报, 40(3):408-415. | |
| [32] |
Xu L F, Xu H, Cao Y W, Yang P P, Feng Y Y, Tang Y C, Yuan S X, Ming J. 2017. Validation of reference genes for quantitative real-time PCR during bicolor tepal development in asiatic hybrid lilies(Lilium spp.). Frontiers in Plant Science, 8:669.
doi: 10.3389/fpls.2017.00669 URL |
| [33] | Xu Ling-yun, Wang Jun-li, Zhou Yi-jun. 2017. Cloning and expression analysis of anthocyanin biosynthetic genes from Iris halophila(Iridaceae). Journal of Plant Genetic Resources, 18(2):340-348. (in Chinese) |
| 徐凌云, 王俊丽, 周宜君. 2017. 喜盐鸢尾花色形成关键基因的克隆及表达分析. 植物遗传资源学报, 18(2):340-348. | |
| [34] | Yang Dan, Li Qing, Wang Gui-xi, Ma Qing-hua, Zhu Li-quan. 2017. Reference genes selection and system establishment for real-time qPCR analysis in ping’ou hybrid hazelnut(C. heterophylla Fisch. × C. avellana L.). Scientia Agricultura Sinica, 50(12):2399-2410. (in Chinese) |
| 杨丹, 李清, 王贵禧, 马庆华, 朱利泉. 2017. 平欧杂种榛实时荧光定量 PCR内参基因的筛选与体系建立. 中国农业科学, 50(12):2399-2410. | |
| [35] | Yi S J, Qian Y Q, Han L, Sun Z Y, Fan C M, Liu J X, Ju G S. 2012. Selection of reliable reference genes for gene expression studies in Rhododendron micranthum Turcz. Scientia Horticulturae, 99(2):128-133. |
| [36] |
Yoshihara N, Imayama T, Fukuchi-Mizutani M, Okuhara H, Tanaka Y, Ino I, Yabuya T. 2005. cDNA cloning and characterization of UDP-glucose:Anthocyanidin 3-O-glucosyltransferase in Iris hollandica. Plant Science, 169(3):496-501.
doi: 10.1016/j.plantsci.2005.04.007 URL |
| [37] |
Yoshihara N, Imayama T, Matsuo Y, Fukuchi-Mizutani M, Tanaka Y, Ino I, Yabuya T. 2006. Characterization of cDNA clones encoding anthocyanin 3-p-coumaroyltransferase from Iris hollandica. Plant Science, 171(5):632-639.
doi: 10.1016/j.plantsci.2006.06.005 URL |
| [38] |
Yuan Wei, Wan Hong-jian, Yang Yue-jian. 2012. Characterization and selection of reference genes for real-time quantitative RT-PCR of plants. Chinese Bulletin of Botany, 47(4):427-436. (in Chinese)
doi: 10.3724/SP.J.1259.2012.00427 URL |
| 袁伟, 万红建, 杨悦俭. 2012. 植物实时荧光定量PCR内参基因的特点及选择. 植物学报, 47(4):427-436. | |
| [39] |
Zhang J, Zhu C S, Huo Y B, Zhang B, Ma Z Q, Feng J T. 2019. Validation of suitable reference genes for quantitative gene expression analysis in Tripterygium wilfordii. Molecular Biology Reports, 46:4161-4174.
doi: 10.1007/s11033-019-04867-8 pmid: 31111371 |
| [40] | Zhao Zhi-gang. 2006. Reproductive strategies of common species of Ranunculaceae at Alpine Meadow in Qinghai Tibetan Plateau[Ph. D. Dissertation]. Lanzhou:Lanzhou University. (in Chinese) |
| 赵志刚. 2006. 青藏高原高寒草甸常见毛茛科植物繁殖对策研究[博士论文]. 兰州:兰州大学. | |
| [41] | Zhu Hai-sheng, Chen Min-dong, Wen Qing-fang, Lan Xin-long, Li Yong-ping, Wang Bin, Zhang Qian-rong, Wu Wei-dong. 2016. Cloning of 18S rRNA gene from Luffa cylindrical and its application as an internal standard. Journal of Nuclear Agricultural Sciences, 30(1):35-41. (in Chinese) |
| 朱海生, 陈敏氡, 温庆放, 蓝新隆, 李永平, 王彬, 张前荣, 吴卫东. 2016. 丝瓜18S rRNA基因克隆及其作为内参基因的应用. 核农学报, 30(1):35-41. |
| [1] | 李镇希, 潘睿翾, 许美容, 郑正, 邓晓玲. 柑橘黄龙病菌双重实时荧光PCR检测方法的建立[J]. 园艺学报, 2023, 50(1): 188-196. |
| [2] | 刘鹏, 李钦, 张维瑞, 贺社起, 张素平, 马晓旭, 袁王俊. 连翘褐斑病病原菌鉴定及其对杀菌剂敏感性研究[J]. 园艺学报, 2022, 49(8): 1805-1814. |
| [3] | 张秋悦, 刘昌来, 于晓晶, 杨甲定, 封超年. 盐胁迫条件下杜梨叶片差异表达基因qRT-PCR内参基因筛选[J]. 园艺学报, 2022, 49(7): 1557-1570. |
| [4] | 孟臻, 张伟萍, 王莹, 李龙, 姬小雪, 董贝, 乔康. 番茄枯萎病菌RT-PCR检测技术的建立与应用[J]. 园艺学报, 2022, 49(11): 2479-2488. |
| [5] | 侯天泽, 易双双, 张志群, 王健, 李崇晖. 秋石斛RT-qPCR内参基因的筛选与验证[J]. 园艺学报, 2022, 49(11): 2489-2501. |
| [6] | 蒋梦琦, 薛晓东, 苏立遥, 陈燕, 张舒婷, 李晓斐, 王培育, 张梓浩, 赖钟雄, 林玉玲. 龙眼TCP家族全基因组鉴定与表达分析[J]. 园艺学报, 2021, 48(12): 2481-2496. |
| [7] | 宋蒙, 徐雷锋, 曹雨薇, 杨盼盼, 毕蒙蒙, 何国仁, 唐玉超, 王静, 明军. 百合中车前草花叶病毒的实时荧光定量PCR检测[J]. 园艺学报, 2021, 48(12): 2497-2505. |
| [8] | 许国春, 罗文彬, 李华伟, 许泳清, 纪荣昌, 张鸿, 邱思鑫, 汤浩. 马铃薯叶片光合效率遗传变异分析及高光效种质筛选[J]. 园艺学报, 2021, 48(11): 2239-2250. |
| [9] | 杨婷, 薛珍珍, 李娜, 郎校安, 李凌飞, 钟春梅. 铁十字秋海棠斑叶发育过程内参基因筛选及验证[J]. 园艺学报, 2021, 48(11): 2251-2261. |
| [10] | 程少禹,宣铃娟,董 彬,顾翠花,申亚梅*,张明如,戴梦怡,王卓为,章颖佳,陆丹迎. ‘红元宝’紫玉兰两次花芽分化差异代谢通路及关键调控基因筛选[J]. 园艺学报, 2020, 47(8): 1490-1504. |
| [11] | 贺 振,董婷婷,吴伟文,陈 雯,李良俊*. 莲藕中甘薯潜隐病毒实时荧光定量PCR检测技术的建立及应用[J]. 园艺学报, 2020, 47(7): 1412-1420. |
| [12] | 乔永刚*,王勇飞,曹亚萍,贺嘉欣,贾孟君,李 政,张鑫瑞,宋 芸. 药用蒲公英低温和高温胁迫下内参基因筛选与相关基因表达分析[J]. 园艺学报, 2020, 47(6): 1153-1164. |
| [13] | 闫思远, 杜 娟, 杨富龙, 任苗苗, 李嘉泓, 顾沛雯. PEG介导枸杞内生真菌NQ8GⅡ4遗传转化及转化子评价[J]. 园艺学报, 2020, 47(12): 2385-2396. |
| [14] | 吕前前, 赵兴刚, 王东东, 冒 霞, 左存武, 杨江山, . 解淀粉芽孢杆菌BaA-007鉴定及其对苹果腐烂病的抑制作用[J]. 园艺学报, 2020, 47(10): 1895-1904. |
| [15] | 李嘉维,徐兰依,王冬霜,吕丽洁,陈晓萌,张冬冬*. 枣炭疽病菌拮抗芽孢杆菌的筛选鉴定及其抑菌物质分析[J]. 园艺学报, 2019, 46(12): 2406-2414. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
版权所有 © 2012 《园艺学报》编辑部 京ICP备10030308号-2 国际联网备案号 11010802023439
编辑部地址: 北京市海淀区中关村南大街12号中国农业科学院蔬菜花卉研究所 邮编: 100081
电话: 010-82109523 E-Mail: yuanyixuebao@126.com
技术支持:北京玛格泰克科技发展有限公司