
Acta Horticulturae Sinica ›› 2026, Vol. 53 ›› Issue (4): 1113-1124.doi: 10.16420/j.issn.0513-353x.2025-0105
• Genetic & Breeding · Germplasm Resources · Molecular Biology • Previous Articles Next Articles
GUO Caizhen1,2, WANG Pengfei1, MU Xiaopeng1, ZHANG Jiancheng1, DU Junjie1,*( )
)
Received:2025-04-17
Revised:2025-12-11
Online:2026-04-25
Published:2026-04-20
Contact:
DU Junjie
GUO Caizhen, WANG Pengfei, MU Xiaopeng, ZHANG Jiancheng, DU Junjie. High-Density Genetic Map Construction and QTL Mapping of Fruit Acidity in the Cerasus humilis[J]. Acta Horticulturae Sinica, 2026, 53(4): 1113-1124.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2025-0105
| 遗传参数Genetic parameter | 2021 | 2022 |
|---|---|---|
| 母本/% Female | 2.06 ± 0.03 | 2.13 ± 0.01 |
| 父本/% Male | 0.90 ± 0.03 | 0.91 ± 0.03 |
| F1株数 F1 Number | 139 | 196 |
| 子代平均值/% Progeny mean | 1.82 | 1.58 |
| 变异范围/% Progenies distribution | 0.82 ~ 2.96 | 0.84 ~ 2.80 |
| 变异系数/% CV | 25.74 | 25.22 |
| 超低亲率/% Ultra-low parent | 1.44 | 2.04 |
| 超高亲率/% Ultra-high parent | 30.94 | 10.72 |
| 遗传传递力/% Ta | 122.88 | 104.40 |
| 广义遗传力/% H2 | 99.70 | 99.73 |
| 年份间相关系数Correlation coefficient | 0.66(P < 0.01) | |
Table 1 Genetic analysis of fruit titratable acid content in Cerasus humilis‘Nongda 4'בDS-1'F1 generation
| 遗传参数Genetic parameter | 2021 | 2022 |
|---|---|---|
| 母本/% Female | 2.06 ± 0.03 | 2.13 ± 0.01 |
| 父本/% Male | 0.90 ± 0.03 | 0.91 ± 0.03 |
| F1株数 F1 Number | 139 | 196 |
| 子代平均值/% Progeny mean | 1.82 | 1.58 |
| 变异范围/% Progenies distribution | 0.82 ~ 2.96 | 0.84 ~ 2.80 |
| 变异系数/% CV | 25.74 | 25.22 |
| 超低亲率/% Ultra-low parent | 1.44 | 2.04 |
| 超高亲率/% Ultra-high parent | 30.94 | 10.72 |
| 遗传传递力/% Ta | 122.88 | 104.40 |
| 广义遗传力/% H2 | 99.70 | 99.73 |
| 年份间相关系数Correlation coefficient | 0.66(P < 0.01) | |
| 样本 Sample | Clean Reads总数 Total clean reads | Clean Bases总数/Gb Total clean bases | Q30/% | GC/% |
|---|---|---|---|---|
| 农大4号Nongda 4 | 35 128 984 | 5.21 | 91.16 | 42.13 |
| DS-1 | 36 044 564 | 5.17 | 89.58 | 40.24 |
| F1子代 F1 Progenies | 1 839 622 152 | 261.89 | 89.29 | 39.61 |
| 合计Total | 1 910 795 700 | 272.27 | 90.01 | 40.66 |
Table 2 The sequencing data of parents and F1 progenies
| 样本 Sample | Clean Reads总数 Total clean reads | Clean Bases总数/Gb Total clean bases | Q30/% | GC/% |
|---|---|---|---|---|
| 农大4号Nongda 4 | 35 128 984 | 5.21 | 91.16 | 42.13 |
| DS-1 | 36 044 564 | 5.17 | 89.58 | 40.24 |
| F1子代 F1 Progenies | 1 839 622 152 | 261.89 | 89.29 | 39.61 |
| 合计Total | 1 910 795 700 | 272.27 | 90.01 | 40.66 |
| 名称 Sample | Clean 序列 Clean reads | Clean reads比对率/% Mapped-rate | 平均测序深度/× Ave-depth | 覆盖5×的碱基比 Cov_ratio_5× |
|---|---|---|---|---|
| 农大4号 Nongda 4 | 35 128 984 | 92.71 | 20.33 | 79.66 |
| DS-1 | 3 604 456 | 93.21 | 20.04 | 80.90 |
| F1均值F1 mean | 8 844 337 | 94.00 | 4.96 | 33.94 |
Table 3 The comparison results of sequencing data and reference genome
| 名称 Sample | Clean 序列 Clean reads | Clean reads比对率/% Mapped-rate | 平均测序深度/× Ave-depth | 覆盖5×的碱基比 Cov_ratio_5× |
|---|---|---|---|---|
| 农大4号 Nongda 4 | 35 128 984 | 92.71 | 20.33 | 79.66 |
| DS-1 | 3 604 456 | 93.21 | 20.04 | 80.90 |
| F1均值F1 mean | 8 844 337 | 94.00 | 4.96 | 33.94 |
| 连锁群 Linkage group | SNP数量 SNP number | 总图距/cM Total distance | 平均图距/cM Average distance | 最大Gap/cM Max Gap | Gaps < 5 cM/% |
|---|---|---|---|---|---|
| LG1 | 828 | 162.43 | 0.20 | 8.82 | 99.88 |
| LG2 | 655 | 197.41 | 0.30 | 12.52 | 99.08 |
| LG3 | 962 | 147.40 | 0.15 | 4.60 | 100.00 |
| LG4 | 446 | 153.86 | 0.34 | 7.67 | 99.78 |
| LG5 | 826 | 173.45 | 0.21 | 8.94 | 99.64 |
| LG6 | 558 | 158.51 | 0.28 | 7.43 | 99.82 |
| LG7 | 504 | 159.51 | 0.32 | 5.12 | 99.80 |
| LG8 | 592 | 168.93 | 0.29 | 7.64 | 99.15 |
| 总计 Total | 5 371 | 1 321.50 | — | — | — |
| 平均 Average | 671 | 165 | 0.26 | 7.84 | 99.64 |
Table 4 The basic information of neutral genetic map of Cerasus humilis‘Nongda 4'and‘DS-1'
| 连锁群 Linkage group | SNP数量 SNP number | 总图距/cM Total distance | 平均图距/cM Average distance | 最大Gap/cM Max Gap | Gaps < 5 cM/% |
|---|---|---|---|---|---|
| LG1 | 828 | 162.43 | 0.20 | 8.82 | 99.88 |
| LG2 | 655 | 197.41 | 0.30 | 12.52 | 99.08 |
| LG3 | 962 | 147.40 | 0.15 | 4.60 | 100.00 |
| LG4 | 446 | 153.86 | 0.34 | 7.67 | 99.78 |
| LG5 | 826 | 173.45 | 0.21 | 8.94 | 99.64 |
| LG6 | 558 | 158.51 | 0.28 | 7.43 | 99.82 |
| LG7 | 504 | 159.51 | 0.32 | 5.12 | 99.80 |
| LG8 | 592 | 168.93 | 0.29 | 7.64 | 99.15 |
| 总计 Total | 5 371 | 1 321.50 | — | — | — |
| 平均 Average | 671 | 165 | 0.26 | 7.84 | 99.64 |

Fig. 3 Marker linkage on the neutral genetic map Each cell represents the recombination rate of pair-wise markers. Yellow indicates a lower recombination rate,suggesting a stronger linkage between markers. Purple represents the opposite situation
| 年份 Year | 连锁群 Linkage group | 位点 QTL | 置信区间/ cM Confidence interval | 包含标记数 Mark number | LOD | 贡献率/% Phenotypic variation explained(PVE) |
|---|---|---|---|---|---|---|
| 2021 | LG1 | 21TA-1 | 67.863 | 1 | 3.35 | 10.5 |
| LG1 | 21TA-2 | 77.669 | 1 | 3.72 | 11.6 | |
| LG6 | 21TA-3 | 44.269 ~ 47.515 | 6 | 3.68 ~ 4.17 | 10.9 ~ 13.1 | |
| LG7 | 21TA-4 | 68.627 ~ 69.023 | 2 | 3.29 ~ 3.52 | 10.3 ~ 11.0 | |
| LG7 | 21TA-5 | 69.610 ~ 69.699 | 1 | 3.60 | 11.0 | |
| 2022 | LG1 | 22TA-1 | 71.508 ~ 71.509 | 2 | 4.57 ~ 4.60 | 10.3 ~ 10.4 |
| LG1 | 22TA-2 | 75.915 ~ 75.929 | 4 | 3.42 ~ 3.60 | 7.8 ~ 8.2 | |
| LG1 | 22TA-3 | 85.683 ~ 85.949 | 4 | 3.88 ~ 4.86 | 8.8 ~ 11.0 | |
| LG1 | 22TA-4 | 91.371 ~ 91.417 | 2 | 3.75 ~ 3.86 | 8.6 ~ 8.8 | |
| LG1 | 22TA-5 | 106.103 ~ 106.335 | 3 | 3.76 ~ 5.18 | 8.6 ~ 11.6 | |
| LG2 | 22TA-6 | 99.136 ~ 99.197 | 2 | 3.64 ~ 3.82 | 8.3 ~ 8.7 | |
| LG2 | 22TA-7 | 134.655 ~ 134.665 | 2 | 3.13 ~ 3.31 | 7.2 ~ 7.6 | |
| LG3 | 22TA-8 | 67.600 ~ 67.883 | 4 | 3.07 ~ 3.13 | 7.1 ~ 7.2 |
Table 5 QTL mapping of fruit acidity on genetic map
| 年份 Year | 连锁群 Linkage group | 位点 QTL | 置信区间/ cM Confidence interval | 包含标记数 Mark number | LOD | 贡献率/% Phenotypic variation explained(PVE) |
|---|---|---|---|---|---|---|
| 2021 | LG1 | 21TA-1 | 67.863 | 1 | 3.35 | 10.5 |
| LG1 | 21TA-2 | 77.669 | 1 | 3.72 | 11.6 | |
| LG6 | 21TA-3 | 44.269 ~ 47.515 | 6 | 3.68 ~ 4.17 | 10.9 ~ 13.1 | |
| LG7 | 21TA-4 | 68.627 ~ 69.023 | 2 | 3.29 ~ 3.52 | 10.3 ~ 11.0 | |
| LG7 | 21TA-5 | 69.610 ~ 69.699 | 1 | 3.60 | 11.0 | |
| 2022 | LG1 | 22TA-1 | 71.508 ~ 71.509 | 2 | 4.57 ~ 4.60 | 10.3 ~ 10.4 |
| LG1 | 22TA-2 | 75.915 ~ 75.929 | 4 | 3.42 ~ 3.60 | 7.8 ~ 8.2 | |
| LG1 | 22TA-3 | 85.683 ~ 85.949 | 4 | 3.88 ~ 4.86 | 8.8 ~ 11.0 | |
| LG1 | 22TA-4 | 91.371 ~ 91.417 | 2 | 3.75 ~ 3.86 | 8.6 ~ 8.8 | |
| LG1 | 22TA-5 | 106.103 ~ 106.335 | 3 | 3.76 ~ 5.18 | 8.6 ~ 11.6 | |
| LG2 | 22TA-6 | 99.136 ~ 99.197 | 2 | 3.64 ~ 3.82 | 8.3 ~ 8.7 | |
| LG2 | 22TA-7 | 134.655 ~ 134.665 | 2 | 3.13 ~ 3.31 | 7.2 ~ 7.6 | |
| LG3 | 22TA-8 | 67.600 ~ 67.883 | 4 | 3.07 ~ 3.13 | 7.1 ~ 7.2 |
| 基因ID Gene ID | 位置 Location | 注释 Annotation |
|---|---|---|
| ouLi_002143 | Chr01:21547678-21553051 | 丙酮酸脱氢酶E1亚基Pyruvate dehydrogenase E1 component subunit |
| ouLi_002975 | Chr01:27180756-27193970 | 铝激活苹果酸转运蛋白Aluminum-activated malate transporter(ALMT9) |
| ouLi_004592 | Chr01:36585973-36587431 | V型质子ATP酶c''1亚基V-type proton ATPase subunit c''1(VHA-c''1) |
| ouLi_003613 | Chr01:30704188-30710433 | V型质子ATP酶a3亚基V-type proton ATPase subunit a3(VHA-a3) |
| ouLi_004286 | Chr01:34805791-3498913 | NADP依赖型苹果酸酶NADP-dependent malic enzyme(ME1) |
| ouLi_005250 | Chr01:41036263-41042630 | 磷酸烯醇式丙酮酸羧激酶Phosphoenolpyruvate carboxykinase(PEPCK) |
| ouLi_005370 | Chr01:42084592-42085645 | V型质子ATP酶G1亚基V-type proton ATPase subunit G1(VHA-G1) |
| ouLi_000951 | Chr01:7754069-7757303 | 丙酮酸脱氢酶激酶Pyruvate dehydrogenase kinase(PDK) |
| ouLi_002922 | Chr01:26894035-26900595 | 磷酸烯醇式丙酮酸羧化酶4 Phosphoenolpyruvate carboxylase 4(PEPC4) |
| ouLi_002547 | Chr01:24612635-24617562 | 丙酮酸脱氢酶E1亚基Pyruvate dehydrogenase E1 component subunit |
| ouLi_006462 | Chr02:3493489-3498913 | V型质子ATP酶E2亚基V-type proton ATPase subunit E2(VHA-E2) |
| ouLi_007563 | Chr02:10814886-10815511 | 琥珀酸脱氢酶Succinate dehydrogenase(SDH2-3) |
| ouLi_007908 | Chr02:16373753-16377457 | 琥珀酸脱氢酶Succinate dehydrogenase(SDH6) |
| ouLi_006283 | Chr02:2636998-2641289 | 焦磷酸驱动液泡膜质子泵Pyrophosphate-energized vacuolar membrane proton pump(AVP1) |
| ouLi_007013 | Chr02:6659085-6661164 | V型质子ATP酶G亚基V-type proton ATPase subunit G(VHA-G) |
| ouLi_007032 | Chr02:6777034-6779884 | NAD依赖型异柠檬酸脱氢酶Isocitrate dehydrogenase [NAD](IDH5) |
| ouLi_007280 | Chr02:8350333-8353503 | 磷酸烯醇式丙酮酸羧激酶Phosphoenolpyruvate carboxykinase(PEPCK) |
| ouLi_007281 | Chr02:8357959-8363295 | 磷酸烯醇式丙酮酸羧激酶Phosphoenolpyruvate carboxykinase(PEPCK) |
| ouLi_007562 | Chr02:10809841-10814853 | 琥珀酸脱氢酶Succinate dehydrogenase(SDH2-3) |
| ouLi_009926 | Chr03:9029672-9035094 | 琥珀酰辅酶A连接酶succinate-CoA ligase |
| ouLi_010115 | Chr03:11715042-11717715 | V型质子ATP酶e1亚基V-type proton ATPase subunit e1(VHA-e1) |
| ouLi_010174 | Chr03:12276025-12278623 | NAD依赖型异柠檬酸脱氢酶Isocitrate dehydrogenase [NAD](IDH1) |
Table 6 Candidate genes selected for fruit acidity
| 基因ID Gene ID | 位置 Location | 注释 Annotation |
|---|---|---|
| ouLi_002143 | Chr01:21547678-21553051 | 丙酮酸脱氢酶E1亚基Pyruvate dehydrogenase E1 component subunit |
| ouLi_002975 | Chr01:27180756-27193970 | 铝激活苹果酸转运蛋白Aluminum-activated malate transporter(ALMT9) |
| ouLi_004592 | Chr01:36585973-36587431 | V型质子ATP酶c''1亚基V-type proton ATPase subunit c''1(VHA-c''1) |
| ouLi_003613 | Chr01:30704188-30710433 | V型质子ATP酶a3亚基V-type proton ATPase subunit a3(VHA-a3) |
| ouLi_004286 | Chr01:34805791-3498913 | NADP依赖型苹果酸酶NADP-dependent malic enzyme(ME1) |
| ouLi_005250 | Chr01:41036263-41042630 | 磷酸烯醇式丙酮酸羧激酶Phosphoenolpyruvate carboxykinase(PEPCK) |
| ouLi_005370 | Chr01:42084592-42085645 | V型质子ATP酶G1亚基V-type proton ATPase subunit G1(VHA-G1) |
| ouLi_000951 | Chr01:7754069-7757303 | 丙酮酸脱氢酶激酶Pyruvate dehydrogenase kinase(PDK) |
| ouLi_002922 | Chr01:26894035-26900595 | 磷酸烯醇式丙酮酸羧化酶4 Phosphoenolpyruvate carboxylase 4(PEPC4) |
| ouLi_002547 | Chr01:24612635-24617562 | 丙酮酸脱氢酶E1亚基Pyruvate dehydrogenase E1 component subunit |
| ouLi_006462 | Chr02:3493489-3498913 | V型质子ATP酶E2亚基V-type proton ATPase subunit E2(VHA-E2) |
| ouLi_007563 | Chr02:10814886-10815511 | 琥珀酸脱氢酶Succinate dehydrogenase(SDH2-3) |
| ouLi_007908 | Chr02:16373753-16377457 | 琥珀酸脱氢酶Succinate dehydrogenase(SDH6) |
| ouLi_006283 | Chr02:2636998-2641289 | 焦磷酸驱动液泡膜质子泵Pyrophosphate-energized vacuolar membrane proton pump(AVP1) |
| ouLi_007013 | Chr02:6659085-6661164 | V型质子ATP酶G亚基V-type proton ATPase subunit G(VHA-G) |
| ouLi_007032 | Chr02:6777034-6779884 | NAD依赖型异柠檬酸脱氢酶Isocitrate dehydrogenase [NAD](IDH5) |
| ouLi_007280 | Chr02:8350333-8353503 | 磷酸烯醇式丙酮酸羧激酶Phosphoenolpyruvate carboxykinase(PEPCK) |
| ouLi_007281 | Chr02:8357959-8363295 | 磷酸烯醇式丙酮酸羧激酶Phosphoenolpyruvate carboxykinase(PEPCK) |
| ouLi_007562 | Chr02:10809841-10814853 | 琥珀酸脱氢酶Succinate dehydrogenase(SDH2-3) |
| ouLi_009926 | Chr03:9029672-9035094 | 琥珀酰辅酶A连接酶succinate-CoA ligase |
| ouLi_010115 | Chr03:11715042-11717715 | V型质子ATP酶e1亚基V-type proton ATPase subunit e1(VHA-e1) |
| ouLi_010174 | Chr03:12276025-12278623 | NAD依赖型异柠檬酸脱氢酶Isocitrate dehydrogenase [NAD](IDH1) |
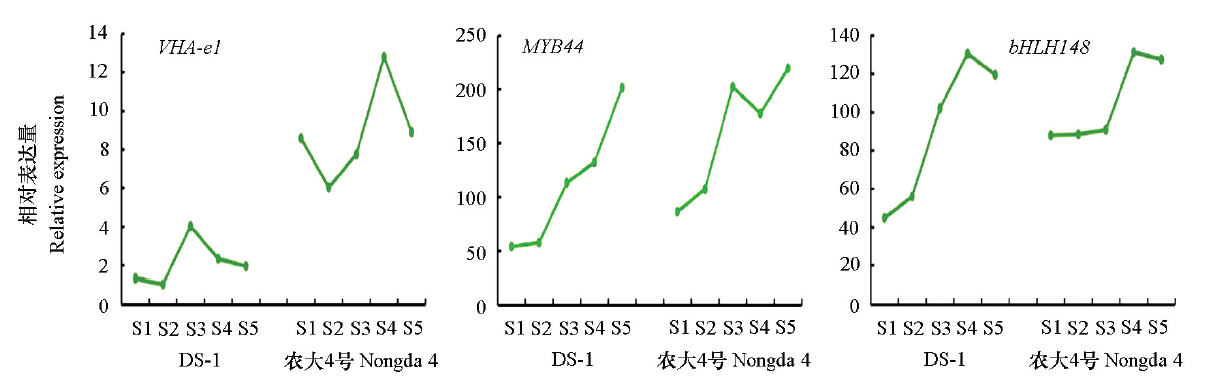
Fig. 5 Dynamic changes of key genes expression during fruit development of different germplasms S1:Young fruit stage;S2:Ard core stage;S3:Turning stage;S4:Coloring stage;S5:Mature stage
| 基因 Gene | 可滴定酸含量 Titrable acid content | 苹果酸含量 Malic acid content |
|---|---|---|
| VHA-e1 | 0.83** | 0.53** |
| MYB44 | 0.69** | 0.80** |
| bHLH148 | 0.56** | 0.68** |
Table 7 Correlation coefficient of titrable acid content and key genes expression
| 基因 Gene | 可滴定酸含量 Titrable acid content | 苹果酸含量 Malic acid content |
|---|---|---|
| VHA-e1 | 0.83** | 0.53** |
| MYB44 | 0.69** | 0.80** |
| bHLH148 | 0.56** | 0.68** |
| [1] |
doi: 10.1080/14620316.2021.1912647 URL |
| [2] |
|
| [3] |
|
|
曹建康, 姜微波, 赵玉梅. 2007. 果蔬采后生理生化实验指导. 北京: 中国轻工业出版社.
|
|
| [4] |
|
| [5] |
|
| [6] |
doi: 10.1590/S1415-47572006000100033 URL |
| [7] |
|
| [8] |
|
| [9] |
|
|
郭彩珍. 2023. 欧李果实有机酸遗传分析和关键基因挖掘及功能验证[博士论文]. 山西: 山西农业大学.
|
|
| [10] |
|
|
[Cerasus humilis(Bge.) Sok.] is controlled by a complex gene regulatory network. Front Plant Sci,13:982112.]
|
|
| [11] |
doi: 10.1111/tpj.2017.91.issue-3 URL |
| [12] |
doi: 10.1104/pp.15.01333 URL |
| [13] |
doi: 10.16420/j.issn.0513-353x.2024-0438 |
|
姜凤超, 杨丽, 张俊环, 张美玲, 于文剑, 孙浩元. 2025. 杏果实中调控有机酸积累的QTL定位及其主效基因筛选. 园艺学报, 52 (4):846-856.
doi: 10.16420/j.issn.0513-353x.2024-0438 |
|
| [14] |
|
| [15] |
|
| [16] |
doi: 10.1007/s13580-022-00473-z |
| [17] |
|
|
彭麟. 2022. 梨高密度SNP图谱构建和果实有机酸的QTL定位[硕士论文]. 杭州: 浙江大学.
|
|
| [18] |
|
|
彭敏, 徐礼羿, 王丽鸳, 韦康, 成浩, 谭礼强. 2018. 杂交一代作图群体大小对茶树QTL效应估计的影响. 植物遗传资源学报, 19 (6):1143-1148.
doi: 10.13430/j.cnki.jpgr.20180406001 |
|
| [19] |
|
| [20] |
|
| [21] |
doi: 10.1093/bioinformatics/btt563 pmid: 24078685 |
| [22] |
|
| [23] |
|
| [24] |
|
| [25] |
doi: 10.16420/j.issn.0513-353x.2020-0933 |
|
唐海霞, 高瑞, 王中堂, 张琼. 2021. 基于SNP标记的枣高密度遗传连锁图谱重新构建. 园艺学报, 48 (11):2275-2285.
doi: 10.16420/j.issn.0513-353x.2020-0933 |
|
| [26] |
doi: 10.1016/S0981-9428(98)80078-8 URL |
| [27] |
|
|
王鹏飞. 2015. 欧李果实转录组测序及苹果酸积累关键酶基因的克隆与表达分析[博士论文]. 山西: 山西农业大学.
|
|
| [28] |
doi: 10.1007/s11676-017-0418-3 URL |
| [29] |
|
| [30] |
doi: 10.1111/pbi.v18.3 URL |
| [31] |
doi: 10.1016/j.hpj.2022.11.005 URL |
| [32] |
|
| [33] |
|
| [34] |
|
|
要燕杰. 2020. 野生大豆和栽培大豆茎杆和籽粒性状的QTL定位和RNA-seq分析[博士论文]. 武汉: 华中农业大学.
|
|
| [35] |
|
|
姚玉新, 李明, 由春香, 刘志, 王冬梅, 郝玉金. 2010. 苹果果实中苹果酸代谢关键酶与苹果酸和可溶性糖积累的关系. 园艺学报, 37 (1):1-8.
|
|
| [36] |
doi: 10.1111/pbi.v19.2 URL |
| [37] |
doi: 10.1093/genetics/136.4.1457 pmid: 8013918 |
| [38] |
doi: 10.1093/dnares/dsv003 pmid: 25776277 |
| [39] |
doi: 10.16420/j.issn.0513-353x.2023-0514 |
|
张立华, 徐玉, 郑丽桐, 王长智, 祝令成, 马百全, 李明军. 2024. 酸转运蛋白与果实酸积累关系的研究进展. 园艺学报, 51 (7):1474-1488.
|
|
| [40] |
|
| [41] |
|
| [42] |
|
| [1] | GAO Xu, HE Daogen, TAN Guoyin, ZHU Changzhi, ZHANG Hongyun, XIAO Jiarui, WANG Hui. QTL Mapping and Candidate Gene Analysis of Related Traits of Broccoli [J]. Acta Horticulturae Sinica, 2025, 52(12): 3189-3201. |
| [2] | JIANG Shuang, WANG Xiaoqing, SHI Chunhui, LI Shuigen, LUO Jun. Detection of SLAF Tag Polymorphism Associated with Pear Russet Peel by PCR Product Pool Resequencing [J]. Acta Horticulturae Sinica, 2024, 51(4): 707-714. |
| [3] | ZHANG Zhenghai, WANG Yongfu, WU Huamao, YU Hailong, CAO Yacong, FENG Xigang, WANG Lihao. Advances in Genetic Mapping and Candidate Gene Analysis of Important Traits in Pepper(Capsicum spp.) [J]. Acta Horticulturae Sinica, 2024, 51(3): 669-696. |
| [4] | YAO Yuan, DENG Lijun, HU Juan, TANG Xiaoyu, WANG Tie, LI Hang, SUN Guochao, XIONG Bo, LIAO Ling, WANG Zhihui. Whole Genome Resequencing Analysis of‘Cuihongli’Plum and Its Early-Ripening Bud Sport Mutation [J]. Acta Horticulturae Sinica, 2024, 51(10): 2255-2266. |
| [5] | NIE Xinghua, ZHANG Yu, LIU Song, YANG Jiabin, HAO Yaqiong, LIU Yang, QIN Ling, XING Yu. Study on Genetic Characteristics and Taxonomic Status of Wild Chinese Chestnut Based on Genome Re-sequencing [J]. Acta Horticulturae Sinica, 2023, 50(8): 1622-1636. |
| [6] | TANG Yuqing, YANG Huidong, YAN Chengpu, WANG Siyu, WANG Yuting, HU Zhongdong, ZHU Fanghong. Development and Application of Jinlan Pummelo(Citrus maxima)InDel Markers Based on Genome Re-sequencing [J]. Acta Horticulturae Sinica, 2023, 50(1): 15-26. |
| [7] | ZHAO Yong, ZHU Hongju, YANG Dongdong, GONG Chengsheng, LIU Wenge. Research Progress of Citric Acid Metabolism in the Fruit [J]. Acta Horticulturae Sinica, 2022, 49(12): 2579-2596. |
| [8] | JIANG Shuang, ZHANG Xueying, AN Haishan, XU Fangjie, ZHANG Jiaying. Development and Analysis of Polymorphism of SSR Markers in the Whole Genome of Loquat [J]. Acta Horticulturae Sinica, 2021, 48(5): 1013-1022. |
| [9] | LIU Mengyu,LIU Xiaofeng,JIANG Dong,ZHU Shiping,SHEN Wanxia,YU Xin,XUE Yang,and ZHAO Xiaochun*. Identification of Genes Related to Oil Gland Development in Kumquat by Using BSA-Seq [J]. ACTA HORTICULTURAE SINICA, 2019, 46(5): 841-854. |
| [10] | ZHANG Xiaojing1,ZHANG Kaige1,ZHU Huayu1,2,ZHANG Yu1,HU Qianmei1,CHENG Siyuan1, ZHANG Minjuan1,HU Jianbin1,2,and YANG Luming1,2,*. QTL Mapping of Lateral Branch Associated Traits in Cucumis melo [J]. ACTA HORTICULTURAE SINICA, 2019, 46(3): 519-528. |
| [11] | OU Chenggang1,SUN Tingting1,LIU Xinyan1,LI Chengjiang2,XU Donghui1,ZHAO Zhiwei1,and ZHUANG Feiyun1,*. Fine Mapping of QTLs Related to Main Carotenoids in Carrot Root [J]. ACTA HORTICULTURAE SINICA, 2017, 44(2): 288-296. |
| [12] | YANG Shu-hua1,LI Shi-chao1,JIA Rui-dong1,ZHAO Xin1,JIN Mao-yong2,LI Feng-jie2,and GE Hong1,*. Primary Construction of a Genetic Linkage Map of Rosa beggeriana‘Aurea’× R. Davurica [J]. ACTA HORTICULTURAE SINICA, 2015, 42(12): 2526-2534. |
| [13] | ZHOU Kun-Hua, CHEN Xue-Jun, FANG Rong, CHEN Li-Zhen, ZONG Hong-Xia, MIAO 南Sheng. Construction and Analysis of an Interspecific Linkage Map of Capsicum annuum × C. frutescens [J]. ACTA HORTICULTURAE SINICA, 2013, 40(11): 2171-2179. |
| [14] | MENG Lin, LIU Bo, LIN Liang-Bin, CHENG Feng, WANG Xiao-Wu, WU Jian. Development of InDel Markers for Brassica campestris and Genetic Linkage Map Construction of the RILs Population [J]. ACTA HORTICULTURAE SINICA, 2012, 39(8): 1491-. |
| [15] | GAO Mei-ling;ZHU Zi-cheng;GAO Peng;and LUAN Fei-shi;. A Microsatellite-based Genetic Map of Melon and Localization of Gene for Gynoecious Sex Expression Using Recombinant Inbred Lines [J]. ACTA HORTICULTURAE SINICA, 2011, 38(7): 1308-1316. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd