
Acta Horticulturae Sinica ›› 2023, Vol. 50 ›› Issue (6): 1230-1242.doi: 10.16420/j.issn.0513-353x.2022-0509
• Genetic & Breeding·Germplasm Resources·Molecular Biology • Previous Articles Next Articles
HUANG Zhihao1, LIU Tingting1, DONG Xujie1, YAN Mingli2, LIUZhixiang 1, ZENG Chaozhen1,*( )
)
Received:2022-10-21
Revised:2023-03-14
Online:2023-06-25
Published:2023-06-27
Contact:
* (E-mail:CLC Number:
HUANG Zhihao, LIU Tingting, DONG Xujie, YAN Mingli, LIUZhixiang , ZENG Chaozhen. Identification and Expression Analysis of HMA Gene Family in Brassica juncea Under Cadmium Stress[J]. Acta Horticulturae Sinica, 2023, 50(6): 1230-1242.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2022-0509
| 基因ID Gene ID | 正向引物(5′-3′) Forward primer | 反向引物(5′-3′) Reverse primer |
|---|---|---|
| BjuB002000 | ATCGTTGGCTTCCCTCTCGTT | ACATCCACCATTGACCGGCTTG |
| BjuA003596 | AAGAACCGTCATCGTCGTCCA | AGAAGTACGCCGGAAACCAC |
| BjuB040613 | TCATCGTCGTCCATGATAGCC | AATTAGTTTCTCCGGTTACCCTC |
| BjuO003139 | CGATACTTGCCAAAGCCGTTG | TGCATAACCGCATTCGCCTT |
| BjuA025267 | GCCTCCTCATCTCTCCGTCCC | TTTGCCGTCCACTTTCACGTT |
| BjuA042408 | CAAGAACCGTCATCGTTGTCC | GCAACAAGAGCGAACCACT |
| Actin | GAATCCACGAGACGACTTACAAC | CGATCCAGACACTGTACTTCCTC |
Table 1 The sequences of qRT-PCR primers
| 基因ID Gene ID | 正向引物(5′-3′) Forward primer | 反向引物(5′-3′) Reverse primer |
|---|---|---|
| BjuB002000 | ATCGTTGGCTTCCCTCTCGTT | ACATCCACCATTGACCGGCTTG |
| BjuA003596 | AAGAACCGTCATCGTCGTCCA | AGAAGTACGCCGGAAACCAC |
| BjuB040613 | TCATCGTCGTCCATGATAGCC | AATTAGTTTCTCCGGTTACCCTC |
| BjuO003139 | CGATACTTGCCAAAGCCGTTG | TGCATAACCGCATTCGCCTT |
| BjuA025267 | GCCTCCTCATCTCTCCGTCCC | TTTGCCGTCCACTTTCACGTT |
| BjuA042408 | CAAGAACCGTCATCGTTGTCC | GCAACAAGAGCGAACCACT |
| Actin | GAATCCACGAGACGACTTACAAC | CGATCCAGACACTGTACTTCCTC |
| 基因Gene | 结构域Domain | 基因Gene | 结构域Domain | |||||
|---|---|---|---|---|---|---|---|---|
| HMA | E1-E2 ATPase | Hydrolase | HMA | E1-E2 ATPase | Hydrolase | |||
| AtHMA1 | - | E1-E2 ATPase | Hydrolase | BjuO003139 | - | E1-E2 ATPase | Hydrolase | |
| AtHMA2 | - | E1-E2 ATPase | Hydrolase | BjuB035256 | - | E1-E2 ATPase | Hydrolase | |
| AtHMA3 | - | E1-E2 ATPase | Hydrolase | BjuA023298 | - | E1-E2 ATPase | Hydrolase | |
| AtHMA4 | - | E1-E2 ATPase | Hydrolase | BjuA025267 | - | E1-E2 ATPase | Hydrolase | |
| AtHMA5 | [HMA]3 | E1-E2 ATPase | Hydrolase | BjuA042408 | HMA | E1-E2 ATPase | Hydrolase | |
| AtHMA6 | HMA | E1-E2 ATPase | Hydrolase | BjuB028207 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| AtHMA7 | [HMA]2 | E1-E2 ATPase | Hydrolase | BjuA046861 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| AtHMA8 | HMA | E1-E2 ATPase | Hydrolase | BjuB035323 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuB002000 | - | E1-E2 ATPase | Hydrolase | BjuA038121 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuB034828 | - | E1-E2 ATPase | Hydrolase | BjuB019118 | HMA | E1-E2 ATPase | Hydrolase | |
| BjuA036645 | - | E1-E2 ATPase | Hydrolase | BjuB027969 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuA002855 | - | E1-E2 ATPase | Hydrolase | BjuA025022 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuO011640 | HMA | E1-E2 ATPase | Hydrolase | BjuB036631 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuA003595 | - | E1-E2 ATPase | Hydrolase | BjuA033764 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuB040614 | - | E1-E2 ATPase | Hydrolase | BjuA003262 | HMA | E1-E2 ATPase | Hydrolase | |
| BjuA003596 | HMA | E1-E2 ATPase | Hydrolase | BjuA038847 | HMA | E1-E2 ATPase | Hydrolase | |
| BjuB040613 | HMA | E1-E2 ATPase | Hydrolase | BjuB045293 | HMA | E1-E2 ATPase | Hydrolase | |
| BjuA042404 | [HMA]3 | E1-E2 ATPase | Hydrolase | |||||
Table 2 Distribution of domains in HMA gene family
| 基因Gene | 结构域Domain | 基因Gene | 结构域Domain | |||||
|---|---|---|---|---|---|---|---|---|
| HMA | E1-E2 ATPase | Hydrolase | HMA | E1-E2 ATPase | Hydrolase | |||
| AtHMA1 | - | E1-E2 ATPase | Hydrolase | BjuO003139 | - | E1-E2 ATPase | Hydrolase | |
| AtHMA2 | - | E1-E2 ATPase | Hydrolase | BjuB035256 | - | E1-E2 ATPase | Hydrolase | |
| AtHMA3 | - | E1-E2 ATPase | Hydrolase | BjuA023298 | - | E1-E2 ATPase | Hydrolase | |
| AtHMA4 | - | E1-E2 ATPase | Hydrolase | BjuA025267 | - | E1-E2 ATPase | Hydrolase | |
| AtHMA5 | [HMA]3 | E1-E2 ATPase | Hydrolase | BjuA042408 | HMA | E1-E2 ATPase | Hydrolase | |
| AtHMA6 | HMA | E1-E2 ATPase | Hydrolase | BjuB028207 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| AtHMA7 | [HMA]2 | E1-E2 ATPase | Hydrolase | BjuA046861 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| AtHMA8 | HMA | E1-E2 ATPase | Hydrolase | BjuB035323 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuB002000 | - | E1-E2 ATPase | Hydrolase | BjuA038121 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuB034828 | - | E1-E2 ATPase | Hydrolase | BjuB019118 | HMA | E1-E2 ATPase | Hydrolase | |
| BjuA036645 | - | E1-E2 ATPase | Hydrolase | BjuB027969 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuA002855 | - | E1-E2 ATPase | Hydrolase | BjuA025022 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuO011640 | HMA | E1-E2 ATPase | Hydrolase | BjuB036631 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuA003595 | - | E1-E2 ATPase | Hydrolase | BjuA033764 | [HMA]2 | E1-E2 ATPase | Hydrolase | |
| BjuB040614 | - | E1-E2 ATPase | Hydrolase | BjuA003262 | HMA | E1-E2 ATPase | Hydrolase | |
| BjuA003596 | HMA | E1-E2 ATPase | Hydrolase | BjuA038847 | HMA | E1-E2 ATPase | Hydrolase | |
| BjuB040613 | HMA | E1-E2 ATPase | Hydrolase | BjuB045293 | HMA | E1-E2 ATPase | Hydrolase | |
| BjuA042404 | [HMA]3 | E1-E2 ATPase | Hydrolase | |||||
| 亚族 Subgroup | 基因 Gene | 长度/aa Length | 分子量/kD MW | 等电点 pI | 不稳定指数Instability index | 疏水性指数 GRAVY | 亚细胞定位 Subcellular location | |
|---|---|---|---|---|---|---|---|---|
| WoLF PSORT | BUSCA | |||||||
| P-1B-1 | BjuB028207 | 998 | 108.48 | 5.72 | 35.69 | 0.16 | PM | PM |
| BjuA046861 | 1 184 | 129.44 | 5.62 | 35.11 | 0.08 | PM | PM | |
| BjuB019118 | 839 | 90.81 | 5.55 | 27.34 | 0.28 | PM | PM | |
| BjuB035323 | 978 | 106.55 | 5.94 | 28.84 | 0.17 | PM | PM | |
| BjuA038121 | 973 | 105.65 | 6.13 | 26.97 | 0.19 | PM | PM | |
| BjuB027969 | 1 006 | 107.54 | 5.02 | 32.02 | 0.22 | PM | PM | |
| BjuA033764 | 980 | 104.40 | 4.93 | 31.54 | 0.23 | PM | PM | |
| BjuA025022 | 999 | 106.84 | 5.00 | 33.47 | 0.25 | PM | PM | |
| BjuB036631 | 1 000 | 106.96 | 5.01 | 33.48 | 0.25 | PM | PM | |
| BjuA003262 | 938 | 98.60 | 8.76 | 41.58 | 0.11 | PM | MM | |
| BjuA038847 | 885 | 94.22 | 5.68 | 37.54 | 0.16 | PM | MM | |
| P-1B-2 | BjuA002855 | 683 | 73.81 | 7.23 | 33.38 | 0.06 | PM | MM |
| BjuO011640 | 801 | 86.29 | 7.10 | 36.86 | 0.22 | PM | PM | |
| BjuA003595 | 758 | 81.39 | 5.98 | 33.75 | 0.26 | PM | PM | |
| BjuB040614 | 760 | 81.84 | 5.88 | 33.83 | 0.25 | PM | PM | |
| BjuA003596 | 872 | 94.33 | 6.93 | 35.83 | 0.04 | PM | PM | |
| BjuB040613 | 928 | 99.86 | 6.78 | 35.05 | 0.04 | PM | PM | |
| BjuA042404 | 903 | 97.14 | 7.97 | 32.33 | 0 | PM | PM | |
| BjuO003139 | 1 271 | 137.78 | 7.79 | 84.05 | −0.24 | PM | PM | |
| BjuB035256 | 1 220 | 132.84 | 6.23 | 43.35 | −0.19 | PM | PM | |
| BjuA023298 | 1 193 | 129.78 | 6.82 | 41.95 | −0.23 | PM | PM | |
| BjuA025267 | 1 162 | 125.17 | 6.31 | 91.34 | −0.08 | PM | PM | |
| BjuA042408 | 795 | 85.53 | 6.80 | 38.11 | 0.25 | PM | PM | |
| BjuB045293 | 613 | 63.83 | 9.29 | 46.64 | 0.22 | PM | MM | |
| P-1B-4 | BjuB002000 | 820 | 88.05 | 7.87 | 35.75 | 0.12 | PM | MM |
| BjuB034828 | 783 | 84.77 | 6.86 | 35.14 | 0.18 | PM | OM | |
| BjuA036645 | 775 | 84.25 | 6.74 | 34.78 | 0.13 | PM | OM | |
Table 3 Protein information and subcellular localization of HMA family
| 亚族 Subgroup | 基因 Gene | 长度/aa Length | 分子量/kD MW | 等电点 pI | 不稳定指数Instability index | 疏水性指数 GRAVY | 亚细胞定位 Subcellular location | |
|---|---|---|---|---|---|---|---|---|
| WoLF PSORT | BUSCA | |||||||
| P-1B-1 | BjuB028207 | 998 | 108.48 | 5.72 | 35.69 | 0.16 | PM | PM |
| BjuA046861 | 1 184 | 129.44 | 5.62 | 35.11 | 0.08 | PM | PM | |
| BjuB019118 | 839 | 90.81 | 5.55 | 27.34 | 0.28 | PM | PM | |
| BjuB035323 | 978 | 106.55 | 5.94 | 28.84 | 0.17 | PM | PM | |
| BjuA038121 | 973 | 105.65 | 6.13 | 26.97 | 0.19 | PM | PM | |
| BjuB027969 | 1 006 | 107.54 | 5.02 | 32.02 | 0.22 | PM | PM | |
| BjuA033764 | 980 | 104.40 | 4.93 | 31.54 | 0.23 | PM | PM | |
| BjuA025022 | 999 | 106.84 | 5.00 | 33.47 | 0.25 | PM | PM | |
| BjuB036631 | 1 000 | 106.96 | 5.01 | 33.48 | 0.25 | PM | PM | |
| BjuA003262 | 938 | 98.60 | 8.76 | 41.58 | 0.11 | PM | MM | |
| BjuA038847 | 885 | 94.22 | 5.68 | 37.54 | 0.16 | PM | MM | |
| P-1B-2 | BjuA002855 | 683 | 73.81 | 7.23 | 33.38 | 0.06 | PM | MM |
| BjuO011640 | 801 | 86.29 | 7.10 | 36.86 | 0.22 | PM | PM | |
| BjuA003595 | 758 | 81.39 | 5.98 | 33.75 | 0.26 | PM | PM | |
| BjuB040614 | 760 | 81.84 | 5.88 | 33.83 | 0.25 | PM | PM | |
| BjuA003596 | 872 | 94.33 | 6.93 | 35.83 | 0.04 | PM | PM | |
| BjuB040613 | 928 | 99.86 | 6.78 | 35.05 | 0.04 | PM | PM | |
| BjuA042404 | 903 | 97.14 | 7.97 | 32.33 | 0 | PM | PM | |
| BjuO003139 | 1 271 | 137.78 | 7.79 | 84.05 | −0.24 | PM | PM | |
| BjuB035256 | 1 220 | 132.84 | 6.23 | 43.35 | −0.19 | PM | PM | |
| BjuA023298 | 1 193 | 129.78 | 6.82 | 41.95 | −0.23 | PM | PM | |
| BjuA025267 | 1 162 | 125.17 | 6.31 | 91.34 | −0.08 | PM | PM | |
| BjuA042408 | 795 | 85.53 | 6.80 | 38.11 | 0.25 | PM | PM | |
| BjuB045293 | 613 | 63.83 | 9.29 | 46.64 | 0.22 | PM | MM | |
| P-1B-4 | BjuB002000 | 820 | 88.05 | 7.87 | 35.75 | 0.12 | PM | MM |
| BjuB034828 | 783 | 84.77 | 6.86 | 35.14 | 0.18 | PM | OM | |
| BjuA036645 | 775 | 84.25 | 6.74 | 34.78 | 0.13 | PM | OM | |
| 亚族 Subgroup | 基因 Gene | 染色体 Chr | 顺式作用元件数量 Number of cis-acting element | |||||
|---|---|---|---|---|---|---|---|---|
| G-Box | ABRE | LTR | TC | MBS | TGA | |||
| P-1B-1 | AtHMA5 | 1 | 1 | 1 | 0 | 2 | 0 | 2 |
| AtHMA6 | 4 | 0 | 1 | 0 | 1 | 1 | 1 | |
| AtHMA7 | 5 | 1 | 1 | 0 | 3 | 1 | 1 | |
| AtHMA8 | 5 | 0 | 1 | 0 | 1 | 2 | 1 | |
| BjuB028207 | B04 | 0 | 1 | 0 | 0 | 0 | 1 | |
| BjuA046861 | A09 | 0 | 0 | 0 | 0 | 0 | 0 | |
| BjuB019118 | B08 | 0 | 0 | 0 | 0 | 1 | 0 | |
| BjuB035323 | B02 | 0 | 0 | 0 | 1 | 0 | 0 | |
| BjuA038121 | A10 | 0 | 0 | 0 | 2 | 0 | 0 | |
| BjuB027969 | B04 | 0 | 0 | 1 | 3 | 1 | 0 | |
| BjuA033764 | A09 | 0 | 0 | 1 | 3 | 1 | 0 | |
| BjuA025022 | A06 | 1 | 1 | 0 | 0 | 1 | 0 | |
| BjuB036631 | B02 | 1 | 1 | 0 | 0 | 1 | 0 | |
| BjuA003262 | A01 | 0 | 0 | 1 | 0 | 0 | 0 | |
| BjuA038847 | A10 | 7 | 7 | 2 | 0 | 1 | 0 | |
| P-1B-2 | AtHMA2 | 4 | 1 | 0 | 0 | 1 | 2 | 0 |
| AtHMA3 | 4 | 1 | 0 | 1 | 0 | 2 | 2 | |
| AtHMA4 | 2 | 0 | 1 | 1 | 1 | 0 | 3 | |
| BjuA002855 | A01 | 0 | 0 | 0 | 0 | 1 | 0 | |
| BjuO011640 | Super_scaffold_184_753382_1581011 | 1 | 1 | 0 | 0 | 1 | 2 | |
| BjuA003595 | A01 | 0 | 0 | 0 | 0 | 0 | 0 | |
| BjuB040614 | B05 | 1 | 1 | 0 | 2 | 0 | 0 | |
| BjuA003596 | A01 | 2 | 2 | 2 | 1 | 0 | 3 | |
| BjuB040613 | B05 | 2 | 2 | 0 | 0 | 0 | 0 | |
| BjuA042404 | A03 | 2 | 3 | 2 | 0 | 1 | 0 | |
| BjuO003139 | Contig2023 | 2 | 2 | 1 | 0 | 0 | 0 | |
| BjuB035256 | B02 | 3 | 1 | 3 | 1 | 0 | 1 | |
| BjuA023298 | A06 | 1 | 1 | 0 | 2 | 0 | 0 | |
| BjuA025267 | A07 | 1 | 1 | 3 | 0 | 0 | 0 | |
| BjuA042408 | A03 | 2 | 0 | 0 | 1 | 1 | 1 | |
| BjuB045293 | B05 | 0 | 0 | 0 | 0 | 0 | 0 | |
| P-1B-4 | AtHMA1 | 4 | 0 | 0 | 1 | 2 | 2 | 2 |
| BjuB002000 | B05 | 1 | 1 | 0 | 0 | 1 | 1 | |
| BjuB034828 | B02 | 0 | 0 | 0 | 0 | 0 | 0 | |
| BjuA036645 | A09 | 1 | 1 | 0 | 0 | 1 | 1 | |
Table 4 The cis-acting element in the promoter region of the HMA gene family in Brassica juncea
| 亚族 Subgroup | 基因 Gene | 染色体 Chr | 顺式作用元件数量 Number of cis-acting element | |||||
|---|---|---|---|---|---|---|---|---|
| G-Box | ABRE | LTR | TC | MBS | TGA | |||
| P-1B-1 | AtHMA5 | 1 | 1 | 1 | 0 | 2 | 0 | 2 |
| AtHMA6 | 4 | 0 | 1 | 0 | 1 | 1 | 1 | |
| AtHMA7 | 5 | 1 | 1 | 0 | 3 | 1 | 1 | |
| AtHMA8 | 5 | 0 | 1 | 0 | 1 | 2 | 1 | |
| BjuB028207 | B04 | 0 | 1 | 0 | 0 | 0 | 1 | |
| BjuA046861 | A09 | 0 | 0 | 0 | 0 | 0 | 0 | |
| BjuB019118 | B08 | 0 | 0 | 0 | 0 | 1 | 0 | |
| BjuB035323 | B02 | 0 | 0 | 0 | 1 | 0 | 0 | |
| BjuA038121 | A10 | 0 | 0 | 0 | 2 | 0 | 0 | |
| BjuB027969 | B04 | 0 | 0 | 1 | 3 | 1 | 0 | |
| BjuA033764 | A09 | 0 | 0 | 1 | 3 | 1 | 0 | |
| BjuA025022 | A06 | 1 | 1 | 0 | 0 | 1 | 0 | |
| BjuB036631 | B02 | 1 | 1 | 0 | 0 | 1 | 0 | |
| BjuA003262 | A01 | 0 | 0 | 1 | 0 | 0 | 0 | |
| BjuA038847 | A10 | 7 | 7 | 2 | 0 | 1 | 0 | |
| P-1B-2 | AtHMA2 | 4 | 1 | 0 | 0 | 1 | 2 | 0 |
| AtHMA3 | 4 | 1 | 0 | 1 | 0 | 2 | 2 | |
| AtHMA4 | 2 | 0 | 1 | 1 | 1 | 0 | 3 | |
| BjuA002855 | A01 | 0 | 0 | 0 | 0 | 1 | 0 | |
| BjuO011640 | Super_scaffold_184_753382_1581011 | 1 | 1 | 0 | 0 | 1 | 2 | |
| BjuA003595 | A01 | 0 | 0 | 0 | 0 | 0 | 0 | |
| BjuB040614 | B05 | 1 | 1 | 0 | 2 | 0 | 0 | |
| BjuA003596 | A01 | 2 | 2 | 2 | 1 | 0 | 3 | |
| BjuB040613 | B05 | 2 | 2 | 0 | 0 | 0 | 0 | |
| BjuA042404 | A03 | 2 | 3 | 2 | 0 | 1 | 0 | |
| BjuO003139 | Contig2023 | 2 | 2 | 1 | 0 | 0 | 0 | |
| BjuB035256 | B02 | 3 | 1 | 3 | 1 | 0 | 1 | |
| BjuA023298 | A06 | 1 | 1 | 0 | 2 | 0 | 0 | |
| BjuA025267 | A07 | 1 | 1 | 3 | 0 | 0 | 0 | |
| BjuA042408 | A03 | 2 | 0 | 0 | 1 | 1 | 1 | |
| BjuB045293 | B05 | 0 | 0 | 0 | 0 | 0 | 0 | |
| P-1B-4 | AtHMA1 | 4 | 0 | 0 | 1 | 2 | 2 | 2 |
| BjuB002000 | B05 | 1 | 1 | 0 | 0 | 1 | 1 | |
| BjuB034828 | B02 | 0 | 0 | 0 | 0 | 0 | 0 | |
| BjuA036645 | A09 | 1 | 1 | 0 | 0 | 1 | 1 | |
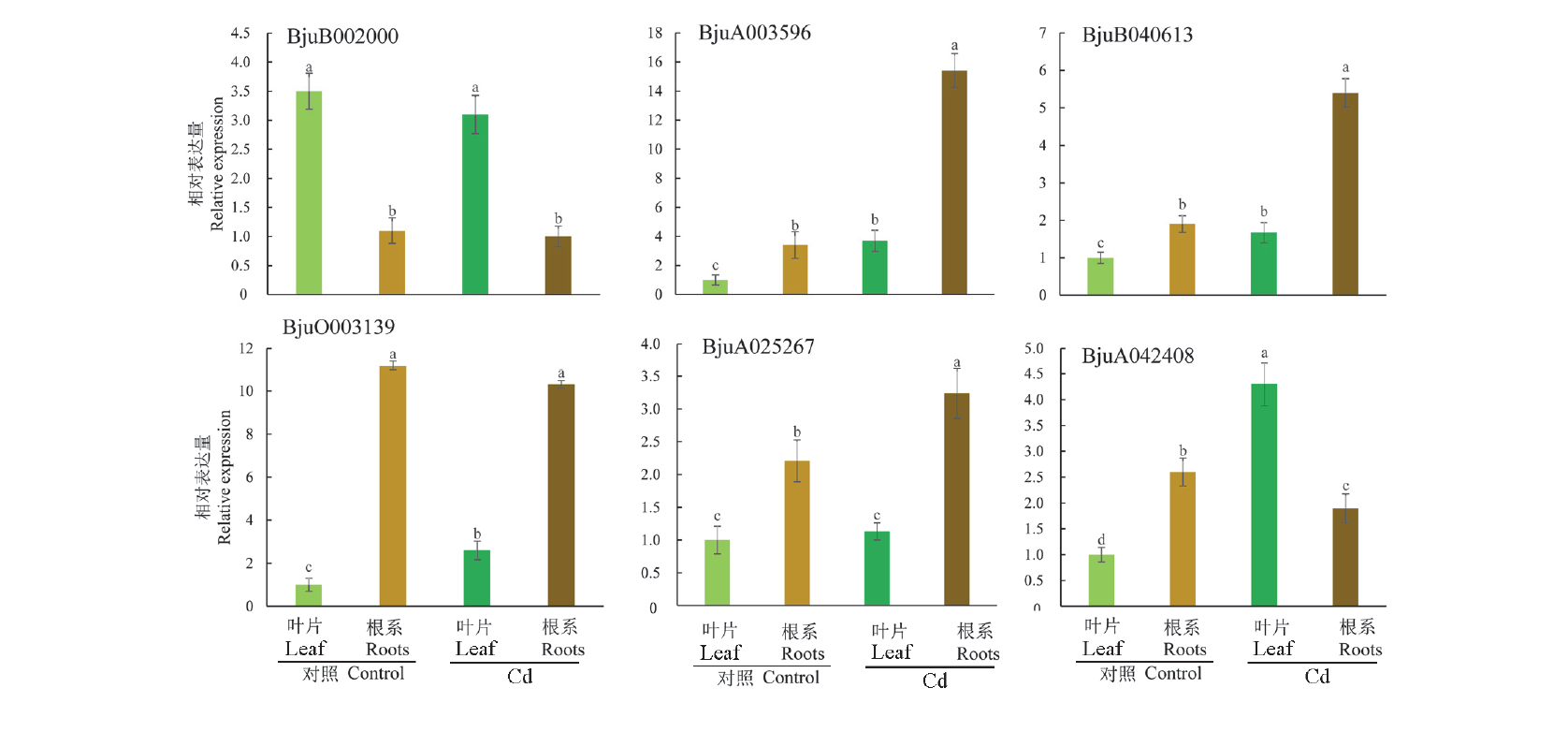
Fig. 5 Expression of HMA gene in Brassica juncea in different locations and under cadmium stress analyzed by qRT-PCR Different letters indicate significantly different values(P < 0.05).
| [1] | Arguëllo J M. 2003. Identification of ion-selectivity determinants in heavy-metal transport P1B-type ATPases. Molecular Membrane Biology, 195 (2):93-108. |
| [2] |
Axelsen K B, Palmgren M G. 2001. Inventory of the superfamily of P-type ion pumps in Arabidopsis. Plant Physiology, 126 (2):696-706.
pmid: 11402198 |
| [3] | Choudhury M R, Islam M S, Ahmed Z U, Nayar F. 2016. Phytoremediation of heavy metal contaminated buriganga riverbed sediment by Indian mustard and marigold plants. Environmental Progress & Sustainable Energy, 35 (1):117-124. |
| [4] |
Clemens S. 2001. Molecular mechanisms of plant metal tolerance and homeostasis. Planta, 212 (4):475-486.
doi: 10.1007/s004250000458 pmid: 11525504 |
| [5] |
Cola N A, Sancenon V, Navarro S R, Mayo S, Thiele D J, Ecker J R, Puig S, Penarrubia L. 2006. The Arabidopsis heavy metal P-type ATPase HMA 5 interacts with metallochaperones and functions in copper detoxification of roots. Plant Journal, 45 (2):225-236.
doi: 10.1111/tpj.2006.45.issue-2 URL |
| [6] |
Courbot M, Willems G, Motte P, Arvidsson S, Roosens N, Laprade P S, erbruggen N V. 2007. A major quantitative trait locus for cadmium tolerance in Arabidopsis halleri colocalizes with HMA4,a gene encoding a heavy metal ATPase. Plant Physiology, 144 (2):1052-1065.
doi: 10.1104/pp.106.095133 pmid: 17434989 |
| [7] |
Ghany S E A, Moule P M, Niyogi K K, Pilon M, Shikanai T. 2005. Two P-type ATPases are required for copper delivery in Arabidopsis thaliana chloroplasts. Plant Cell, 17 (4):1233-1251.
doi: 10.1105/tpc.104.030452 URL |
| [8] |
Goswami S, Das S. 2015. A study on cadmium phytoremediation potential of indian mustard,Brassica juncea. International Journal of Phytoremediation, 17 (6):583-588.
doi: 10.1080/15226514.2014.935289 URL |
| [9] |
Gravot A, Lieutaud A, Verret F, Auroy P, Vavasseur A, Richaud P. 2004. AtHMA3,a plant P1B-ATPase,functions as a Cd/Pb transporter in yeast. Febs Letters, 561 (1-3):22-28.
doi: 10.1016/S0014-5793(04)00072-9 URL |
| [10] |
Hanikenne M, Talke I N, Haydon M J, Lanz C, Nolte A, Motte P, Kroymann J, Weigel D, Kramer U. 2008. Evolution of metal hyperaccumulation required cis-regulatory changes and triplication of HMA4. Nature, 453 (7193):391-395.
doi: 10.1038/nature06877 |
| [11] |
Hussain D, Haydon M J, Wang Y, Wong E, Sherson S M, Young J, Camakaris J, Harper J F, Cobbett C S. 2004. P-type ATPase heavy metal transporters with roles in essential zinc homeostasis in Arabidopsis. Plant Cell, 16 (5):1327-1339.
doi: 10.1105/tpc.020487 pmid: 15100400 |
| [12] |
Kim Y Y, Choi H, Segami S, Cho H T, Martinoia E, Masayoshi, Maeshima, Lee Y. 2009. AtHMA1 contributes to the detoxification of excess Zn(II) in Arabidopsis. Plant Journal, 58 (5):737-753.
doi: 10.1111/tpj.2009.58.issue-5 URL |
| [13] | Liao Tian-lu, Liao Tian-jiang, Lü Lin-hua. 2015. Application of phytoremediation technology in heavy metal contaminated soil. Beijing Agriculture, 6 (17):192. (in Chinese) |
| 廖天录, 廖天江, 吕麟华. 2015. 植物修复技术在重金属污染土壤中的应用. 北京农业, 6 (17):192. | |
| [14] |
Liu L, Yin H, Liu Y, Shen L, Yang X, Zhang D, Li M, Yan M. 2021. Analysis of cadmium-stress-induced microRNAs and their targets reveals bra-miR172b-3p as a potential Cd2+-specific resistance factor in Brassica juncea. Processes, 9 (7):1099.
doi: 10.3390/pr9071099 URL |
| [15] |
Miyadate H, Adachi S, Hiraizumi A, Tezuka K, Nakazawa N, Kawamoto T, Katou K, Kodama I, Sakurai K, Takahashi H, Nagasawa N S, Watanabe A, Fujimura T, Akagi H. 2011. OsHMA3,a P1B-type of ATPase affects root-to-shoot cadmium translocation in rice by mediating efflux into vacuoles. New Phytologist, 189 (1):190-199.
doi: 10.1111/nph.2010.189.issue-1 URL |
| [16] |
Morel M, Crouzet J, Gravot A, Auroy P, Leonhardt N, Vavasseur A, Richaud P. 2009. AtHMA3,a P1B-ATPase allowing Cd/Zn/Co/Pb vacuolar storage in Arabidopsis. Plant Physiology, 149 (2):894-904.
doi: 10.1104/pp.108.130294 URL |
| [17] |
Mourato M P, Moreira I N, Leitao I, Pinto F R, Sales J R, Martins L L. 2015. Effect of heavy metals in plants of the genus Brassica. International Journal of Molecular Sciences, 16 (8):17975-17998.
doi: 10.3390/ijms160817975 pmid: 26247945 |
| [18] |
Palmgren M G, Clemens S, Williams L E, Kramer U, Borg S, Schjorring J K, Sanders D. 2008. Zinc biofortification of cereals:problems and solutions. Trends In Plant Science, 13 (9):464-473.
doi: 10.1016/j.tplants.2008.06.005 pmid: 18701340 |
| [19] |
Papoyan A, Kochian L V. 2004. Identification of Thlaspi caerulescens genes that may be involved in heavy metal hyperaccumulation and tolerance Characterization of a novel heavy metal transporting ATPase. Plant Physiology, 136 (3):3814-3823.
pmid: 15516513 |
| [20] |
Parkin I A P, Gulden S M, Sharpe A G, Lukens L, Trick M, Osborn T C, Lydiate D J. 2005. Segmental structure of the Brassica napus genome based on comparative analysis with Arabidopsis thaliana. Genetics, 171 (2):765-781.
doi: 10.1534/genetics.105.042093 URL |
| [21] |
Puig S, Colas N A, Molina A G, Pearrubia L. 2007. Copper and iron homeostasis in Arabidopsis:responses to metal deficiencies,interactions and biotechnological applications. Plant Cell and Environment, 30 (3):271-290.
doi: 10.1111/j.1365-3040.2007.01642.x URL |
| [22] | Sankaran R P, Ebbs S D. 2008. Transport of Cd and Zn to seeds of Indian mustard(Brassica juncea)during specific stages of plant growth and development. Physiologia Plantarum, 132 (1):69-78. |
| [23] | Sun Tao, Zhang Yu-xiu, Chai Tuan-yao. 2011. Research progress on tolerance of Indian mustard(Brassica juncea L.) to heavy metal. Chinese Journal of Eco-Agriculture, 19 (1):226-234. (in Chinese) |
| 孙涛, 张玉秀, 柴团耀. 2011. 印度芥菜(Brassica juncea L.)重金属耐性机理研究进展. 中国生态农业学报, 19 (1):226-234. | |
| [24] |
Tan P, Zeng C, Wan C, Liu Z, Dong X, Peng J, Lin H, Li M, Liu Z, Yan M. 2021. Metabolic profiles of Brassica juncea roots in response to cadmium stress. Metabolites, 11 (6):383.
doi: 10.3390/metabo11060383 URL |
| [25] |
Verret F, Gravot A, Auroy P, Preveral S, Forestier C, Vavasseur A, Richaud P. 2005. Heavy metal transport by AtHMA4 involves the N-terminal degenerated metal binding domain and the C-terminal His11 stretch. Febs Letters, 579 (6):1515-1522.
doi: 10.1016/j.febslet.2005.01.065 URL |
| [26] |
Verreta F, Gravota A, Auroya P, Leonhardta N, Davidb P, Nussaumeb L, Vavasseura A, Richaud P. 2004. Overexpression of AtHMA4enhances root-to-shoot translocation of zinc and cadmium and plant metal tolerance. Febs Letters, 576 (3):306-312.
doi: 10.1016/j.febslet.2004.09.023 URL |
| [27] |
Wang Ruonan, Nie Lanchun, Zhang Shuangshuang, Cui Qiang, Jia Mingfei. 2019. Research progress on plant resistance to heavy metal stress. Acta Horticulturae Sinica, 46 (1):157-170. (in Chinese)
doi: 10.16420/j.issn.0513-353x.2018-0585 URL |
|
王若男, 乜兰春, 张双双, 崔强, 贾明飞. 2019. 植物抗重金属胁迫研究进展. 园艺学报, 46 (1):157-170.
doi: 10.16420/j.issn.0513-353x.2018-0585 URL |
|
| [28] | Wang Xiao-tong, Li Hao-yang, Xu Ji-chen. 2014. Bioinformatics analysis of the heavy metal transporting ATPase gene family in poplar genome. Plant Physiology Journal, 50 (7):891-900. (in Chinese) |
| 王晓桐, 李昊阳, 徐吉臣. 2014. 毛果杨HMA基因家族的生物信息学分析. 植物生理学报, 50 (7):891-900. | |
| [29] |
Williams L E, Mills R F. 2005. P(1B)-ATPases--an ancient family of transition metal pumps with diverse functions in plants. Trends In Plant Science, 10 (10):491-502.
doi: 10.1016/j.tplants.2005.08.008 pmid: 16154798 |
| [30] |
Wong C K E, Cobbett C S. 2009. HMA P-type ATPases are the major mechanism for root-to-shoot Cd translocation in Arabidopsis thaliana. New Phytologist, 181 (1):71-78.
doi: 10.1111/nph.2009.181.issue-1 URL |
| [31] |
Wong C K E, Jarvis R S, Sherson S M, Cobbett C S. 2009. Functional analysis of the heavy metal binding domains of the Zn/Cd-transporting ATPase,HMA2,in Arabidopsis thaliana. New Phytologist, 181 (1):79-88.
doi: 10.1111/nph.2009.181.issue-1 URL |
| [32] |
Xu D, Lu Z C, Jin K M, Qiu W M, Qiao G R, Han X J, Zhuo R Y. 2020. SPDE:a multi-functional software for sequence processing and data extraction. Bioinformatics, 37 (20):3686-3687.
doi: 10.1093/bioinformatics/btab235 URL |
| [33] | Zhang Fu-gui, Xiao Xin, Yan Gui-xin. 2017. Evolution of HMA gene family in Brassica AC genomes. Chinese Journal of Oil Crop Sciences, 39 (3):294-307. (in Chinese) |
| 张付贵, 肖欣, 闫贵欣. 2017. HMA基因家族在芸薹属AC基因组中的进化分析. 中国油料作物学报, 39 (3):294-307. | |
| [34] | Zhang Yu-xiu, Chai Tuan-yao. 2006. Isolation and function of heavy metal responsive gene in plant. Beijing: China Agriculture Press. (in Chinese) |
| 张玉秀, 柴团耀. 2006. 植物重金属调节基因的分离和功能. 北京: 中国农业出版社. |
| [1] | GAO Pengfei, GAO Bing, FENG Zhenghong, WU Jianhui. Cloning and Cd-resistant Analysis of PsWRKY40 in Potentilla sericea [J]. Acta Horticulturae Sinica, 2023, 50(6): 1269-1283. |
| [2] | ZHANG Kun, SI Binbin, ZHOU Jun, REN Yufeng, ZHANG Xin, XU Wendi, WANG Jiawei, QIAO Shuai, WANG Huiran. Construction of cDNA Library of Apple Rootstock‘Qingzhen 1’Leaf and Screen of MdMLO Genes’ Upstream Regulator [J]. Acta Horticulturae Sinica, 2023, 50(5): 933-946. |
| [3] | CHEN Daozong, LIU Yi, SHEN Wenjie, ZHU Bo, TAN Chen. Identification and Analysis of PAP1/2 Homologous Genes in Brassica rapa,B. oleracea and B. napus [J]. Acta Horticulturae Sinica, 2022, 49(6): 1301-1312. |
| [4] | YANG Tianchen, CHEN Xiaotong, LÜ Ke, ZHANG Di. Expression Pattern and Regulation Mechanism of ApSK3 Dehydrin (Agapanthus praecox)Response to Abiotic Stress and Hormone Signals [J]. Acta Horticulturae Sinica, 2021, 48(8): 1565-1578. |
| [5] | BIAN Kun1,2,SHAO Jiarong2,XU Liwei2,SHEN Ziming1,2,CHEN Wei2,and YANG Zhenfeng2,*. Bioinformatic and Expression Analysis of Fasciclin-like Arabinogalactan Proteins(FLAs)Gene Family in Peach [J]. ACTA HORTICULTURAE SINICA, 2017, 44(4): 653-663. |
| [6] | WANG Honglei,WANG Yanping,YU Tingqiao,LIU Shengli,CHEN Yuzhen,and LU Cunfu*. Characterization of AmCSDP Gene from Ammopiptanthus mongolicus and the Analysis of Cold Resistance in Transgenic Tobacco Plants [J]. ACTA HORTICULTURAE SINICA, 2017, 44(4): 712-722. |
| [7] | CAO Ming-ming,YANG Jia,LI Xiao-yu,LI Yu-hua,and LAN Xing-guo. Analysis and Characterization of Interaction Domain Between ARC1 and Exo70A1 from Ornamental Kale(Brassica oleracea var. acephala) [J]. ACTA HORTICULTURAE SINICA, 2015, 42(4): 791-798. |
| [8] | CHEN Xiu, RAO Xue-Qin, RUAN Xiao-Lei, LIU Fu-Xiu, LI Hua-Ping. Prokaryotic Expression and Antiserum Preparation of Functional Domain of Banana streak virus Coat Protein Gene [J]. ACTA HORTICULTURAE SINICA, 2013, 40(12): 2401-2408. |
| [9] | LIANG Chun-bo;CHEN Yi-li;LI Ying;and HUANG San-wen;. Overexpression of the Potato Late Blight Resistance Genes RB and R3a and Identification of Domains Required for Recognition Specificity of R3a [J]. ACTA HORTICULTURAE SINICA, 2011, 38(1): 87-94. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd